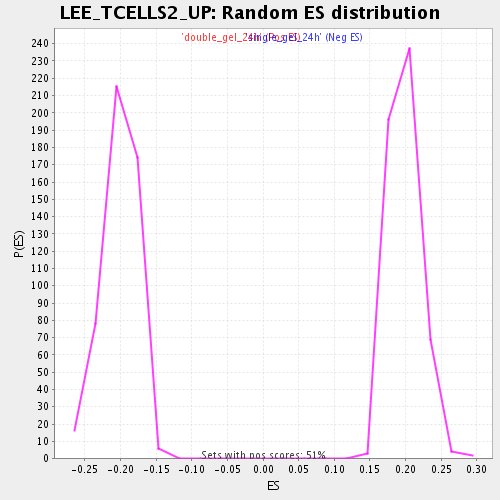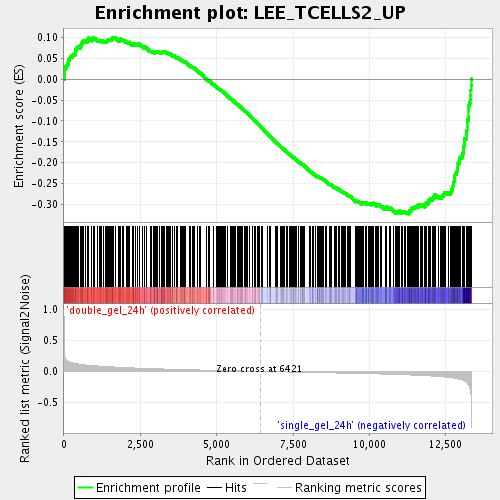
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | LEE_TCELLS2_UP |
| Enrichment Score (ES) | -0.3234 |
| Normalized Enrichment Score (NES) | -1.6048115 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.23308289 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 10 | 0.347 | 0.0113 | No |
| 2 | TUBB3 | TUBB3 Entrez, Source | tubulin, beta 3 | 32 | 0.237 | 0.0179 | No |
| 3 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 35 | 0.229 | 0.0257 | No |
| 4 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 57 | 0.202 | 0.0311 | No |
| 5 | LRRN3 | LRRN3 Entrez, Source | leucine rich repeat neuronal 3 | 99 | 0.178 | 0.0341 | No |
| 6 | PLEKHB1 | PLEKHB1 Entrez, Source | pleckstrin homology domain containing, family B (evectins) member 1 | 117 | 0.170 | 0.0387 | No |
| 7 | IFI44 | IFI44 Entrez, Source | interferon-induced protein 44 | 157 | 0.157 | 0.0411 | No |
| 8 | AQP3 | AQP3 Entrez, Source | aquaporin 3 (Gill blood group) | 158 | 0.157 | 0.0465 | No |
| 9 | TXK | TXK Entrez, Source | TXK tyrosine kinase | 190 | 0.151 | 0.0494 | No |
| 10 | IGF2BP3 | IGF2BP3 Entrez, Source | insulin-like growth factor 2 mRNA binding protein 3 | 199 | 0.149 | 0.0539 | No |
| 11 | SATB1 | SATB1 Entrez, Source | special AT-rich sequence binding protein 1 (binds to nuclear matrix/scaffold-associating DNA's) | 234 | 0.143 | 0.0563 | No |
| 12 | MLLT11 | MLLT11 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 11 | 295 | 0.136 | 0.0563 | No |
| 13 | UPB1 | UPB1 Entrez, Source | ureidopropionase, beta | 300 | 0.135 | 0.0607 | No |
| 14 | RASD1 | RASD1 Entrez, Source | RAS, dexamethasone-induced 1 | 357 | 0.129 | 0.0608 | No |
| 15 | MPP7 | MPP7 Entrez, Source | membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) | 378 | 0.128 | 0.0637 | No |
| 16 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 379 | 0.127 | 0.0681 | No |
| 17 | FIGNL1 | FIGNL1 Entrez, Source | fidgetin-like 1 | 383 | 0.127 | 0.0723 | No |
| 18 | SELL | SELL Entrez, Source | selectin L (lymphocyte adhesion molecule 1) | 417 | 0.125 | 0.0741 | No |
| 19 | ENPP2 | ENPP2 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 2 (autotaxin) | 429 | 0.123 | 0.0775 | No |
| 20 | CTSS | CTSS Entrez, Source | cathepsin S | 462 | 0.120 | 0.0792 | No |
| 21 | IER2 | IER2 Entrez, Source | immediate early response 2 | 528 | 0.115 | 0.0781 | No |
| 22 | IRS1 | IRS1 Entrez, Source | insulin receptor substrate 1 | 549 | 0.114 | 0.0805 | No |
| 23 | SYK | SYK Entrez, Source | spleen tyrosine kinase | 561 | 0.113 | 0.0836 | No |
| 24 | PTCH1 | PTCH1 Entrez, Source | patched homolog 1 (Drosophila) | 566 | 0.112 | 0.0872 | No |
| 25 | MAFK | MAFK Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog K (avian) | 599 | 0.110 | 0.0885 | No |
| 26 | HSPA5 | HSPA5 Entrez, Source | heat shock 70kDa protein 5 (glucose-regulated protein, 78kDa) | 618 | 0.109 | 0.0909 | No |
| 27 | SQLE | SQLE Entrez, Source | squalene epoxidase | 634 | 0.108 | 0.0935 | No |
| 28 | IL6R | IL6R Entrez, Source | interleukin 6 receptor | 695 | 0.104 | 0.0925 | No |
| 29 | IGJ | IGJ Entrez, Source | immunoglobulin J polypeptide, linker protein for immunoglobulin alpha and mu polypeptides | 764 | 0.101 | 0.0907 | No |
| 30 | GIMAP4 | GIMAP4 Entrez, Source | GTPase, IMAP family member 4 | 772 | 0.101 | 0.0937 | No |
| 31 | BAG3 | BAG3 Entrez, Source | BCL2-associated athanogene 3 | 783 | 0.100 | 0.0964 | No |
| 32 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 789 | 0.100 | 0.0995 | No |
| 33 | GIMAP5 | GIMAP5 Entrez, Source | GTPase, IMAP family member 5 | 888 | 0.096 | 0.0952 | No |
| 34 | IER3 | IER3 Entrez, Source | immediate early response 3 | 890 | 0.096 | 0.0984 | No |
| 35 | FDFT1 | FDFT1 Entrez, Source | farnesyl-diphosphate farnesyltransferase 1 | 952 | 0.094 | 0.0969 | No |
| 36 | ABP1 | ABP1 Entrez, Source | amiloride binding protein 1 (amine oxidase (copper-containing)) | 959 | 0.093 | 0.0997 | No |
| 37 | NR4A2 | NR4A2 Entrez, Source | nuclear receptor subfamily 4, group A, member 2 | 1011 | 0.091 | 0.0989 | No |
| 38 | STS | STS Entrez, Source | steroid sulfatase (microsomal), arylsulfatase C, isozyme S | 1112 | 0.087 | 0.0941 | No |
| 39 | CAMK4 | CAMK4 Entrez, Source | calcium/calmodulin-dependent protein kinase IV | 1150 | 0.086 | 0.0942 | No |
| 40 | LAPTM5 | LAPTM5 Entrez, Source | lysosomal associated multispanning membrane protein 5 | 1212 | 0.083 | 0.0924 | No |
| 41 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 1238 | 0.082 | 0.0933 | No |
| 42 | CEBPB | CEBPB Entrez, Source | CCAAT/enhancer binding protein (C/EBP), beta | 1304 | 0.080 | 0.0910 | No |
| 43 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 1348 | 0.079 | 0.0904 | No |
| 44 | GPR56 | GPR56 Entrez, Source | G protein-coupled receptor 56 | 1381 | 0.078 | 0.0906 | No |
| 45 | CD8B | CD8B Entrez, Source | CD8b molecule | 1397 | 0.078 | 0.0922 | No |
| 46 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 1411 | 0.077 | 0.0939 | No |
| 47 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 1441 | 0.077 | 0.0943 | No |
| 48 | CYP2U1 | CYP2U1 Entrez, Source | cytochrome P450, family 2, subfamily U, polypeptide 1 | 1454 | 0.076 | 0.0960 | No |
| 49 | PCSK5 | PCSK5 Entrez, Source | proprotein convertase subtilisin/kexin type 5 | 1480 | 0.075 | 0.0967 | No |
| 50 | KIF4A | KIF4A Entrez, Source | kinesin family member 4A | 1522 | 0.074 | 0.0960 | No |
| 51 | GZMB | GZMB Entrez, Source | granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) | 1567 | 0.073 | 0.0952 | No |
| 52 | ST6GAL1 | ST6GAL1 Entrez, Source | ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 | 1574 | 0.073 | 0.0972 | No |
| 53 | ADORA2A | ADORA2A Entrez, Source | adenosine A2a receptor | 1586 | 0.073 | 0.0989 | No |
| 54 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 1593 | 0.073 | 0.1010 | No |
| 55 | CD38 | CD38 Entrez, Source | CD38 molecule | 1632 | 0.072 | 0.1005 | No |
| 56 | LFNG | LFNG Entrez, Source | lunatic fringe homolog (Drosophila) | 1673 | 0.071 | 0.0998 | No |
| 57 | RUNX2 | RUNX2 Entrez, Source | runt-related transcription factor 2 | 1784 | 0.068 | 0.0936 | No |
| 58 | NUCB2 | NUCB2 Entrez, Source | nucleobindin 2 | 1811 | 0.067 | 0.0940 | No |
| 59 | WSB1 | WSB1 Entrez, Source | WD repeat and SOCS box-containing 1 | 1817 | 0.067 | 0.0959 | No |
| 60 | HES1 | HES1 Entrez, Source | hairy and enhancer of split 1, (Drosophila) | 1845 | 0.066 | 0.0961 | No |
