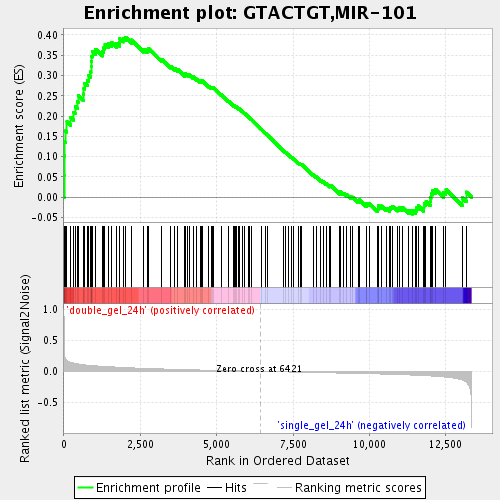
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | double_gel_24h |
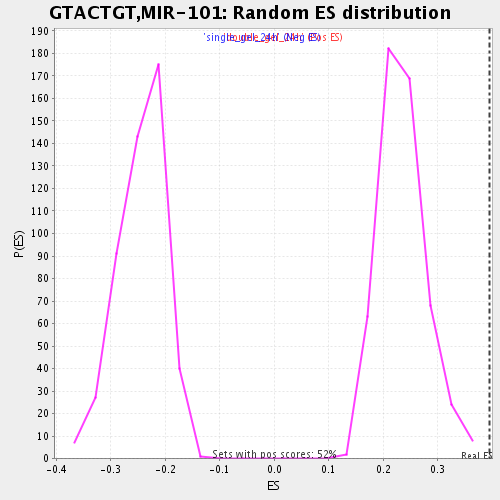
| GeneSet | GTACTGT,MIR-101 |
| Enrichment Score (ES) | 0.39424843 |
| Normalized Enrichment Score (NES) | 1.6764573 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.14301711 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PHLDA1 | PHLDA1 Entrez, Source | pleckstrin homology-like domain, family A, member 1 | 6 | 0.386 | 0.0548 | Yes |
| 2 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 11 | 0.331 | 0.1020 | Yes |
| 3 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 25 | 0.248 | 0.1365 | Yes |
| 4 | PCDH8 | PCDH8 Entrez, Source | protocadherin 8 | 56 | 0.205 | 0.1636 | Yes |
| 5 | LRRN3 | LRRN3 Entrez, Source | leucine rich repeat neuronal 3 | 99 | 0.178 | 0.1859 | Yes |
| 6 | KCNH7 | KCNH7 Entrez, Source | potassium voltage-gated channel, subfamily H (eag-related), member 7 | 224 | 0.145 | 0.1974 | Yes |
| 7 | MARK1 | MARK1 Entrez, Source | MAP/microtubule affinity-regulating kinase 1 | 323 | 0.132 | 0.2089 | Yes |
| 8 | FBN2 | FBN2 Entrez, Source | fibrillin 2 (congenital contractural arachnodactyly) | 368 | 0.128 | 0.2240 | Yes |
| 9 | CEBPA | CEBPA Entrez, Source | CCAAT/enhancer binding protein (C/EBP), alpha | 443 | 0.122 | 0.2359 | Yes |
| 10 | PRKCE | PRKCE Entrez, Source | protein kinase C, epsilon | 467 | 0.120 | 0.2513 | Yes |
| 11 | CDK5R1 | CDK5R1 Entrez, Source | cyclin-dependent kinase 5, regulatory subunit 1 (p35) | 630 | 0.108 | 0.2546 | Yes |
| 12 | ACCN2 | ACCN2 Entrez, Source | amiloride-sensitive cation channel 2, neuronal | 653 | 0.107 | 0.2682 | Yes |
| 13 | KHDRBS3 | KHDRBS3 Entrez, Source | KH domain containing, RNA binding, signal transduction associated 3 | 681 | 0.105 | 0.2813 | Yes |
| 14 | NEGR1 | NEGR1 Entrez, Source | neuronal growth regulator 1 | 775 | 0.100 | 0.2886 | Yes |
| 15 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 816 | 0.099 | 0.2998 | Yes |
| 16 | SLC19A2 | SLC19A2 Entrez, Source | solute carrier family 19 (thiamine transporter), member 2 | 878 | 0.096 | 0.3089 | Yes |
| 17 | SYNGAP1 | SYNGAP1 Entrez, Source | synaptic Ras GTPase activating protein 1 homolog (rat) | 887 | 0.096 | 0.3221 | Yes |
| 18 | IGFBP5 | IGFBP5 Entrez, Source | insulin-like growth factor binding protein 5 | 892 | 0.096 | 0.3355 | Yes |
| 19 | COL10A1 | COL10A1 Entrez, Source | collagen, type X, alpha 1(Schmid metaphyseal chondrodysplasia) | 908 | 0.095 | 0.3480 | Yes |
| 20 | LRRC4 | LRRC4 Entrez, Source | leucine rich repeat containing 4 | 942 | 0.094 | 0.3589 | Yes |
| 21 | RNF38 | RNF38 Entrez, Source | ring finger protein 38 | 1038 | 0.090 | 0.3646 | Yes |
