Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
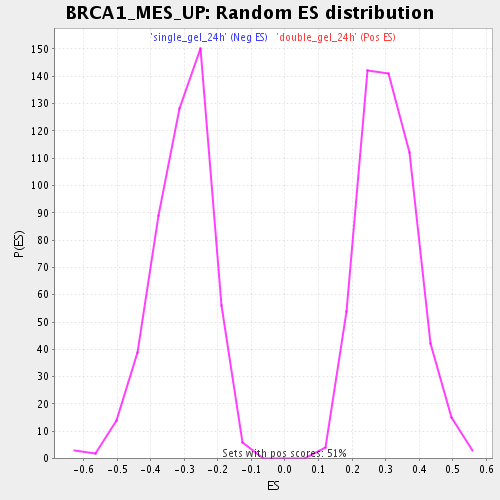
| GeneSet | BRCA1_MES_UP |
| Enrichment Score (ES) | -0.70800227 |
| Normalized Enrichment Score (NES) | -2.2990549 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0010741539 |
| FWER p-Value | 0.0010 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FGFBP1 | FGFBP1 Entrez, Source | fibroblast growth factor binding protein 1 | 3131 | 0.041 | -0.2170 | No |
| 2 | PDIA3 | PDIA3 Entrez, Source | protein disulfide isomerase family A, member 3 | 3281 | 0.039 | -0.2111 | No |
| 3 | DNAJA1 | DNAJA1 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 1 | 4679 | 0.020 | -0.3073 | No |
| 4 | FGFR1 | FGFR1 Entrez, Source | fibroblast growth factor receptor 1 (fms-related tyrosine kinase 2, Pfeiffer syndrome) | 4922 | 0.017 | -0.3180 | No |
| 5 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 5728 | 0.008 | -0.3751 | No |
| 6 | NUMB | NUMB Entrez, Source | numb homolog (Drosophila) | 6445 | -0.000 | -0.4288 | No |
| 7 | LXN | LXN Entrez, Source | latexin | 6581 | -0.002 | -0.4382 | No |
| 8 | TCEA1 | TCEA1 Entrez, Source | transcription elongation factor A (SII), 1 | 7023 | -0.006 | -0.4687 | No |
| 9 | PSMC2 | PSMC2 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 2 | 7074 | -0.006 | -0.4697 | No |
| 10 | EIF3S8 | EIF3S8 Entrez, Source | eukaryotic translation initiation factor 3, subunit 8, 110kDa | 7460 | -0.010 | -0.4942 | No |
| 11 | EIF2S1 | EIF2S1 Entrez, Source | eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa | 7641 | -0.012 | -0.5024 | No |
| 12 | DDX1 | DDX1 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 1 | 7656 | -0.012 | -0.4981 | No |
| 13 | EPRS | EPRS Entrez, Source | glutamyl-prolyl-tRNA synthetase | 7670 | -0.012 | -0.4937 | No |
| 14 | RPA2 | RPA2 Entrez, Source | replication protein A2, 32kDa | 7682 | -0.012 | -0.4892 | No |
| 15 | DDX24 | DDX24 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 24 | 7803 | -0.014 | -0.4922 | No |
| 16 | ARF1 | ARF1 Entrez, Source | ADP-ribosylation factor 1 | 7820 | -0.014 | -0.4873 | No |
| 17 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 7897 | -0.014 | -0.4866 | No |
| 18 | PSMC1 | PSMC1 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 1 | 8459 | -0.020 | -0.5198 | No |
| 19 | DLG1 | DLG1 Entrez, Source | discs, large homolog 1 (Drosophila) | 9446 | -0.031 | -0.5803 | No |
| 20 | ATP6AP1 | ATP6AP1 Entrez, Source | ATPase, H+ transporting, lysosomal accessory protein 1 | 10377 | -0.042 | -0.6318 | No |
| 21 | NEFL | NEFL Entrez, Source | neurofilament, light polypeptide 68kDa | 10506 | -0.044 | -0.6222 | No |
| 22 | CA2 | CA2 Entrez, Source | carbonic anhydrase II | 10802 | -0.048 | -0.6232 | No |
| 23 | ACTC1 | ACTC1 Entrez, Source | actin, alpha, cardiac muscle 1 | 11932 | -0.071 | -0.6767 | Yes |
| 24 | APP | APP Entrez, Source | amyloid beta (A4) precursor protein (peptidase nexin-II, Alzheimer disease) | 12014 | -0.073 | -0.6505 | Yes |
| 25 | VIM | VIM Entrez, Source | vimentin | 12125 | -0.077 | -0.6248 | Yes |
| 26 | ANXA5 | ANXA5 Entrez, Source | annexin A5 | 12733 | -0.105 | -0.6241 | Yes |
| 27 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 12937 | -0.125 | -0.5841 | Yes |
| 28 | ROCK2 | ROCK2 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 2 | 13146 | -0.160 | -0.5290 | Yes |
| 29 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 13255 | -0.224 | -0.4381 | Yes |
| 30 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 13314 | -0.341 | -0.2921 | Yes |
| 31 | TAGLN | TAGLN Entrez, Source | transgelin | 13340 | -0.666 | 0.0002 | Yes |