| 61 | AMPH | AMPH Entrez, Source | amphiphysin (Stiff-Man syndrome with breast cancer 128kDa autoantigen) | 1857 | 0.066 | 0.0975 | No |
| 62 | TRAF3IP3 | TRAF3IP3 Entrez, Source | TRAF3 interacting protein 3 | 1910 | 0.064 | 0.0957 | No |
| 63 | TNFSF4 | TNFSF4 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 4 (tax-transcriptionally activated glycoprotein 1, 34kDa) | 1960 | 0.063 | 0.0941 | No |
| 64 | ITGAE | ITGAE Entrez, Source | integrin, alpha E (antigen CD103, human mucosal lymphocyte antigen 1; alpha polypeptide) | 2045 | 0.061 | 0.0897 | No |
| 65 | TMEM66 | TMEM66 Entrez, Source | transmembrane protein 66 | 2082 | 0.060 | 0.0890 | No |
| 66 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 2110 | 0.060 | 0.0890 | No |
| 67 | IL4R | IL4R Entrez, Source | interleukin 4 receptor | 2130 | 0.059 | 0.0896 | No |
| 68 | ICAM2 | ICAM2 Entrez, Source | intercellular adhesion molecule 2 | 2246 | 0.056 | 0.0826 | No |
| 69 | SOCS3 | SOCS3 Entrez, Source | suppressor of cytokine signaling 3 | 2264 | 0.056 | 0.0832 | No |
| 70 | ICOS | ICOS Entrez, Source | inducible T-cell co-stimulator | 2280 | 0.056 | 0.0840 | No |
| 71 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 2289 | 0.056 | 0.0853 | No |
| 72 | PRSS1 | PRSS1 Entrez, Source | protease, serine, 1 (trypsin 1) | 2329 | 0.055 | 0.0842 | No |
| 73 | STK17B | STK17B Entrez, Source | serine/threonine kinase 17b (apoptosis-inducing) | 2348 | 0.055 | 0.0847 | No |
| 74 | ARRDC2 | ARRDC2 Entrez, Source | arrestin domain containing 2 | 2392 | 0.054 | 0.0832 | No |
| 75 | GSTK1 | GSTK1 Entrez, Source | glutathione S-transferase kappa 1 | 2396 | 0.054 | 0.0849 | No |
| 76 | RASGRF2 | RASGRF2 Entrez, Source | Ras protein-specific guanine nucleotide-releasing factor 2 | 2458 | 0.052 | 0.0819 | No |
| 77 | EDG1 | EDG1 Entrez, Source | endothelial differentiation, sphingolipid G-protein-coupled receptor, 1 | 2463 | 0.052 | 0.0834 | No |
| 78 | GALNT7 | GALNT7 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 7 (GalNAc-T7) | 2468 | 0.052 | 0.0849 | No |
| 79 | LRRC8C | LRRC8C Entrez, Source | leucine rich repeat containing 8 family, member C | 2562 | 0.050 | 0.0795 | No |
| 80 | CD93 | CD93 Entrez, Source | CD93 molecule | 2625 | 0.049 | 0.0764 | No |
| 81 | BTG3 | BTG3 Entrez, Source | BTG family, member 3 | 2641 | 0.049 | 0.0769 | No |
| 82 | SCN3A | SCN3A Entrez, Source | sodium channel, voltage-gated, type III, alpha | 2647 | 0.049 | 0.0782 | No |
| 83 | KLF9 | KLF9 Entrez, Source | Kruppel-like factor 9 | 2689 | 0.048 | 0.0767 | No |
| 84 | SLC38A2 | SLC38A2 Entrez, Source | solute carrier family 38, member 2 | 2830 | 0.046 | 0.0674 | No |
| 85 | TCF4 | TCF4 Entrez, Source | transcription factor 4 | 2845 | 0.045 | 0.0679 | No |
| 86 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 2878 | 0.045 | 0.0670 | No |
| 87 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 2916 | 0.044 | 0.0656 | No |
| 88 | CYSLTR1 | CYSLTR1 Entrez, Source | cysteinyl leukotriene receptor 1 | 2972 | 0.043 | 0.0629 | No |
| 89 | DNAJB6 | DNAJB6 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 6 | 2973 | 0.043 | 0.0644 | No |
| 90 | GIMAP6 | GIMAP6 Entrez, Source | GTPase, IMAP family member 6 | 2978 | 0.043 | 0.0656 | No |
| 91 | ARL6IP2 | ARL6IP2 Entrez, Source | ADP-ribosylation factor-like 6 interacting protein 2 | 2995 | 0.043 | 0.0658 | No |
| 92 | CPLX1 | CPLX1 Entrez, Source | complexin 1 | 3026 | 0.043 | 0.0650 | No |
| 93 | SCP2 | SCP2 Entrez, Source | sterol carrier protein 2 | 3048 | 0.042 | 0.0648 | No |
| 94 | IL2RB | IL2RB Entrez, Source | interleukin 2 receptor, beta | 3050 | 0.042 | 0.0662 | No |
| 95 | SLC16A10 | SLC16A10 Entrez, Source | solute carrier family 16, member 10 (aromatic amino acid transporter) | 3067 | 0.042 | 0.0664 | No |
| 96 | CD200 | CD200 Entrez, Source | CD200 molecule | 3078 | 0.042 | 0.0671 | No |
| 97 | RABEP1 | RABEP1 Entrez, Source | rabaptin, RAB GTPase binding effector protein 1 | 3117 | 0.041 | 0.0656 | No |
| 98 | SMAP1L | SMAP1L Entrez, Source | stromal membrane-associated protein 1-like | 3177 | 0.040 | 0.0624 | No |
| 99 | IFITM2 | IFITM2 Entrez, Source | interferon induced transmembrane protein 2 (1-8D) | 3184 | 0.040 | 0.0633 | No |
| 100 | ENC1 | ENC1 Entrez, Source | ectodermal-neural cortex (with BTB-like domain) | 3196 | 0.040 | 0.0639 | No |
| 101 | SOCS2 | SOCS2 Entrez, Source | suppressor of cytokine signaling 2 | 3200 | 0.040 | 0.0650 | No |
| 102 | EPHX2 | EPHX2 Entrez, Source | epoxide hydrolase 2, cytoplasmic | 3215 | 0.040 | 0.0653 | No |
| 103 | TIMP2 | TIMP2 Entrez, Source | TIMP metallopeptidase inhibitor 2 | 3227 | 0.040 | 0.0658 | No |
| 104 | NUP107 | NUP107 Entrez, Source | nucleoporin 107kDa | 3252 | 0.039 | 0.0653 | No |
| 105 | CD6 | CD6 Entrez, Source | CD6 molecule | 3260 | 0.039 | 0.0661 | No |
| 106 | VDP | VDP Entrez, Source | - | 3277 | 0.039 | 0.0663 | No |
| 107 | PFKFB3 | PFKFB3 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3 | 3285 | 0.039 | 0.0671 | No |
| 108 | CDKN3 | CDKN3 Entrez, Source | cyclin-dependent kinase inhibitor 3 (CDK2-associated dual specificity phosphatase) | 3296 | 0.038 | 0.0676 | No |
| 109 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 3360 | 0.037 | 0.0640 | No |
| 110 | DBN1 | DBN1 Entrez, Source | drebrin 1 | 3383 | 0.037 | 0.0636 | No |
| 111 | MAL | MAL Entrez, Source | mal, T-cell differentiation protein | 3407 | 0.037 | 0.0631 | No |
| 112 | CD24 | CD24 Entrez, Source | CD24 molecule | 3463 | 0.036 | 0.0600 | No |
| 113 | PTGER2 | PTGER2 Entrez, Source | prostaglandin E receptor 2 (subtype EP2), 53kDa | 3478 | 0.036 | 0.0602 | No |
| 114 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 3481 | 0.035 | 0.0612 | No |
| 115 | ATP10A | ATP10A Entrez, Source | ATPase, Class V, type 10A | 3557 | 0.034 | 0.0566 | No |
| 116 | ZFP36L1 | ZFP36L1 Entrez, Source | zinc finger protein 36, C3H type-like 1 | 3560 | 0.034 | 0.0577 | No |
| 117 | DUSP4 | DUSP4 Entrez, Source | dual specificity phosphatase 4 | 3568 | 0.034 | 0.0583 | No |
| 118 | PXMP2 | PXMP2 Entrez, Source | peroxisomal membrane protein 2, 22kDa | 3604 | 0.034 | 0.0568 | No |
| 119 | TAX1BP1 | TAX1BP1 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 1 | 3683 | 0.033 | 0.0518 | No |
| 120 | HTR2B | HTR2B Entrez, Source | 5-hydroxytryptamine (serotonin) receptor 2B | 3684 | 0.033 | 0.0530 | No |
| 121 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 3699 | 0.033 | 0.0530 | No |
| 122 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 3724 | 0.032 | 0.0523 | No |
| 123 | GAS5 | GAS5 Entrez, Source | growth arrest-specific 5 | 3804 | 0.031 | 0.0472 | No |
