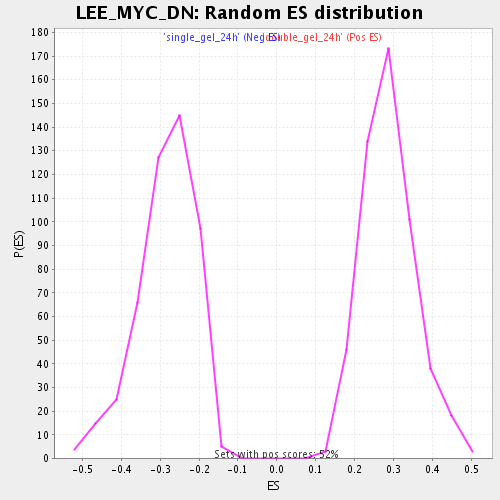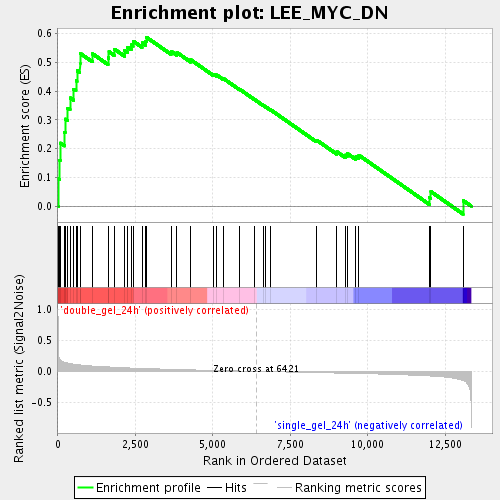
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | LEE_MYC_DN |
| Enrichment Score (ES) | 0.58699447 |
| Normalized Enrichment Score (NES) | 2.0341356 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.022163546 |
| FWER p-Value | 0.238 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | G6PC | G6PC Entrez, Source | glucose-6-phosphatase, catalytic subunit | 14 | 0.295 | 0.0960 | Yes |
| 2 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 59 | 0.201 | 0.1586 | Yes |
| 3 | HERPUD1 | HERPUD1 Entrez, Source | homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 | 74 | 0.189 | 0.2197 | Yes |
| 4 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 215 | 0.147 | 0.2573 | Yes |
| 5 | SATB1 | SATB1 Entrez, Source | special AT-rich sequence binding protein 1 (binds to nuclear matrix/scaffold-associating DNA's) | 234 | 0.143 | 0.3031 | Yes |
| 6 | BHMT | BHMT Entrez, Source | betaine-homocysteine methyltransferase | 317 | 0.133 | 0.3406 | Yes |
| 7 | HSPH1 | HSPH1 Entrez, Source | heat shock 105kDa/110kDa protein 1 | 393 | 0.126 | 0.3763 | Yes |
| 8 | TDO2 | TDO2 Entrez, Source | tryptophan 2,3-dioxygenase | 504 | 0.117 | 0.4065 | Yes |
| 9 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 584 | 0.111 | 0.4371 | Yes |
| 10 | HSPA5 | HSPA5 Entrez, Source | heat shock 70kDa protein 5 (glucose-regulated protein, 78kDa) | 618 | 0.109 | 0.4705 | Yes |
| 11 | C4BPA | C4BPA Entrez, Source | complement component 4 binding protein, alpha | 712 | 0.104 | 0.4975 | Yes |
| 12 | ELA2 | ELA2 Entrez, Source | elastase 2, neutrophil | 716 | 0.103 | 0.5312 | Yes |
| 13 | PCOLCE | PCOLCE Entrez, Source | procollagen C-endopeptidase enhancer | 1118 | 0.087 | 0.5295 | Yes |
| 14 | FGG | FGG Entrez, Source | fibrinogen gamma chain | 1619 | 0.072 | 0.5155 | Yes |
| 15 | ZG16 | ZG16 Entrez, Source | - | 1647 | 0.071 | 0.5368 | Yes |
| 16 | SLC37A4 | SLC37A4 Entrez, Source | solute carrier family 37 (glycerol-6-phosphate transporter), member 4 | 1818 | 0.067 | 0.5461 | Yes |
| 17 | OTC | OTC Entrez, Source | ornithine carbamoyltransferase | 2152 | 0.059 | 0.5404 | Yes |
| 18 | FKBP5 | FKBP5 Entrez, Source | FK506 binding protein 5 | 2248 | 0.056 | 0.5517 | Yes |
| 19 | DMBT1 | DMBT1 Entrez, Source | deleted in malignant brain tumors 1 | 2368 | 0.054 | 0.5606 | Yes |
