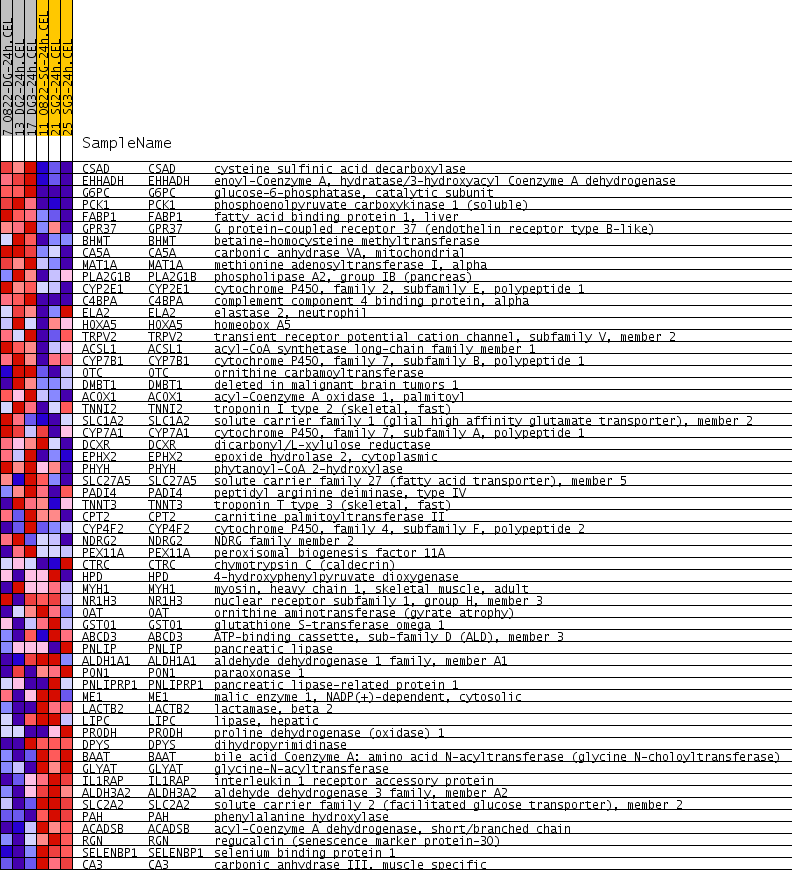
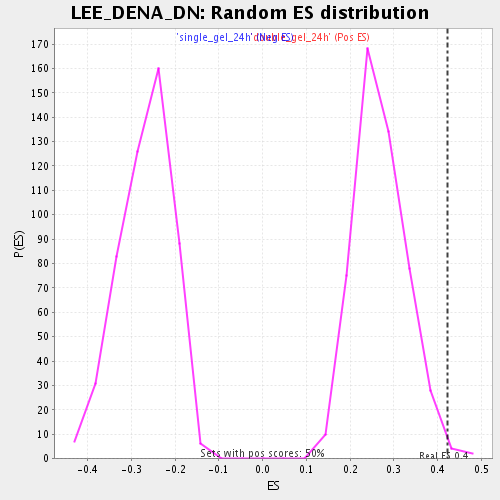
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | LEE_DENA_DN |
| Enrichment Score (ES) | 0.4217981 |
| Normalized Enrichment Score (NES) | 1.5668926 |
| Nominal p-value | 0.006012024 |
| FDR q-value | 0.22999562 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CSAD | CSAD Entrez, Source | cysteine sulfinic acid decarboxylase | 4 | 0.412 | 0.0812 | Yes |
| 2 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 7 | 0.362 | 0.1525 | Yes |
| 3 | G6PC | G6PC Entrez, Source | glucose-6-phosphatase, catalytic subunit | 14 | 0.295 | 0.2105 | Yes |
| 4 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 59 | 0.201 | 0.2469 | Yes |
| 5 | FABP1 | FABP1 Entrez, Source | fatty acid binding protein 1, liver | 63 | 0.197 | 0.2855 | Yes |
| 6 | GPR37 | GPR37 Entrez, Source | G protein-coupled receptor 37 (endothelin receptor type B-like) | 236 | 0.143 | 0.3008 | Yes |
| 7 | BHMT | BHMT Entrez, Source | betaine-homocysteine methyltransferase | 317 | 0.133 | 0.3211 | Yes |
| 8 | CA5A | CA5A Entrez, Source | carbonic anhydrase VA, mitochondrial | 509 | 0.117 | 0.3298 | Yes |
| 9 | MAT1A | MAT1A Entrez, Source | methionine adenosyltransferase I, alpha | 530 | 0.115 | 0.3510 | Yes |
| 10 | PLA2G1B | PLA2G1B Entrez, Source | phospholipase A2, group IB (pancreas) | 597 | 0.111 | 0.3679 | Yes |
| 11 | CYP2E1 | CYP2E1 Entrez, Source | cytochrome P450, family 2, subfamily E, polypeptide 1 | 601 | 0.110 | 0.3894 | Yes |
| 12 | C4BPA | C4BPA Entrez, Source | complement component 4 binding protein, alpha | 712 | 0.104 | 0.4016 | Yes |
| 13 | ELA2 | ELA2 Entrez, Source | elastase 2, neutrophil | 716 | 0.103 | 0.4218 | Yes |
| 14 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 1427 | 0.077 | 0.3835 | No |
| 15 | TRPV2 | TRPV2 Entrez, Source | transient receptor potential cation channel, subfamily V, member 2 | 1634 | 0.072 | 0.3822 | No |
| 16 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 1761 | 0.068 | 0.3862 | No |
| 17 | CYP7B1 | CYP7B1 Entrez, Source | cytochrome P450, family 7, subfamily B, polypeptide 1 | 1796 | 0.068 | 0.3970 | No |