| 124 | APBA2 | APBA2 Entrez, Source | amyloid beta (A4) precursor protein-binding, family A, member 2 (X11-like) | 3832 | 0.031 | 0.0462 | No |
| 125 | GP5 | GP5 Entrez, Source | glycoprotein V (platelet) | 3840 | 0.030 | 0.0467 | No |
| 126 | REPIN1 | REPIN1 Entrez, Source | replication initiator 1 | 3882 | 0.030 | 0.0445 | No |
| 127 | NANP | NANP Entrez, Source | N-acetylneuraminic acid phosphatase | 3910 | 0.030 | 0.0434 | No |
| 128 | PTPRC | PTPRC Entrez, Source | protein tyrosine phosphatase, receptor type, C | 3940 | 0.029 | 0.0422 | No |
| 129 | IL10RA | IL10RA Entrez, Source | interleukin 10 receptor, alpha | 3942 | 0.029 | 0.0431 | No |
| 130 | CEACAM1 | CEACAM1 Entrez, Source | carcinoembryonic antigen-related cell adhesion molecule 1 (biliary glycoprotein) | 3951 | 0.029 | 0.0435 | No |
| 131 | PHACTR2 | PHACTR2 Entrez, Source | phosphatase and actin regulator 2 | 3976 | 0.029 | 0.0427 | No |
| 132 | LAP3 | LAP3 Entrez, Source | leucine aminopeptidase 3 | 4119 | 0.027 | 0.0326 | No |
| 133 | GMPS | GMPS Entrez, Source | guanine monphosphate synthetase | 4135 | 0.026 | 0.0323 | No |
| 134 | ELOVL6 | ELOVL6 Entrez, Source | ELOVL family member 6, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 4158 | 0.026 | 0.0315 | No |
| 135 | CD28 | CD28 Entrez, Source | CD28 molecule | 4201 | 0.026 | 0.0291 | No |
| 136 | JAK3 | JAK3 Entrez, Source | Janus kinase 3 (a protein tyrosine kinase, leukocyte) | 4223 | 0.026 | 0.0284 | No |
| 137 | CUTL1 | CUTL1 Entrez, Source | cut-like 1, CCAAT displacement protein (Drosophila) | 4235 | 0.025 | 0.0284 | No |
| 138 | FBXL12 | FBXL12 Entrez, Source | F-box and leucine-rich repeat protein 12 | 4261 | 0.025 | 0.0273 | No |
| 139 | ST6GALNAC1 | ST6GALNAC1 Entrez, Source | ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 1 | 4373 | 0.023 | 0.0195 | No |
| 140 | STK38 | STK38 Entrez, Source | serine/threonine kinase 38 | 4377 | 0.023 | 0.0201 | No |
| 141 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 4450 | 0.023 | 0.0153 | No |
| 142 | ADK | ADK Entrez, Source | adenosine kinase | 4457 | 0.022 | 0.0156 | No |
| 143 | BCL2L1 | BCL2L1 Entrez, Source | BCL2-like 1 | 4653 | 0.020 | 0.0011 | No |
| 144 | GPRASP1 | GPRASP1 Entrez, Source | G protein-coupled receptor associated sorting protein 1 | 4675 | 0.020 | 0.0002 | No |
| 145 | USP11 | USP11 Entrez, Source | ubiquitin specific peptidase 11 | 4726 | 0.019 | -0.0030 | No |
| 146 | SLC7A3 | SLC7A3 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 3 | 4745 | 0.019 | -0.0038 | No |
| 147 | IRF8 | IRF8 Entrez, Source | interferon regulatory factor 8 | 4750 | 0.019 | -0.0034 | No |
| 148 | GAS7 | GAS7 Entrez, Source | growth arrest-specific 7 | 4771 | 0.019 | -0.0043 | No |
| 149 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 4780 | 0.019 | -0.0043 | No |
| 150 | PRKCA | PRKCA Entrez, Source | protein kinase C, alpha | 4883 | 0.018 | -0.0116 | No |
| 151 | PRKAR2A | PRKAR2A Entrez, Source | protein kinase, cAMP-dependent, regulatory, type II, alpha | 4888 | 0.018 | -0.0113 | No |
| 152 | NOG | NOG Entrez, Source | noggin | 4995 | 0.016 | -0.0190 | No |
| 153 | PDE4DIP | PDE4DIP Entrez, Source | phosphodiesterase 4D interacting protein (myomegalin) | 5022 | 0.016 | -0.0205 | No |
| 154 | GUCY1A3 | GUCY1A3 Entrez, Source | guanylate cyclase 1, soluble, alpha 3 | 5030 | 0.016 | -0.0205 | No |
| 155 | NELL2 | NELL2 Entrez, Source | NEL-like 2 (chicken) | 5064 | 0.015 | -0.0225 | No |
| 156 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 5083 | 0.015 | -0.0234 | No |
| 157 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 5100 | 0.015 | -0.0241 | No |
| 158 | STAT4 | STAT4 Entrez, Source | signal transducer and activator of transcription 4 | 5129 | 0.015 | -0.0258 | No |
| 159 | TRIB1 | TRIB1 Entrez, Source | tribbles homolog 1 (Drosophila) | 5150 | 0.014 | -0.0268 | No |
| 160 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 5161 | 0.014 | -0.0271 | No |
| 161 | GZMA | GZMA Entrez, Source | granzyme A (granzyme 1, cytotoxic T-lymphocyte-associated serine esterase 3) | 5197 | 0.014 | -0.0294 | No |
| 162 | LCP2 | LCP2 Entrez, Source | lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa) | 5226 | 0.013 | -0.0311 | No |
| 163 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 5269 | 0.013 | -0.0339 | No |
| 164 | LRRC59 | LRRC59 Entrez, Source | leucine rich repeat containing 59 | 5300 | 0.013 | -0.0358 | No |
| 165 | GZMK | GZMK Entrez, Source | granzyme K (granzyme 3; tryptase II) | 5359 | 0.012 | -0.0399 | No |
| 166 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 5438 | 0.011 | -0.0456 | No |
| 167 | UGP2 | UGP2 Entrez, Source | UDP-glucose pyrophosphorylase 2 | 5449 | 0.011 | -0.0460 | No |
| 168 | PCBP3 | PCBP3 Entrez, Source | poly(rC) binding protein 3 | 5478 | 0.010 | -0.0478 | No |
| 169 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 5488 | 0.010 | -0.0481 | No |
| 170 | ADA | ADA Entrez, Source | adenosine deaminase | 5503 | 0.010 | -0.0489 | No |
| 171 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 5517 | 0.010 | -0.0495 | No |
| 172 | ITPR2 | ITPR2 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 2 | 5533 | 0.010 | -0.0503 | No |
| 173 | TSC22D3 | TSC22D3 Entrez, Source | TSC22 domain family, member 3 | 5572 | 0.009 | -0.0530 | No |
| 174 | PSMD6 | PSMD6 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 6 | 5605 | 0.009 | -0.0552 | No |
| 175 | EIF3S10 | EIF3S10 Entrez, Source | eukaryotic translation initiation factor 3, subunit 10 theta, 150/170kDa | 5672 | 0.008 | -0.0600 | No |
| 176 | PDK1 | PDK1 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 1 | 5686 | 0.008 | -0.0607 | No |
| 177 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 5696 | 0.008 | -0.0612 | No |
| 178 | HNRPC | HNRPC Entrez, Source | heterogeneous nuclear ribonucleoprotein C (C1/C2) | 5697 | 0.008 | -0.0609 | No |
| 179 | HCST | HCST Entrez, Source | hematopoietic cell signal transducer | 5756 | 0.007 | -0.0651 | No |
| 180 | BZRAP1 | BZRAP1 Entrez, Source | benzodiazapine receptor (peripheral) associated protein 1 | 5774 | 0.007 | -0.0662 | No |
| 181 | UBE2F | UBE2F Entrez, Source | ubiquitin-conjugating enzyme E2F (putative) | 5778 | 0.007 | -0.0662 | No |
| 182 | CD8A | CD8A Entrez, Source | CD8a molecule | 5814 | 0.007 | -0.0687 | No |
| 183 | DPEP2 | DPEP2 Entrez, Source | dipeptidase 2 | 5824 | 0.006 | -0.0692 | No |
| 184 | MCL1 | MCL1 Entrez, Source | myeloid cell leukemia sequence 1 (BCL2-related) | 5839 | 0.006 | -0.0700 | No |