| 22 | THRB | THRB Entrez, Source | thyroid hormone receptor, beta (erythroblastic leukemia viral (v-erb-a) oncogene homolog 2, avian) | 1259 | 0.082 | 0.3597 | Yes |
| 23 | FGA | FGA Entrez, Source | fibrinogen alpha chain | 1302 | 0.080 | 0.3680 | Yes |
| 24 | HAS2 | HAS2 Entrez, Source | hyaluronan synthase 2 | 1343 | 0.079 | 0.3763 | Yes |
| 25 | SLC1A1 | SLC1A1 Entrez, Source | solute carrier family 1 (neuronal/epithelial high affinity glutamate transporter, system Xag), member 1 | 1453 | 0.076 | 0.3790 | Yes |
| 26 | MAGI2 | MAGI2 Entrez, Source | membrane associated guanylate kinase, WW and PDZ domain containing 2 | 1563 | 0.073 | 0.3812 | Yes |
| 27 | XKR6 | XKR6 Entrez, Source | XK, Kell blood group complex subunit-related family, member 6 | 1721 | 0.069 | 0.3793 | Yes |
| 28 | WSB1 | WSB1 Entrez, Source | WD repeat and SOCS box-containing 1 | 1817 | 0.067 | 0.3817 | Yes |
| 29 | RASD2 | RASD2 Entrez, Source | RASD family, member 2 | 1830 | 0.067 | 0.3903 | Yes |
| 30 | BCL2L11 | BCL2L11 Entrez, Source | BCL2-like 11 (apoptosis facilitator) | 1935 | 0.064 | 0.3916 | Yes |
| 31 | RIPK5 | RIPK5 Entrez, Source | receptor interacting protein kinase 5 | 2019 | 0.062 | 0.3942 | Yes |
| 32 | CDH11 | CDH11 Entrez, Source | cadherin 11, type 2, OB-cadherin (osteoblast) | 2216 | 0.057 | 0.3876 | No |
| 33 | MAK | MAK Entrez, Source | male germ cell-associated kinase | 2614 | 0.049 | 0.3647 | No |
| 34 | RAB5A | RAB5A Entrez, Source | RAB5A, member RAS oncogene family | 2725 | 0.048 | 0.3632 | No |
| 35 | LRRN1 | LRRN1 Entrez, Source | leucine rich repeat neuronal 1 | 2774 | 0.047 | 0.3662 | No |
| 36 | SOCS2 | SOCS2 Entrez, Source | suppressor of cytokine signaling 2 | 3200 | 0.040 | 0.3398 | No |
| 37 | KIF1A | KIF1A Entrez, Source | kinesin family member 1A | 3486 | 0.035 | 0.3233 | No |
| 38 | SEMA3G | SEMA3G Entrez, Source | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3G | 3621 | 0.033 | 0.3180 | No |
| 39 | GPR85 | GPR85 Entrez, Source | G protein-coupled receptor 85 | 3730 | 0.032 | 0.3144 | No |
| 40 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 3952 | 0.029 | 0.3019 | No |
| 41 | PREI3 | PREI3 Entrez, Source | preimplantation protein 3 | 3967 | 0.029 | 0.3049 | No |
| 42 | PDE4A | PDE4A Entrez, Source | phosphodiesterase 4A, cAMP-specific (phosphodiesterase E2 dunce homolog, Drosophila) | 4052 | 0.028 | 0.3025 | No |
| 43 | NEUROD1 | NEUROD1 Entrez, Source | neurogenic differentiation 1 | 4116 | 0.027 | 0.3016 | No |
| 44 | ZFAND3 | ZFAND3 Entrez, Source | zinc finger, AN1-type domain 3 | 4225 | 0.026 | 0.2971 | No |
| 45 | CTTNBP2 | CTTNBP2 Entrez, Source | cortactin binding protein 2 | 4345 | 0.024 | 0.2915 | No |
| 46 | SFRS5 | SFRS5 Entrez, Source | splicing factor, arginine/serine-rich 5 | 4458 | 0.022 | 0.2863 | No |