| 18 | OTC | OTC Entrez, Source | ornithine carbamoyltransferase | 2152 | 0.059 | 0.3819 | No |
| 19 | DMBT1 | DMBT1 Entrez, Source | deleted in malignant brain tumors 1 | 2368 | 0.054 | 0.3764 | No |
| 20 | ACOX1 | ACOX1 Entrez, Source | acyl-Coenzyme A oxidase 1, palmitoyl | 2371 | 0.054 | 0.3870 | No |
| 21 | TNNI2 | TNNI2 Entrez, Source | troponin I type 2 (skeletal, fast) | 2401 | 0.054 | 0.3955 | No |
| 22 | SLC1A2 | SLC1A2 Entrez, Source | solute carrier family 1 (glial high affinity glutamate transporter), member 2 | 2558 | 0.051 | 0.3937 | No |
| 23 | CYP7A1 | CYP7A1 Entrez, Source | cytochrome P450, family 7, subfamily A, polypeptide 1 | 2589 | 0.050 | 0.4013 | No |
| 24 | DCXR | DCXR Entrez, Source | dicarbonyl/L-xylulose reductase | 2731 | 0.047 | 0.4001 | No |
| 25 | EPHX2 | EPHX2 Entrez, Source | epoxide hydrolase 2, cytoplasmic | 3215 | 0.040 | 0.3716 | No |
| 26 | PHYH | PHYH Entrez, Source | phytanoyl-CoA 2-hydroxylase | 3341 | 0.038 | 0.3696 | No |
| 27 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 3698 | 0.033 | 0.3492 | No |
| 28 | PADI4 | PADI4 Entrez, Source | peptidyl arginine deiminase, type IV | 3776 | 0.031 | 0.3497 | No |
| 29 | TNNT3 | TNNT3 Entrez, Source | troponin T type 3 (skeletal, fast) | 3949 | 0.029 | 0.3425 | No |
| 30 | CPT2 | CPT2 Entrez, Source | carnitine palmitoyltransferase II | 4497 | 0.022 | 0.3056 | No |
| 31 | CYP4F2 | CYP4F2 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 2 | 5234 | 0.013 | 0.2529 | No |
| 32 | NDRG2 | NDRG2 Entrez, Source | NDRG family member 2 | 5272 | 0.013 | 0.2526 | No |
| 33 | PEX11A | PEX11A Entrez, Source | peroxisomal biogenesis factor 11A | 5307 | 0.013 | 0.2525 | No |
| 34 | CTRC | CTRC Entrez, Source | chymotrypsin C (caldecrin) | 6708 | -0.003 | 0.1477 | No |
| 35 | HPD | HPD Entrez, Source | 4-hydroxyphenylpyruvate dioxygenase | 6761 | -0.003 | 0.1445 | No |
| 36 | MYH1 | MYH1 Entrez, Source | myosin, heavy chain 1, skeletal muscle, adult | 7518 | -0.011 | 0.0897 | No |
| 37 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 7622 | -0.012 | 0.0843 | No |
| 38 | OAT | OAT Entrez, Source | ornithine aminotransferase (gyrate atrophy) | 7916 | -0.015 | 0.0651 | No |