| 185 | SERPINE2 | SERPINE2 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 2 | 5843 | 0.006 | -0.0701 | No |
| 186 | NPC2 | NPC2 Entrez, Source | Niemann-Pick disease, type C2 | 5920 | 0.005 | -0.0758 | No |
| 187 | RGC32 | RGC32 Entrez, Source | - | 5929 | 0.005 | -0.0762 | No |
| 188 | KATNAL1 | KATNAL1 Entrez, Source | katanin p60 subunit A-like 1 | 5940 | 0.005 | -0.0768 | No |
| 189 | TSHR | TSHR Entrez, Source | thyroid stimulating hormone receptor | 5969 | 0.005 | -0.0788 | No |
| 190 | CDCA2 | CDCA2 Entrez, Source | cell division cycle associated 2 | 6018 | 0.004 | -0.0824 | No |
| 191 | FOXO1A | FOXO1A Entrez, Source | forkhead box O1A (rhabdomyosarcoma) | 6085 | 0.004 | -0.0874 | No |
| 192 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 6161 | 0.003 | -0.0932 | No |
| 193 | METAP2 | METAP2 Entrez, Source | methionyl aminopeptidase 2 | 6178 | 0.003 | -0.0943 | No |
| 194 | FXYD6 | FXYD6 Entrez, Source | FXYD domain containing ion transport regulator 6 | 6228 | 0.002 | -0.0980 | No |
| 195 | ATP1B3 | ATP1B3 Entrez, Source | ATPase, Na+/K+ transporting, beta 3 polypeptide | 6238 | 0.002 | -0.0987 | No |
| 196 | CTSL | CTSL Entrez, Source | cathepsin L | 6275 | 0.002 | -0.1014 | No |
| 197 | FBXL20 | FBXL20 Entrez, Source | F-box and leucine-rich repeat protein 20 | 6285 | 0.001 | -0.1021 | No |
| 198 | ADARB1 | ADARB1 Entrez, Source | adenosine deaminase, RNA-specific, B1 (RED1 homolog rat) | 6339 | 0.001 | -0.1062 | No |
| 199 | LITAF | LITAF Entrez, Source | lipopolysaccharide-induced TNF factor | 6376 | 0.000 | -0.1090 | No |
| 200 | BCL11A | BCL11A Entrez, Source | B-cell CLL/lymphoma 11A (zinc finger protein) | 6391 | 0.000 | -0.1100 | No |
| 201 | TUBB2C | TUBB2C Entrez, Source | tubulin, beta 2C | 6401 | 0.000 | -0.1107 | No |
| 202 | ST3GAL5 | ST3GAL5 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 5 | 6457 | -0.000 | -0.1150 | No |
| 203 | CDC42SE1 | CDC42SE1 Entrez, Source | CDC42 small effector 1 | 6481 | -0.001 | -0.1168 | No |
| 204 | INPP4B | INPP4B Entrez, Source | inositol polyphosphate-4-phosphatase, type II, 105kDa | 6486 | -0.001 | -0.1170 | No |
| 205 | AURKB | AURKB Entrez, Source | aurora kinase B | 6500 | -0.001 | -0.1180 | No |
| 206 | MKNK2 | MKNK2 Entrez, Source | MAP kinase interacting serine/threonine kinase 2 | 6661 | -0.003 | -0.1304 | No |
| 207 | GAPDH | GAPDH Entrez, Source | glyceraldehyde-3-phosphate dehydrogenase | 6726 | -0.003 | -0.1353 | No |
| 208 | LEPROTL1 | LEPROTL1 Entrez, Source | leptin receptor overlapping transcript-like 1 | 6749 | -0.003 | -0.1369 | No |
| 209 | HNRPAB | HNRPAB Entrez, Source | heterogeneous nuclear ribonucleoprotein A/B | 6758 | -0.003 | -0.1374 | No |
| 210 | MX1 | MX1 Entrez, Source | myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) | 6762 | -0.003 | -0.1375 | No |
| 211 | YWHAE | YWHAE Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, epsilon polypeptide | 6914 | -0.005 | -0.1491 | No |
| 212 | TTL | TTL Entrez, Source | tubulin tyrosine ligase | 6964 | -0.005 | -0.1527 | No |
| 213 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 6967 | -0.005 | -0.1527 | No |
| 214 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 6990 | -0.006 | -0.1542 | No |
| 215 | SLA | SLA Entrez, Source | Src-like-adaptor | 6999 | -0.006 | -0.1546 | No |
| 216 | INSIG2 | INSIG2 Entrez, Source | insulin induced gene 2 | 7082 | -0.006 | -0.1608 | No |
| 217 | CCNB1 | CCNB1 Entrez, Source | cyclin B1 | 7114 | -0.007 | -0.1629 | No |
| 218 | GARS | GARS Entrez, Source | glycyl-tRNA synthetase | 7116 | -0.007 | -0.1628 | No |
| 219 | RPL18 | RPL18 Entrez, Source | ribosomal protein L18 | 7152 | -0.007 | -0.1653 | No |
| 220 | IFITM3 | IFITM3 Entrez, Source | interferon induced transmembrane protein 3 (1-8U) | 7163 | -0.007 | -0.1658 | No |
| 221 | TCEA3 | TCEA3 Entrez, Source | transcription elongation factor A (SII), 3 | 7173 | -0.007 | -0.1662 | No |
| 222 | ITGB7 | ITGB7 Entrez, Source | integrin, beta 7 | 7193 | -0.007 | -0.1674 | No |
| 223 | SCARB2 | SCARB2 Entrez, Source | scavenger receptor class B, member 2 | 7224 | -0.008 | -0.1695 | No |
| 224 | ADRBK2 | ADRBK2 Entrez, Source | adrenergic, beta, receptor kinase 2 | 7276 | -0.008 | -0.1732 | No |
| 225 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 7292 | -0.009 | -0.1741 | No |
| 226 | SNX10 | SNX10 Entrez, Source | sorting nexin 10 | 7325 | -0.009 | -0.1762 | No |
| 227 | CDCA1 | CDCA1 Entrez, Source | cell division cycle associated 1 | 7368 | -0.009 | -0.1792 | No |
| 228 | RPL13 | RPL13 Entrez, Source | ribosomal protein L13 | 7403 | -0.010 | -0.1815 | No |
| 229 | RPS5 | RPS5 Entrez, Source | ribosomal protein S5 | 7442 | -0.010 | -0.1841 | No |
| 230 | RPL35A | RPL35A Entrez, Source | ribosomal protein L35a | 7449 | -0.010 | -0.1842 | No |
| 231 | SMAD1 | SMAD1 Entrez, Source | SMAD, mothers against DPP homolog 1 (Drosophila) | 7489 | -0.010 | -0.1869 | No |
| 232 | TGFBR1 | TGFBR1 Entrez, Source | transforming growth factor, beta receptor I (activin A receptor type II-like kinase, 53kDa) | 7514 | -0.011 | -0.1884 | No |
| 233 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 7517 | -0.011 | -0.1882 | No |
| 234 | ASRGL1 | ASRGL1 Entrez, Source | asparaginase like 1 | 7528 | -0.011 | -0.1886 | No |
| 235 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 7532 | -0.011 | -0.1884 | No |
| 236 | RASA3 | RASA3 Entrez, Source | RAS p21 protein activator 3 | 7585 | -0.011 | -0.1921 | No |
| 237 | SCCPDH | SCCPDH Entrez, Source | saccharopine dehydrogenase (putative) | 7589 | -0.011 | -0.1919 | No |
| 238 | ATIC | ATIC Entrez, Source | 5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase | 7625 | -0.012 | -0.1942 | No |
| 239 | ATP5B | ATP5B Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide | 7688 | -0.012 | -0.1986 | No |
| 240 | SORD | SORD Entrez, Source | sorbitol dehydrogenase | 7689 | -0.012 | -0.1982 | No |
| 241 | TCP1 | TCP1 Entrez, Source | t-complex 1 | 7728 | -0.013 | -0.2007 | No |
| 242 | BIN1 | BIN1 Entrez, Source | bridging integrator 1 | 7729 | -0.013 | -0.2003 | No |
| 243 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 7778 | -0.013 | -0.2035 | No |
| 244 | RPL28 | RPL28 Entrez, Source | ribosomal protein L28 | 7793 | -0.013 | -0.2042 | No |
| 245 | NEDD9 | NEDD9 Entrez, Source | neural precursor cell expressed, developmentally down-regulated 9 | 7794 | -0.013 | -0.2037 | No |
| 246 | MTAC2D1 | MTAC2D1 Entrez, Source | membrane targeting (tandem) C2 domain containing 1 | 7815 | -0.014 | -0.2048 | No |
| 247 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 7827 | -0.014 | -0.2052 | No |