| 47 | MYRIP | MYRIP Entrez, Source | myosin VIIA and Rab interacting protein | 4486 | 0.022 | 0.2874 | No |
| 48 | SOX9 | SOX9 Entrez, Source | SRY (sex determining region Y)-box 9 (campomelic dysplasia, autosomal sex-reversal) | 4534 | 0.021 | 0.2869 | No |
| 49 | DNMT3A | DNMT3A Entrez, Source | DNA (cytosine-5-)-methyltransferase 3 alpha | 4742 | 0.019 | 0.2740 | No |
| 50 | GLRA2 | GLRA2 Entrez, Source | glycine receptor, alpha 2 | 4835 | 0.018 | 0.2697 | No |
| 51 | RXRB | RXRB Entrez, Source | retinoid X receptor, beta | 4872 | 0.018 | 0.2695 | No |
| 52 | GJA1 | GJA1 Entrez, Source | gap junction protein, alpha 1, 43kDa (connexin 43) | 4902 | 0.017 | 0.2698 | No |
| 53 | TRIB1 | TRIB1 Entrez, Source | tribbles homolog 1 (Drosophila) | 5150 | 0.014 | 0.2531 | No |
| 54 | PRKAA1 | PRKAA1 Entrez, Source | protein kinase, AMP-activated, alpha 1 catalytic subunit | 5388 | 0.011 | 0.2368 | No |
| 55 | RAB1A | RAB1A Entrez, Source | RAB1A, member RAS oncogene family | 5561 | 0.009 | 0.2252 | No |
| 56 | MYST2 | MYST2 Entrez, Source | MYST histone acetyltransferase 2 | 5566 | 0.009 | 0.2262 | No |
| 57 | SUB1 | SUB1 Entrez, Source | SUB1 homolog (S. cerevisiae) | 5627 | 0.009 | 0.2230 | No |
| 58 | PPFIA4 | PPFIA4 Entrez, Source | protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 4 | 5649 | 0.009 | 0.2226 | No |
| 59 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 5728 | 0.008 | 0.2178 | No |
| 60 | SPOP | SPOP Entrez, Source | speckle-type POZ protein | 5734 | 0.008 | 0.2185 | No |
| 61 | HNRPF | HNRPF Entrez, Source | heterogeneous nuclear ribonucleoprotein F | 5833 | 0.006 | 0.2120 | No |
| 62 | SOX11 | SOX11 Entrez, Source | SRY (sex determining region Y)-box 11 | 5904 | 0.005 | 0.2075 | No |
| 63 | CAV3 | CAV3 Entrez, Source | caveolin 3 | 6031 | 0.004 | 0.1986 | No |
| 64 | PUM2 | PUM2 Entrez, Source | pumilio homolog 2 (Drosophila) | 6077 | 0.004 | 0.1957 | No |
| 65 | LASS2 | LASS2 Entrez, Source | LAG1 homolog, ceramide synthase 2 (S. cerevisiae) | 6148 | 0.003 | 0.1909 | No |
| 66 | CACNB2 | CACNB2 Entrez, Source | calcium channel, voltage-dependent, beta 2 subunit | 6462 | -0.000 | 0.1672 | No |
| 67 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 6595 | -0.002 | 0.1575 | No |
| 68 | ROBO2 | ROBO2 Entrez, Source | roundabout, axon guidance receptor, homolog 2 (Drosophila) | 6677 | -0.003 | 0.1518 | No |
| 69 | UBE2D3 | UBE2D3 Entrez, Source | ubiquitin-conjugating enzyme E2D 3 (UBC4/5 homolog, yeast) | 7172 | -0.007 | 0.1155 | No |
| 70 | ICK | ICK Entrez, Source | intestinal cell (MAK-like) kinase | 7255 | -0.008 | 0.1104 | No |
| 71 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 7336 | -0.009 | 0.1057 | No |
| 72 | TMED5 | TMED5 Entrez, Source | transmembrane emp24 protein transport domain containing 5 | 7444 | -0.010 | 0.0990 | No |
