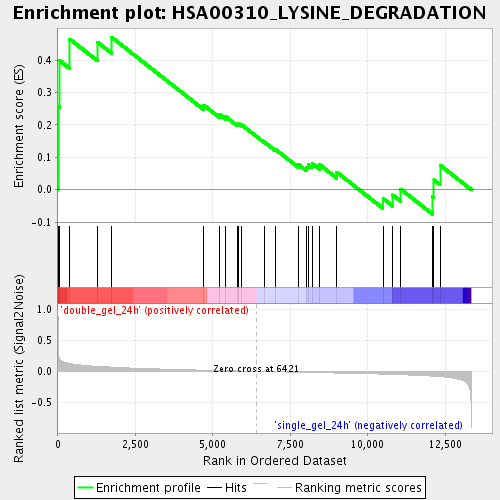
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | HSA00310_LYSINE_DEGRADATION |
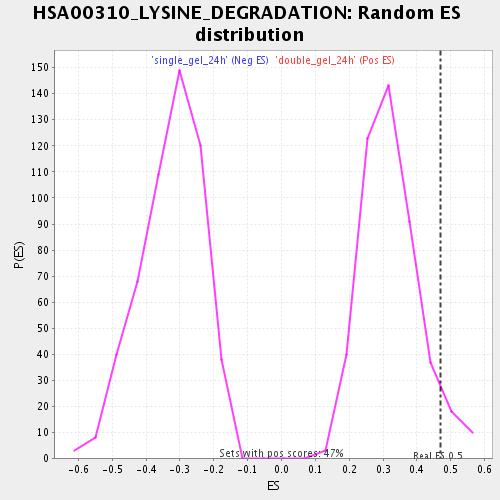
| Enrichment Score (ES) | 0.4701807 |
| Normalized Enrichment Score (NES) | 1.4603825 |
| Nominal p-value | 0.060215052 |
| FDR q-value | 0.29042685 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 7 | 0.362 | 0.2575 | Yes |
| 2 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 57 | 0.202 | 0.3979 | Yes |
| 3 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 379 | 0.127 | 0.4647 | Yes |
| 4 | AKR1B10 | AKR1B10 Entrez, Source | aldo-keto reductase family 1, member B10 (aldose reductase) | 1279 | 0.081 | 0.4550 | Yes |
| 5 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 1732 | 0.069 | 0.4702 | Yes |
| 6 | BBOX1 | BBOX1 Entrez, Source | butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 | 4701 | 0.020 | 0.2613 | No |
| 7 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 5228 | 0.013 | 0.2314 | No |
| 8 | PLOD1 | PLOD1 Entrez, Source | procollagen-lysine 1, 2-oxoglutarate 5-dioxygenase 1 | 5411 | 0.011 | 0.2257 | No |
| 9 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 5790 | 0.007 | 0.2022 | No |
| 10 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 5819 | 0.007 | 0.2047 | No |
| 11 | SHMT2 | SHMT2 Entrez, Source | serine hydroxymethyltransferase 2 (mitochondrial) | 5935 | 0.005 | 0.1998 | No |
| 12 | HSD17B4 | HSD17B4 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 4 | 6653 | -0.002 | 0.1477 | No |
| 13 | PLOD3 | PLOD3 Entrez, Source | procollagen-lysine, 2-oxoglutarate 5-dioxygenase 3 | 7020 | -0.006 | 0.1244 | No |
| 14 | AADAT | AADAT Entrez, Source | aminoadipate aminotransferase | 7758 | -0.013 | 0.0784 | No |
| 15 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 8031 | -0.016 | 0.0694 | No |
| 16 | RDH11 | RDH11 Entrez, Source | retinol dehydrogenase 11 (all-trans/9-cis/11-cis) | 8080 | -0.017 | 0.0776 | No |
| 17 | ALDH1B1 | ALDH1B1 Entrez, Source | aldehyde dehydrogenase 1 family, member B1 | 8208 | -0.018 | 0.0807 | No |
| 18 | HSD3B7 | HSD3B7 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 | 8441 | -0.020 | 0.0776 | No |
| 19 | DLST | DLST Entrez, Source | dihydrolipoamide S-succinyltransferase (E2 component of 2-oxo-glutarate complex) | 9003 | -0.026 | 0.0541 | No |
| 20 | EHMT2 | EHMT2 Entrez, Source | euchromatic histone-lysine N-methyltransferase 2 | 10492 | -0.043 | -0.0268 | No |
| 21 | TMLHE | TMLHE Entrez, Source | trimethyllysine hydroxylase, epsilon | 10808 | -0.048 | -0.0162 | No |
| 22 | SHMT1 | SHMT1 Entrez, Source | serine hydroxymethyltransferase 1 (soluble) | 11060 | -0.052 | 0.0020 | No |
| 23 | ALDH7A1 | ALDH7A1 Entrez, Source | aldehyde dehydrogenase 7 family, member A1 | 12088 | -0.076 | -0.0212 | No |
| 24 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 12137 | -0.077 | 0.0302 | No |
| 25 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 12333 | -0.084 | 0.0758 | No |