| 248 | CD48 | CD48 Entrez, Source | CD48 molecule | 7841 | -0.014 | -0.2057 | No |
| 249 | CCT6A | CCT6A Entrez, Source | chaperonin containing TCP1, subunit 6A (zeta 1) | 7874 | -0.014 | -0.2077 | No |
| 250 | ARHGEF7 | ARHGEF7 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 7 | 7877 | -0.014 | -0.2073 | No |
| 251 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 8031 | -0.016 | -0.2187 | No |
| 252 | SKAP1 | SKAP1 Entrez, Source | src kinase associated phosphoprotein 1 | 8081 | -0.017 | -0.2219 | No |
| 253 | THEM4 | THEM4 Entrez, Source | thioesterase superfamily member 4 | 8121 | -0.017 | -0.2244 | No |
| 254 | CRBN | CRBN Entrez, Source | cereblon | 8123 | -0.017 | -0.2239 | No |
| 255 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 8126 | -0.017 | -0.2234 | No |
| 256 | PPP2CA | PPP2CA Entrez, Source | protein phosphatase 2 (formerly 2A), catalytic subunit, alpha isoform | 8183 | -0.017 | -0.2272 | No |
| 257 | CAPG | CAPG Entrez, Source | capping protein (actin filament), gelsolin-like | 8248 | -0.018 | -0.2315 | No |
| 258 | ABCD3 | ABCD3 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 3 | 8286 | -0.019 | -0.2338 | No |
| 259 | LYST | LYST Entrez, Source | lysosomal trafficking regulator | 8300 | -0.019 | -0.2341 | No |
| 260 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 8303 | -0.019 | -0.2336 | No |
| 261 | CLEC2D | CLEC2D Entrez, Source | C-type lectin domain family 2, member D | 8305 | -0.019 | -0.2330 | No |
| 262 | C1ORF121 | C1ORF121 Entrez, Source | chromosome 1 open reading frame 121 | 8323 | -0.019 | -0.2337 | No |
| 263 | TTC7A | TTC7A Entrez, Source | tetratricopeptide repeat domain 7A | 8353 | -0.019 | -0.2353 | No |
| 264 | C1QBP | C1QBP Entrez, Source | complement component 1, q subcomponent binding protein | 8371 | -0.019 | -0.2359 | No |
| 265 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 8374 | -0.019 | -0.2354 | No |
| 266 | CTSB | CTSB Entrez, Source | cathepsin B | 8382 | -0.019 | -0.2353 | No |
| 267 | E2F8 | E2F8 Entrez, Source | E2F transcription factor 8 | 8431 | -0.020 | -0.2383 | No |
| 268 | VDAC1 | VDAC1 Entrez, Source | voltage-dependent anion channel 1 | 8435 | -0.020 | -0.2379 | No |
| 269 | SSR4 | SSR4 Entrez, Source | signal sequence receptor, delta (translocon-associated protein delta) | 8447 | -0.020 | -0.2380 | No |
| 270 | CXXC5 | CXXC5 Entrez, Source | CXXC finger 5 | 8472 | -0.020 | -0.2392 | No |
| 271 | CSDA | CSDA Entrez, Source | cold shock domain protein A | 8491 | -0.021 | -0.2399 | No |
| 272 | PIGK | PIGK Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class K | 8508 | -0.021 | -0.2404 | No |
| 273 | PCCB | PCCB Entrez, Source | propionyl Coenzyme A carboxylase, beta polypeptide | 8561 | -0.021 | -0.2437 | No |
| 274 | SSRP1 | SSRP1 Entrez, Source | structure specific recognition protein 1 | 8587 | -0.022 | -0.2449 | No |
| 275 | PSMA4 | PSMA4 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 4 | 8680 | -0.023 | -0.2512 | No |
| 276 | KPNB1 | KPNB1 Entrez, Source | karyopherin (importin) beta 1 | 8707 | -0.023 | -0.2525 | No |
| 277 | ORC6L | ORC6L Entrez, Source | origin recognition complex, subunit 6 like (yeast) | 8724 | -0.023 | -0.2529 | No |
| 278 | MAGED1 | MAGED1 Entrez, Source | melanoma antigen family D, 1 | 8735 | -0.023 | -0.2529 | No |
| 279 | RPS15 | RPS15 Entrez, Source | ribosomal protein S15 | 8752 | -0.023 | -0.2533 | No |
| 280 | NCBP1 | NCBP1 Entrez, Source | nuclear cap binding protein subunit 1, 80kDa | 8762 | -0.023 | -0.2532 | No |
| 281 | RPL29 | RPL29 Entrez, Source | ribosomal protein L29 | 8850 | -0.024 | -0.2591 | No |
| 282 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 8894 | -0.025 | -0.2616 | No |
| 283 | ABAT | ABAT Entrez, Source | 4-aminobutyrate aminotransferase | 8902 | -0.025 | -0.2613 | No |
| 284 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 8905 | -0.025 | -0.2606 | No |
| 285 | PON1 | PON1 Entrez, Source | paraoxonase 1 | 8923 | -0.025 | -0.2610 | No |
| 286 | ACSBG1 | ACSBG1 Entrez, Source | acyl-CoA synthetase bubblegum family member 1 | 8970 | -0.026 | -0.2637 | No |
| 287 | STAU2 | STAU2 Entrez, Source | staufen, RNA binding protein, homolog 2 (Drosophila) | 8971 | -0.026 | -0.2628 | No |
| 288 | HSP90AB1 | HSP90AB1 Entrez, Source | heat shock protein 90kDa alpha (cytosolic), class B member 1 | 9017 | -0.026 | -0.2654 | No |
| 289 | UBE2A | UBE2A Entrez, Source | ubiquitin-conjugating enzyme E2A (RAD6 homolog) | 9034 | -0.026 | -0.2657 | No |
| 290 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 9093 | -0.027 | -0.2693 | No |
| 291 | PSAP | PSAP Entrez, Source | prosaposin (variant Gaucher disease and variant metachromatic leukodystrophy) | 9105 | -0.027 | -0.2692 | No |
| 292 | PSMA6 | PSMA6 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 6 | 9141 | -0.027 | -0.2710 | No |
| 293 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 9157 | -0.028 | -0.2712 | No |
| 294 | JAM3 | JAM3 Entrez, Source | junctional adhesion molecule 3 | 9183 | -0.028 | -0.2722 | No |
| 295 | MDH2 | MDH2 Entrez, Source | malate dehydrogenase 2, NAD (mitochondrial) | 9221 | -0.028 | -0.2740 | No |
| 296 | CALM1 | CALM1 Entrez, Source | calmodulin 1 (phosphorylase kinase, delta) | 9285 | -0.029 | -0.2779 | No |
| 297 | LETM1 | LETM1 Entrez, Source | leucine zipper-EF-hand containing transmembrane protein 1 | 9304 | -0.029 | -0.2783 | No |
| 298 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9337 | -0.029 | -0.2798 | No |
| 299 | CIB2 | CIB2 Entrez, Source | calcium and integrin binding family member 2 | 9352 | -0.030 | -0.2799 | No |
| 300 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 9381 | -0.030 | -0.2810 | No |
| 301 | ODC1 | ODC1 Entrez, Source | ornithine decarboxylase 1 | 9554 | -0.032 | -0.2933 | No |
| 302 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 9559 | -0.032 | -0.2925 | No |
| 303 | RAB7L1 | RAB7L1 Entrez, Source | RAB7, member RAS oncogene family-like 1 | 9566 | -0.032 | -0.2918 | No |
| 304 | PTPN2 | PTPN2 Entrez, Source | protein tyrosine phosphatase, non-receptor type 2 | 9568 | -0.032 | -0.2908 | No |
| 305 | VAT1 | VAT1 Entrez, Source | vesicle amine transport protein 1 homolog (T californica) | 9603 | -0.033 | -0.2923 | No |
| 306 | RAB13 | RAB13 Entrez, Source | RAB13, member RAS oncogene family | 9626 | -0.033 | -0.2928 | No |
| 307 | AURKA | AURKA Entrez, Source | aurora kinase A | 9667 | -0.034 | -0.2948 | No |
| 308 | CTTN | CTTN Entrez, Source | cortactin | 9710 | -0.034 | -0.2969 | No |