| 20 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 2425 | 0.053 | 0.5738 | Yes |
| 21 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 2716 | 0.048 | 0.5677 | Yes |
| 22 | ELOVL5 | ELOVL5 Entrez, Source | ELOVL family member 5, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 2833 | 0.046 | 0.5740 | Yes |
| 23 | IGF2 | IGF2 Entrez, Source | insulin-like growth factor 2 (somatomedin A) | 2858 | 0.045 | 0.5870 | Yes |
| 24 | HBP1 | HBP1 Entrez, Source | HMG-box transcription factor 1 | 3652 | 0.033 | 0.5382 | No |
| 25 | CFH | CFH Entrez, Source | complement factor H | 3841 | 0.030 | 0.5341 | No |
| 26 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 4287 | 0.025 | 0.5088 | No |
| 27 | IGFBP2 | IGFBP2 Entrez, Source | insulin-like growth factor binding protein 2, 36kDa | 5034 | 0.016 | 0.4578 | No |
| 28 | HAL | HAL Entrez, Source | histidine ammonia-lyase | 5120 | 0.015 | 0.4563 | No |
| 29 | AQP8 | AQP8 Entrez, Source | aquaporin 8 | 5336 | 0.012 | 0.4441 | No |
| 30 | IL1R1 | IL1R1 Entrez, Source | interleukin 1 receptor, type I | 5860 | 0.006 | 0.4067 | No |
| 31 | HRSP12 | HRSP12 Entrez, Source | heat-responsive protein 12 | 6345 | 0.001 | 0.3706 | No |
| 32 | MEOX2 | MEOX2 Entrez, Source | mesenchyme homeobox 2 | 6638 | -0.002 | 0.3494 | No |
| 33 | CTRC | CTRC Entrez, Source | chymotrypsin C (caldecrin) | 6708 | -0.003 | 0.3452 | No |
| 34 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 6867 | -0.004 | 0.3347 | No |
| 35 | PNLIP | PNLIP Entrez, Source | pancreatic lipase | 8346 | -0.019 | 0.2299 | No |
| 36 | PNLIPRP1 | PNLIPRP1 Entrez, Source | pancreatic lipase-related protein 1 | 9006 | -0.026 | 0.1889 | No |
| 37 | DPYD | DPYD Entrez, Source | dihydropyrimidine dehydrogenase | 9279 | -0.029 | 0.1779 | No |
| 38 | MAPKAPK2 | MAPKAPK2 Entrez, Source | mitogen-activated protein kinase-activated protein kinase 2 | 9335 | -0.029 | 0.1835 | No |
| 39 | RARRES1 | RARRES1 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 1 | 9619 | -0.033 | 0.1730 | No |
| 40 | SFXN1 | SFXN1 Entrez, Source | sideroflexin 1 | 9712 | -0.034 | 0.1773 | No |
| 41 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 11987 | -0.072 | 0.0302 | No |
| 42 | KLF15 | KLF15 Entrez, Source | Kruppel-like factor 15 | 12038 | -0.074 | 0.0507 | No |
| 43 | EFEMP1 | EFEMP1 Entrez, Source | EGF-containing fibulin-like extracellular matrix protein 1 | 13077 | -0.144 | 0.0199 | No |