| 39 | GSTO1 | GSTO1 Entrez, Source | glutathione S-transferase omega 1 | 8085 | -0.017 | 0.0557 | No |
| 40 | ABCD3 | ABCD3 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 3 | 8286 | -0.019 | 0.0444 | No |
| 41 | PNLIP | PNLIP Entrez, Source | pancreatic lipase | 8346 | -0.019 | 0.0437 | No |
| 42 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 8802 | -0.024 | 0.0142 | No |
| 43 | PON1 | PON1 Entrez, Source | paraoxonase 1 | 8923 | -0.025 | 0.0102 | No |
| 44 | PNLIPRP1 | PNLIPRP1 Entrez, Source | pancreatic lipase-related protein 1 | 9006 | -0.026 | 0.0091 | No |
| 45 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 9698 | -0.034 | -0.0362 | No |
| 46 | LACTB2 | LACTB2 Entrez, Source | lactamase, beta 2 | 10311 | -0.041 | -0.0742 | No |
| 47 | LIPC | LIPC Entrez, Source | lipase, hepatic | 10652 | -0.046 | -0.0908 | No |
| 48 | PRODH | PRODH Entrez, Source | proline dehydrogenase (oxidase) 1 | 10801 | -0.048 | -0.0924 | No |
| 49 | DPYS | DPYS Entrez, Source | dihydropyrimidinase | 11440 | -0.059 | -0.1287 | No |
| 50 | BAAT | BAAT Entrez, Source | bile acid Coenzyme A: amino acid N-acyltransferase (glycine N-choloyltransferase) | 11730 | -0.065 | -0.1375 | No |
| 51 | GLYAT | GLYAT Entrez, Source | glycine-N-acyltransferase | 11991 | -0.073 | -0.1428 | No |
| 52 | IL1RAP | IL1RAP Entrez, Source | interleukin 1 receptor accessory protein | 12123 | -0.077 | -0.1375 | No |
| 53 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 12137 | -0.077 | -0.1232 | No |
| 54 | SLC2A2 | SLC2A2 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 2 | 12210 | -0.080 | -0.1128 | No |
| 55 | PAH | PAH Entrez, Source | phenylalanine hydroxylase | 12643 | -0.099 | -0.1258 | No |
| 56 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 13150 | -0.162 | -0.1319 | No |
| 57 | RGN | RGN Entrez, Source | regucalcin (senescence marker protein-30) | 13176 | -0.172 | -0.0997 | No |
| 58 | SELENBP1 | SELENBP1 Entrez, Source | selenium binding protein 1 | 13197 | -0.186 | -0.0643 | No |
| 59 | CA3 | CA3 Entrez, Source | carbonic anhydrase III, muscle specific | 13324 | -0.380 | 0.0014 | No |