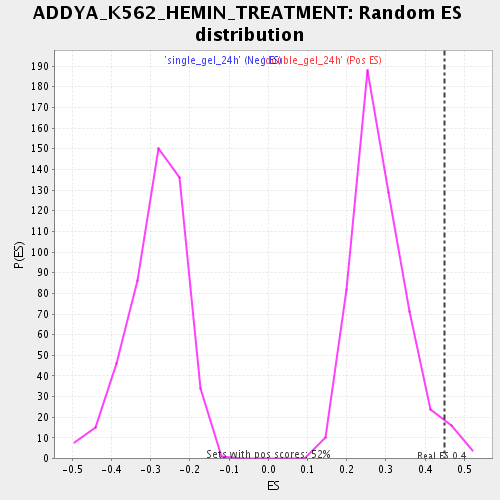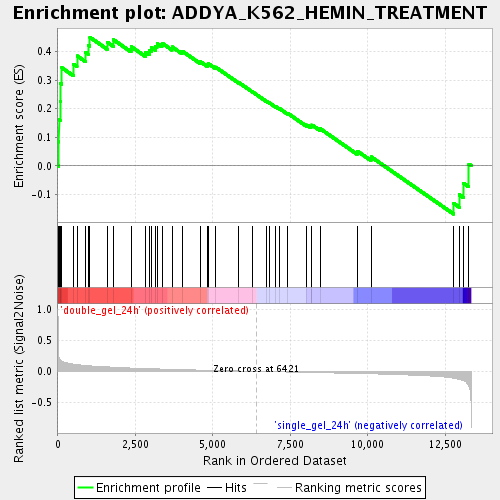
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | ADDYA_K562_HEMIN_TREATMENT |
| Enrichment Score (ES) | 0.44974303 |
| Normalized Enrichment Score (NES) | 1.580418 |
| Nominal p-value | 0.030534351 |
| FDR q-value | 0.22555201 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 25 | 0.248 | 0.0837 | Yes |
| 2 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 35 | 0.229 | 0.1621 | Yes |
| 3 | HERPUD1 | HERPUD1 Entrez, Source | homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 | 74 | 0.189 | 0.2246 | Yes |
| 4 | TXNIP | TXNIP Entrez, Source | thioredoxin interacting protein | 82 | 0.186 | 0.2882 | Yes |
| 5 | RPP25 | RPP25 Entrez, Source | ribonuclease P 25kDa subunit | 125 | 0.168 | 0.3430 | Yes |
| 6 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 502 | 0.118 | 0.3554 | Yes |
| 7 | HSPA5 | HSPA5 Entrez, Source | heat shock 70kDa protein 5 (glucose-regulated protein, 78kDa) | 618 | 0.109 | 0.3844 | Yes |
| 8 | IER3 | IER3 Entrez, Source | immediate early response 3 | 890 | 0.096 | 0.3971 | Yes |
| 9 | INSIG1 | INSIG1 Entrez, Source | insulin induced gene 1 | 978 | 0.092 | 0.4224 | Yes |
| 10 | NEFH | NEFH Entrez, Source | neurofilament, heavy polypeptide 200kDa | 1031 | 0.090 | 0.4497 | Yes |
| 11 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 1593 | 0.073 | 0.4326 | No |
| 12 | OSGIN1 | OSGIN1 Entrez, Source | oxidative stress induced growth inhibitor 1 | 1795 | 0.068 | 0.4409 | No |
| 13 | LMO2 | LMO2 Entrez, Source | LIM domain only 2 (rhombotin-like 1) | 2364 | 0.054 | 0.4169 | No |
| 14 | APOC1 | APOC1 Entrez, Source | apolipoprotein C-I | 2829 | 0.046 | 0.3979 | No |
| 15 | KLF6 | KLF6 Entrez, Source | Kruppel-like factor 6 | 2960 | 0.044 | 0.4032 | No |
| 16 | MAFG | MAFG Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog G (avian) | 3017 | 0.043 | 0.4137 | No |
| 17 | NCF1 | NCF1 Entrez, Source | neutrophil cytosolic factor 1, (chronic granulomatous disease, autosomal 1) | 3150 | 0.041 | 0.4179 | No |
| 18 | SOCS2 | SOCS2 Entrez, Source | suppressor of cytokine signaling 2 | 3200 | 0.040 | 0.4281 | No |
| 19 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 3360 | 0.037 | 0.4290 | No |
| 20 | TAX1BP1 | TAX1BP1 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 1 | 3683 | 0.033 | 0.4161 | No |
| 21 | ATF5 | ATF5 Entrez, Source | activating transcription factor 5 | 4006 | 0.028 | 0.4016 | No |
| 22 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 4604 | 0.021 | 0.3639 | No |
| 23 | JUN | JUN Entrez, Source | jun oncogene | 4841 | 0.018 | 0.3524 | No |
| 24 | FTH1 | FTH1 Entrez, Source | ferritin, heavy polypeptide 1 | 4858 | 0.018 | 0.3574 | No |
| 25 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 5083 | 0.015 | 0.3457 | No |
| 26 | MCL1 | MCL1 Entrez, Source | myeloid cell leukemia sequence 1 (BCL2-related) | 5839 | 0.006 | 0.2911 | No |
| 27 | CTSL | CTSL Entrez, Source | cathepsin L | 6275 | 0.002 | 0.2589 | No |
| 28 | UCP2 | UCP2 Entrez, Source | uncoupling protein 2 (mitochondrial, proton carrier) | 6742 | -0.003 | 0.2250 | No |
| 29 | IER5 | IER5 Entrez, Source | immediate early response 5 | 6813 | -0.004 | 0.2211 | No |
| 30 | CPOX | CPOX Entrez, Source | coproporphyrinogen oxidase | 7027 | -0.006 | 0.2071 | No |
| 31 | ANGPT1 | ANGPT1 Entrez, Source | angiopoietin 1 | 7146 | -0.007 | 0.2006 | No |
| 32 | ITGB5 | ITGB5 Entrez, Source | integrin, beta 5 | 7411 | -0.010 | 0.1841 | No |
| 33 | PTPN7 | PTPN7 Entrez, Source | protein tyrosine phosphatase, non-receptor type 7 | 8035 | -0.016 | 0.1428 | No |
| 34 | HBZ | HBZ Entrez, Source | hemoglobin, zeta | 8170 | -0.017 | 0.1387 | No |
| 35 | FDXR | FDXR Entrez, Source | ferredoxin reductase | 8198 | -0.018 | 0.1428 | No |
| 36 | NFYA | NFYA Entrez, Source | nuclear transcription factor Y, alpha | 8466 | -0.020 | 0.1297 | No |
| 37 | CTSH | CTSH Entrez, Source | cathepsin H | 9659 | -0.033 | 0.0517 | No |
| 38 | ADAM10 | ADAM10 Entrez, Source | ADAM metallopeptidase domain 10 | 10107 | -0.038 | 0.0313 | No |
| 39 | RTN4 | RTN4 Entrez, Source | reticulon 4 | 12773 | -0.109 | -0.1315 | No |
| 40 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 12953 | -0.126 | -0.1013 | No |
| 41 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 13084 | -0.146 | -0.0605 | No |
| 42 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 13262 | -0.231 | 0.0060 | No |