| 309 | FHIT | FHIT Entrez, Source | fragile histidine triad gene | 9754 | -0.034 | -0.2990 | No |
| 310 | WDR34 | WDR34 Entrez, Source | WD repeat domain 34 | 9757 | -0.035 | -0.2980 | No |
| 311 | CAPN2 | CAPN2 Entrez, Source | calpain 2, (m/II) large subunit | 9763 | -0.035 | -0.2971 | No |
| 312 | ATXN10 | ATXN10 Entrez, Source | ataxin 10 | 9765 | -0.035 | -0.2960 | No |
| 313 | CIB1 | CIB1 Entrez, Source | calcium and integrin binding 1 (calmyrin) | 9792 | -0.035 | -0.2968 | No |
| 314 | FTS | FTS Entrez, Source | fused toes homolog (mouse) | 9805 | -0.035 | -0.2965 | No |
| 315 | H2AFY | H2AFY Entrez, Source | H2A histone family, member Y | 9820 | -0.035 | -0.2964 | No |
| 316 | RAPGEF5 | RAPGEF5 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 5 | 9858 | -0.036 | -0.2980 | No |
| 317 | CD99 | CD99 Entrez, Source | CD99 molecule | 9867 | -0.036 | -0.2974 | No |
| 318 | NR3C2 | NR3C2 Entrez, Source | nuclear receptor subfamily 3, group C, member 2 | 9890 | -0.036 | -0.2979 | No |
| 319 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 9894 | -0.036 | -0.2968 | No |
| 320 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 9904 | -0.036 | -0.2963 | No |
| 321 | IL6ST | IL6ST Entrez, Source | interleukin 6 signal transducer (gp130, oncostatin M receptor) | 9951 | -0.037 | -0.2986 | No |
| 322 | SLC25A3 | SLC25A3 Entrez, Source | solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 3 | 9966 | -0.037 | -0.2984 | No |
| 323 | ASNS | ASNS Entrez, Source | asparagine synthetase | 9972 | -0.037 | -0.2975 | No |
| 324 | NDFIP1 | NDFIP1 Entrez, Source | Nedd4 family interacting protein 1 | 10002 | -0.037 | -0.2985 | No |
| 325 | RNASEH2A | RNASEH2A Entrez, Source | ribonuclease H2, subunit A | 10032 | -0.037 | -0.2994 | No |
| 326 | UPP1 | UPP1 Entrez, Source | uridine phosphorylase 1 | 10062 | -0.038 | -0.3004 | No |
| 327 | CDCA8 | CDCA8 Entrez, Source | cell division cycle associated 8 | 10068 | -0.038 | -0.2994 | No |
| 328 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 10085 | -0.038 | -0.2993 | No |
| 329 | SLC40A1 | SLC40A1 Entrez, Source | solute carrier family 40 (iron-regulated transporter), member 1 | 10093 | -0.038 | -0.2986 | No |
| 330 | PTP4A2 | PTP4A2 Entrez, Source | protein tyrosine phosphatase type IVA, member 2 | 10102 | -0.038 | -0.2978 | No |
| 331 | CAST | CAST Entrez, Source | calpastatin | 10118 | -0.039 | -0.2977 | No |
| 332 | CD5 | CD5 Entrez, Source | CD5 molecule | 10124 | -0.039 | -0.2967 | No |
| 333 | SKAP2 | SKAP2 Entrez, Source | src kinase associated phosphoprotein 2 | 10200 | -0.039 | -0.3012 | No |
| 334 | GIMAP7 | GIMAP7 Entrez, Source | GTPase, IMAP family member 7 | 10243 | -0.040 | -0.3031 | No |
| 335 | PSMC3 | PSMC3 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 3 | 10244 | -0.040 | -0.3017 | No |
| 336 | TMEM106B | TMEM106B Entrez, Source | transmembrane protein 106B | 10252 | -0.040 | -0.3008 | No |
| 337 | NME3 | NME3 Entrez, Source | non-metastatic cells 3, protein expressed in | 10266 | -0.040 | -0.3004 | No |
| 338 | ATP1A1 | ATP1A1 Entrez, Source | ATPase, Na+/K+ transporting, alpha 1 polypeptide | 10291 | -0.041 | -0.3009 | No |
| 339 | SLC1A4 | SLC1A4 Entrez, Source | solute carrier family 1 (glutamate/neutral amino acid transporter), member 4 | 10292 | -0.041 | -0.2995 | No |
| 340 | NDN | NDN Entrez, Source | necdin homolog (mouse) | 10375 | -0.042 | -0.3044 | No |
| 341 | MBP | MBP Entrez, Source | myelin basic protein | 10406 | -0.042 | -0.3053 | No |
| 342 | RALY | RALY Entrez, Source | RNA binding protein, autoantigenic (hnRNP-associated with lethal yellow homolog (mouse)) | 10512 | -0.044 | -0.3119 | No |
| 343 | MNS1 | MNS1 Entrez, Source | meiosis-specific nuclear structural 1 | 10516 | -0.044 | -0.3107 | No |
| 344 | RAB43 | RAB43 Entrez, Source | RAB43, member RAS oncogene family | 10522 | -0.044 | -0.3095 | No |
| 345 | IDH3A | IDH3A Entrez, Source | isocitrate dehydrogenase 3 (NAD+) alpha | 10525 | -0.044 | -0.3082 | No |
| 346 | SFPQ | SFPQ Entrez, Source | splicing factor proline/glutamine-rich (polypyrimidine tract binding protein associated) | 10541 | -0.044 | -0.3078 | No |
| 347 | TLOC1 | TLOC1 Entrez, Source | translocation protein 1 | 10559 | -0.044 | -0.3076 | No |
| 348 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 10560 | -0.044 | -0.3060 | No |
| 349 | CASP6 | CASP6 Entrez, Source | caspase 6, apoptosis-related cysteine peptidase | 10649 | -0.045 | -0.3113 | No |
| 350 | HDGF | HDGF Entrez, Source | hepatoma-derived growth factor (high-mobility group protein 1-like) | 10656 | -0.046 | -0.3102 | No |
| 351 | CD96 | CD96 Entrez, Source | CD96 molecule | 10674 | -0.046 | -0.3099 | No |
| 352 | EMP3 | EMP3 Entrez, Source | epithelial membrane protein 3 | 10678 | -0.046 | -0.3086 | No |
| 353 | NME1 | NME1 Entrez, Source | non-metastatic cells 1, protein (NM23A) expressed in | 10783 | -0.048 | -0.3150 | No |
| 354 | CD52 | CD52 Entrez, Source | CD52 molecule | 10867 | -0.049 | -0.3197 | No |
| 355 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 10869 | -0.049 | -0.3181 | No |
| 356 | CDR2 | CDR2 Entrez, Source | cerebellar degeneration-related protein 2, 62kDa | 10906 | -0.050 | -0.3192 | No |
| 357 | ARL3 | ARL3 Entrez, Source | ADP-ribosylation factor-like 3 | 10937 | -0.050 | -0.3198 | No |
| 358 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 10939 | -0.050 | -0.3181 | No |
| 359 | PLEKHA5 | PLEKHA5 Entrez, Source | pleckstrin homology domain containing, family A member 5 | 10971 | -0.051 | -0.3188 | No |
| 360 | SPRED2 | SPRED2 Entrez, Source | sprouty-related, EVH1 domain containing 2 | 10977 | -0.051 | -0.3174 | No |
| 361 | ZWINT | ZWINT Entrez, Source | ZW10 interactor | 10978 | -0.051 | -0.3156 | No |
| 362 | SHMT1 | SHMT1 Entrez, Source | serine hydroxymethyltransferase 1 (soluble) | 11060 | -0.052 | -0.3201 | No |
| 363 | PRMT2 | PRMT2 Entrez, Source | protein arginine methyltransferase 2 | 11068 | -0.052 | -0.3188 | No |
| 364 | BARD1 | BARD1 Entrez, Source | BRCA1 associated RING domain 1 | 11085 | -0.053 | -0.3183 | No |
| 365 | NME7 | NME7 Entrez, Source | non-metastatic cells 7, protein expressed in (nucleoside-diphosphate kinase) | 11095 | -0.053 | -0.3171 | No |
| 366 | RAB3D | RAB3D Entrez, Source | RAB3D, member RAS oncogene family | 11143 | -0.054 | -0.3189 | No |
| 367 | GLUL | GLUL Entrez, Source | glutamate-ammonia ligase (glutamine synthetase) | 11146 | -0.054 | -0.3172 | No |
| 368 | CREBL2 | CREBL2 Entrez, Source | cAMP responsive element binding protein-like 2 | 11190 | -0.055 | -0.3186 | No |