| 73 | TGFBR1 | TGFBR1 Entrez, Source | transforming growth factor, beta receptor I (activin A receptor type II-like kinase, 53kDa) | 7514 | -0.011 | 0.0953 | No |
| 74 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 7684 | -0.012 | 0.0843 | No |
| 75 | NOVA1 | NOVA1 Entrez, Source | neuro-oncological ventral antigen 1 | 7743 | -0.013 | 0.0818 | No |
| 76 | RBBP7 | RBBP7 Entrez, Source | retinoblastoma binding protein 7 | 7768 | -0.013 | 0.0818 | No |
| 77 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 8152 | -0.017 | 0.0553 | No |
| 78 | LRRFIP2 | LRRFIP2 Entrez, Source | leucine rich repeat (in FLII) interacting protein 2 | 8250 | -0.018 | 0.0506 | No |
| 79 | POMGNT1 | POMGNT1 Entrez, Source | protein O-linked mannose beta1,2-N-acetylglucosaminyltransferase | 8405 | -0.020 | 0.0418 | No |
| 80 | EXOC5 | EXOC5 Entrez, Source | exocyst complex component 5 | 8479 | -0.020 | 0.0392 | No |
| 81 | TXNDC4 | TXNDC4 Entrez, Source | thioredoxin domain containing 4 (endoplasmic reticulum) | 8591 | -0.022 | 0.0339 | No |
| 82 | KPNB1 | KPNB1 Entrez, Source | karyopherin (importin) beta 1 | 8707 | -0.023 | 0.0285 | No |
| 83 | ABCC5 | ABCC5 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 5 | 8737 | -0.023 | 0.0296 | No |
| 84 | UBE2A | UBE2A Entrez, Source | ubiquitin-conjugating enzyme E2A (RAD6 homolog) | 9034 | -0.026 | 0.0110 | No |
| 85 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 9041 | -0.026 | 0.0144 | No |
| 86 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 9157 | -0.028 | 0.0096 | No |
| 87 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 9251 | -0.029 | 0.0067 | No |
| 88 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 9381 | -0.030 | 0.0012 | No |
| 89 | PLCG1 | PLCG1 Entrez, Source | phospholipase C, gamma 1 | 9447 | -0.031 | 0.0007 | No |
| 90 | OGT | OGT Entrez, Source | O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) | 9640 | -0.033 | -0.0091 | No |
| 91 | PIP5K1C | PIP5K1C Entrez, Source | phosphatidylinositol-4-phosphate 5-kinase, type I, gamma | 9663 | -0.034 | -0.0059 | No |
| 92 | SGPL1 | SGPL1 Entrez, Source | sphingosine-1-phosphate lyase 1 | 9899 | -0.036 | -0.0185 | No |
| 93 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 9911 | -0.036 | -0.0142 | No |
| 94 | NDFIP1 | NDFIP1 Entrez, Source | Nedd4 family interacting protein 1 | 10002 | -0.037 | -0.0157 | No |
| 95 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 10273 | -0.040 | -0.0303 | No |
| 96 | QKI | QKI Entrez, Source | quaking homolog, KH domain RNA binding (mouse) | 10287 | -0.041 | -0.0255 | No |
| 97 | STC1 | STC1 Entrez, Source | stanniocalcin 1 | 10297 | -0.041 | -0.0203 | No |
| 98 | ADCY6 | ADCY6 Entrez, Source | adenylate cyclase 6 | 10390 | -0.042 | -0.0213 | No |