| 369 | TOR1B | TOR1B Entrez, Source | torsin family 1, member B (torsin B) | 11237 | -0.056 | -0.3203 | No |
| 370 | TAX1BP3 | TAX1BP3 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 3 | 11278 | -0.056 | -0.3214 | Yes |
| 371 | PDCD8 | PDCD8 Entrez, Source | programmed cell death 8 (apoptosis-inducing factor) | 11289 | -0.057 | -0.3202 | Yes |
| 372 | GFI1 | GFI1 Entrez, Source | growth factor independent 1 | 11310 | -0.057 | -0.3198 | Yes |
| 373 | YWHAQ | YWHAQ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta polypeptide | 11312 | -0.057 | -0.3179 | Yes |
| 374 | MTF2 | MTF2 Entrez, Source | metal response element binding transcription factor 2 | 11313 | -0.057 | -0.3159 | Yes |
| 375 | SEPHS1 | SEPHS1 Entrez, Source | selenophosphate synthetase 1 | 11314 | -0.057 | -0.3139 | Yes |
| 376 | FXYD5 | FXYD5 Entrez, Source | FXYD domain containing ion transport regulator 5 | 11354 | -0.058 | -0.3150 | Yes |
| 377 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 11369 | -0.058 | -0.3140 | Yes |
| 378 | PAICS | PAICS Entrez, Source | phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoimidazole succinocarboxamide synthetase | 11374 | -0.058 | -0.3123 | Yes |
| 379 | PSMD1 | PSMD1 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 1 | 11377 | -0.058 | -0.3104 | Yes |
| 380 | TUBB | TUBB Entrez, Source | tubulin, beta | 11384 | -0.058 | -0.3089 | Yes |
| 381 | RIMS3 | RIMS3 Entrez, Source | regulating synaptic membrane exocytosis 3 | 11395 | -0.059 | -0.3076 | Yes |
| 382 | MAP3K1 | MAP3K1 Entrez, Source | mitogen-activated protein kinase kinase kinase 1 | 11443 | -0.059 | -0.3092 | Yes |
| 383 | PPP1R14B | PPP1R14B Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 14B | 11464 | -0.060 | -0.3087 | Yes |
| 384 | CHN2 | CHN2 Entrez, Source | chimerin (chimaerin) 2 | 11487 | -0.060 | -0.3083 | Yes |
| 385 | MARCKS | MARCKS Entrez, Source | myristoylated alanine-rich protein kinase C substrate | 11501 | -0.060 | -0.3072 | Yes |
| 386 | MPEG1 | MPEG1 Entrez, Source | - | 11505 | -0.060 | -0.3053 | Yes |
| 387 | TSC22D1 | TSC22D1 Entrez, Source | TSC22 domain family, member 1 | 11547 | -0.061 | -0.3064 | Yes |
| 388 | PASK | PASK Entrez, Source | PAS domain containing serine/threonine kinase | 11561 | -0.062 | -0.3053 | Yes |
| 389 | UBE2E2 | UBE2E2 Entrez, Source | ubiquitin-conjugating enzyme E2E 2 (UBC4/5 homolog, yeast) | 11565 | -0.062 | -0.3034 | Yes |
| 390 | PTK2 | PTK2 Entrez, Source | PTK2 protein tyrosine kinase 2 | 11607 | -0.063 | -0.3044 | Yes |
| 391 | ACACB | ACACB Entrez, Source | acetyl-Coenzyme A carboxylase beta | 11612 | -0.063 | -0.3025 | Yes |
| 392 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 11618 | -0.063 | -0.3007 | Yes |
| 393 | DPP3 | DPP3 Entrez, Source | dipeptidyl-peptidase 3 | 11670 | -0.064 | -0.3024 | Yes |
| 394 | ZNF14 | ZNF14 Entrez, Source | zinc finger protein 14 | 11701 | -0.065 | -0.3025 | Yes |
| 395 | PGGT1B | PGGT1B Entrez, Source | protein geranylgeranyltransferase type I, beta subunit | 11723 | -0.065 | -0.3019 | Yes |
| 396 | GUSB | GUSB Entrez, Source | glucuronidase, beta | 11743 | -0.066 | -0.3011 | Yes |
| 397 | SFRS9 | SFRS9 Entrez, Source | splicing factor, arginine/serine-rich 9 | 11816 | -0.068 | -0.3043 | Yes |
| 398 | SPBC25 | SPBC25 Entrez, Source | spindle pole body component 25 homolog (S. cerevisiae) | 11828 | -0.068 | -0.3028 | Yes |
| 399 | OSBPL5 | OSBPL5 Entrez, Source | oxysterol binding protein-like 5 | 11832 | -0.068 | -0.3007 | Yes |
| 400 | B4GALT6 | B4GALT6 Entrez, Source | UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 6 | 11834 | -0.068 | -0.2984 | Yes |
| 401 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 11849 | -0.069 | -0.2971 | Yes |
| 402 | SH3BP5 | SH3BP5 Entrez, Source | SH3-domain binding protein 5 (BTK-associated) | 11864 | -0.069 | -0.2958 | Yes |
| 403 | INPP1 | INPP1 Entrez, Source | inositol polyphosphate-1-phosphatase | 11919 | -0.070 | -0.2975 | Yes |
| 404 | CBX7 | CBX7 Entrez, Source | chromobox homolog 7 | 11922 | -0.070 | -0.2952 | Yes |
| 405 | TM7SF3 | TM7SF3 Entrez, Source | transmembrane 7 superfamily member 3 | 11923 | -0.070 | -0.2928 | Yes |
| 406 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 11928 | -0.071 | -0.2906 | Yes |
| 407 | LIMK2 | LIMK2 Entrez, Source | LIM domain kinase 2 | 11957 | -0.071 | -0.2903 | Yes |
| 408 | FXYD2 | FXYD2 Entrez, Source | FXYD domain containing ion transport regulator 2 | 11964 | -0.072 | -0.2883 | Yes |
| 409 | PTPRK | PTPRK Entrez, Source | protein tyrosine phosphatase, receptor type, K | 11989 | -0.073 | -0.2877 | Yes |
| 410 | CTSC | CTSC Entrez, Source | cathepsin C | 11993 | -0.073 | -0.2854 | Yes |
| 411 | SIGIRR | SIGIRR Entrez, Source | single immunoglobulin and toll-interleukin 1 receptor (TIR) domain | 12066 | -0.075 | -0.2884 | Yes |
| 412 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 12078 | -0.075 | -0.2866 | Yes |
| 413 | ALDH7A1 | ALDH7A1 Entrez, Source | aldehyde dehydrogenase 7 family, member A1 | 12088 | -0.076 | -0.2847 | Yes |
| 414 | VRK1 | VRK1 Entrez, Source | vaccinia related kinase 1 | 12091 | -0.076 | -0.2822 | Yes |
| 415 | VIM | VIM Entrez, Source | vimentin | 12125 | -0.077 | -0.2821 | Yes |
| 416 | GGH | GGH Entrez, Source | gamma-glutamyl hydrolase (conjugase, folylpolygammaglutamyl hydrolase) | 12131 | -0.077 | -0.2798 | Yes |
| 417 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 12137 | -0.077 | -0.2775 | Yes |
| 418 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 12169 | -0.078 | -0.2772 | Yes |
| 419 | RUFY3 | RUFY3 Entrez, Source | RUN and FYVE domain containing 3 | 12269 | -0.082 | -0.2821 | Yes |
| 420 | PXN | PXN Entrez, Source | paxillin | 12322 | -0.084 | -0.2832 | Yes |
| 421 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 12364 | -0.086 | -0.2834 | Yes |
| 422 | MYEF2 | MYEF2 Entrez, Source | myelin expression factor 2 | 12369 | -0.086 | -0.2807 | Yes |
| 423 | CLIC4 | CLIC4 Entrez, Source | chloride intracellular channel 4 | 12392 | -0.087 | -0.2794 | Yes |
| 424 | TIMP1 | TIMP1 Entrez, Source | TIMP metallopeptidase inhibitor 1 | 12415 | -0.087 | -0.2781 | Yes |
| 425 | PHGDH | PHGDH Entrez, Source | phosphoglycerate dehydrogenase | 12426 | -0.088 | -0.2758 | Yes |
| 426 | PRNP | PRNP Entrez, Source | prion protein (p27-30) (Creutzfeldt-Jakob disease, Gerstmann-Strausler-Scheinker syndrome, fatal familial insomnia) | 12448 | -0.089 | -0.2744 | Yes |
| 427 | AHI1 | AHI1 Entrez, Source | Abelson helper integration site 1 | 12457 | -0.089 | -0.2719 | Yes |