| 99 | GNB1 | GNB1 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 1 | 10542 | -0.044 | -0.0264 | No |
| 100 | DR1 | DR1 Entrez, Source | down-regulator of transcription 1, TBP-binding (negative cofactor 2) | 10653 | -0.046 | -0.0282 | No |
| 101 | CLCN3 | CLCN3 Entrez, Source | chloride channel 3 | 10689 | -0.046 | -0.0243 | No |
| 102 | ZNF24 | ZNF24 Entrez, Source | zinc finger protein 24 | 10748 | -0.047 | -0.0219 | No |
| 103 | RAP1B | RAP1B Entrez, Source | RAP1B, member of RAS oncogene family | 10907 | -0.050 | -0.0268 | No |
| 104 | TEAD3 | TEAD3 Entrez, Source | TEA domain family member 3 | 10976 | -0.051 | -0.0247 | No |
| 105 | SPRED1 | SPRED1 Entrez, Source | sprouty-related, EVH1 domain containing 1 | 11076 | -0.052 | -0.0247 | No |
| 106 | SLC30A7 | SLC30A7 Entrez, Source | solute carrier family 30 (zinc transporter), member 7 | 11283 | -0.057 | -0.0321 | No |
| 107 | OTUD4 | OTUD4 Entrez, Source | OTU domain containing 4 | 11407 | -0.059 | -0.0330 | No |
| 108 | REV3L | REV3L Entrez, Source | REV3-like, catalytic subunit of DNA polymerase zeta (yeast) | 11512 | -0.061 | -0.0322 | No |
| 109 | TSC22D1 | TSC22D1 Entrez, Source | TSC22 domain family, member 1 | 11547 | -0.061 | -0.0260 | No |
| 110 | PRPF4B | PRPF4B Entrez, Source | PRP4 pre-mRNA processing factor 4 homolog B (yeast) | 11602 | -0.063 | -0.0211 | No |
| 111 | TMEFF1 | TMEFF1 Entrez, Source | transmembrane protein with EGF-like and two follistatin-like domains 1 | 11783 | -0.067 | -0.0251 | No |
| 112 | DAG1 | DAG1 Entrez, Source | dystroglycan 1 (dystrophin-associated glycoprotein 1) | 11785 | -0.067 | -0.0156 | No |
| 113 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 11849 | -0.069 | -0.0106 | No |
| 114 | DCBLD2 | DCBLD2 Entrez, Source | discoidin, CUB and LCCL domain containing 2 | 11988 | -0.072 | -0.0106 | No |
| 115 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 11994 | -0.073 | -0.0006 | No |
| 116 | APP | APP Entrez, Source | amyloid beta (A4) precursor protein (peptidase nexin-II, Alzheimer disease) | 12014 | -0.073 | 0.0085 | No |
| 117 | FZD6 | FZD6 Entrez, Source | frizzled homolog 6 (Drosophila) | 12054 | -0.074 | 0.0162 | No |
| 118 | BZW2 | BZW2 Entrez, Source | basic leucine zipper and W2 domains 2 | 12150 | -0.078 | 0.0202 | No |
| 119 | PTGS2 | PTGS2 Entrez, Source | prostaglandin-endoperoxide synthase 2 (prostaglandin G/H synthase and cyclooxygenase) | 12422 | -0.088 | 0.0122 | No |
| 120 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 12503 | -0.092 | 0.0193 | No |
| 121 | BICD2 | BICD2 Entrez, Source | bicaudal D homolog 2 (Drosophila) | 13035 | -0.137 | -0.0012 | No |
| 122 | FZD4 | FZD4 Entrez, Source | frizzled homolog 4 (Drosophila) | 13170 | -0.170 | 0.0130 | No |