| 428 | GNA15 | GNA15 Entrez, Source | guanine nucleotide binding protein (G protein), alpha 15 (Gq class) | 12502 | -0.092 | -0.2721 | Yes |
| 429 | PSMC3IP | PSMC3IP Entrez, Source | PSMC3 interacting protein | 12571 | -0.095 | -0.2741 | Yes |
| 430 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 12577 | -0.095 | -0.2712 | Yes |
| 431 | CLDN1 | CLDN1 Entrez, Source | claudin 1 | 12658 | -0.100 | -0.2739 | Yes |
| 432 | PLSCR3 | PLSCR3 Entrez, Source | phospholipid scramblase 3 | 12667 | -0.100 | -0.2711 | Yes |
| 433 | AMIGO2 | AMIGO2 Entrez, Source | adhesion molecule with Ig-like domain 2 | 12673 | -0.101 | -0.2680 | Yes |
| 434 | ACPL2 | ACPL2 Entrez, Source | acid phosphatase-like 2 | 12680 | -0.101 | -0.2649 | Yes |
| 435 | PPP4R1 | PPP4R1 Entrez, Source | protein phosphatase 4, regulatory subunit 1 | 12714 | -0.104 | -0.2638 | Yes |
| 436 | PDCD4 | PDCD4 Entrez, Source | programmed cell death 4 (neoplastic transformation inhibitor) | 12722 | -0.105 | -0.2608 | Yes |
| 437 | HSPA4 | HSPA4 Entrez, Source | heat shock 70kDa protein 4 | 12729 | -0.105 | -0.2576 | Yes |
| 438 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 12735 | -0.105 | -0.2543 | Yes |
| 439 | CD69 | CD69 Entrez, Source | CD69 molecule | 12743 | -0.106 | -0.2512 | Yes |
| 440 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 12758 | -0.107 | -0.2485 | Yes |
| 441 | PAK1 | PAK1 Entrez, Source | p21/Cdc42/Rac1-activated kinase 1 (STE20 homolog, yeast) | 12764 | -0.108 | -0.2451 | Yes |
| 442 | PDE3B | PDE3B Entrez, Source | phosphodiesterase 3B, cGMP-inhibited | 12777 | -0.109 | -0.2423 | Yes |
| 443 | TAGLN2 | TAGLN2 Entrez, Source | transgelin 2 | 12779 | -0.109 | -0.2386 | Yes |
| 444 | ASAH1 | ASAH1 Entrez, Source | N-acylsphingosine amidohydrolase (acid ceramidase) 1 | 12784 | -0.110 | -0.2351 | Yes |
| 445 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 12790 | -0.110 | -0.2316 | Yes |
| 446 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 12799 | -0.110 | -0.2284 | Yes |
| 447 | TMPO | TMPO Entrez, Source | thymopoietin | 12812 | -0.111 | -0.2255 | Yes |
| 448 | IRAK2 | IRAK2 Entrez, Source | interleukin-1 receptor-associated kinase 2 | 12849 | -0.115 | -0.2243 | Yes |
| 449 | P2RY1 | P2RY1 Entrez, Source | purinergic receptor P2Y, G-protein coupled, 1 | 12861 | -0.116 | -0.2211 | Yes |
| 450 | PAWR | PAWR Entrez, Source | PRKC, apoptosis, WT1, regulator | 12871 | -0.118 | -0.2177 | Yes |
| 451 | PLDN | PLDN Entrez, Source | pallidin homolog (mouse) | 12874 | -0.118 | -0.2137 | Yes |
| 452 | TMSB10 | TMSB10 Entrez, Source | thymosin, beta 10 | 12891 | -0.120 | -0.2108 | Yes |
| 453 | RASA2 | RASA2 Entrez, Source | RAS p21 protein activator 2 | 12895 | -0.120 | -0.2069 | Yes |
| 454 | PSCD1 | PSCD1 Entrez, Source | pleckstrin homology, Sec7 and coiled-coil domains 1(cytohesin 1) | 12896 | -0.121 | -0.2027 | Yes |
| 455 | RFC3 | RFC3 Entrez, Source | replication factor C (activator 1) 3, 38kDa | 12915 | -0.123 | -0.1998 | Yes |
| 456 | DPP4 | DPP4 Entrez, Source | dipeptidyl-peptidase 4 (CD26, adenosine deaminase complexing protein 2) | 12933 | -0.125 | -0.1968 | Yes |
| 457 | DPYSL2 | DPYSL2 Entrez, Source | dihydropyrimidinase-like 2 | 12948 | -0.126 | -0.1935 | Yes |
| 458 | FAIM | FAIM Entrez, Source | Fas apoptotic inhibitory molecule | 12956 | -0.127 | -0.1896 | Yes |
| 459 | TPM4 | TPM4 Entrez, Source | tropomyosin 4 | 12972 | -0.128 | -0.1863 | Yes |
| 460 | PSIP1 | PSIP1 Entrez, Source | PC4 and SFRS1 interacting protein 1 | 13029 | -0.136 | -0.1860 | Yes |
| 461 | LCP1 | LCP1 Entrez, Source | lymphocyte cytosolic protein 1 (L-plastin) | 13041 | -0.138 | -0.1820 | Yes |
| 462 | LZTFL1 | LZTFL1 Entrez, Source | leucine zipper transcription factor-like 1 | 13050 | -0.140 | -0.1778 | Yes |
| 463 | BIRC3 | BIRC3 Entrez, Source | baculoviral IAP repeat-containing 3 | 13071 | -0.143 | -0.1743 | Yes |
| 464 | TRIP13 | TRIP13 Entrez, Source | thyroid hormone receptor interactor 13 | 13078 | -0.144 | -0.1698 | Yes |
| 465 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 13084 | -0.146 | -0.1651 | Yes |
| 466 | RRAS2 | RRAS2 Entrez, Source | related RAS viral (r-ras) oncogene homolog 2 | 13088 | -0.147 | -0.1602 | Yes |
| 467 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 13097 | -0.150 | -0.1556 | Yes |
| 468 | MARCKSL1 | MARCKSL1 Entrez, Source | MARCKS-like 1 | 13105 | -0.150 | -0.1509 | Yes |
| 469 | PRPS2 | PRPS2 Entrez, Source | phosphoribosyl pyrophosphate synthetase 2 | 13107 | -0.151 | -0.1457 | Yes |
| 470 | PLSCR1 | PLSCR1 Entrez, Source | phospholipid scramblase 1 | 13117 | -0.153 | -0.1411 | Yes |
| 471 | CD44 | CD44 Entrez, Source | CD44 molecule (Indian blood group) | 13165 | -0.168 | -0.1389 | Yes |
| 472 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 13167 | -0.169 | -0.1331 | Yes |
| 473 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 13174 | -0.172 | -0.1276 | Yes |
| 474 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 13185 | -0.179 | -0.1222 | Yes |
| 475 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 13192 | -0.182 | -0.1163 | Yes |
| 476 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13194 | -0.185 | -0.1100 | Yes |
| 477 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13196 | -0.186 | -0.1036 | Yes |
| 478 | SERINC5 | SERINC5 Entrez, Source | serine incorporator 5 | 13200 | -0.190 | -0.0972 | Yes |
| 479 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 13232 | -0.213 | -0.0922 | Yes |
| 480 | SPRY2 | SPRY2 Entrez, Source | sprouty homolog 2 (Drosophila) | 13247 | -0.219 | -0.0857 | Yes |
| 481 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 13248 | -0.219 | -0.0781 | Yes |
| 482 | LTB | LTB Entrez, Source | lymphotoxin beta (TNF superfamily, member 3) | 13250 | -0.220 | -0.0705 | Yes |
| 483 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 13253 | -0.222 | -0.0629 | Yes |
| 484 | PDE4B | PDE4B Entrez, Source | phosphodiesterase 4B, cAMP-specific (phosphodiesterase E4 dunce homolog, Drosophila) | 13271 | -0.241 | -0.0559 | Yes |
| 485 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 13295 | -0.275 | -0.0481 | Yes |
| 486 | S100A4 | S100A4 Entrez, Source | S100 calcium binding protein A4 | 13306 | -0.310 | -0.0381 | Yes |
| 487 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 13314 | -0.341 | -0.0268 | Yes |
| 488 | GBP2 | GBP2 Entrez, Source | guanylate binding protein 2, interferon-inducible | 13323 | -0.371 | -0.0145 | Yes |
| 489 | S100A6 | S100A6 Entrez, Source | S100 calcium binding protein A6 | 13330 | -0.458 | 0.0009 | Yes |