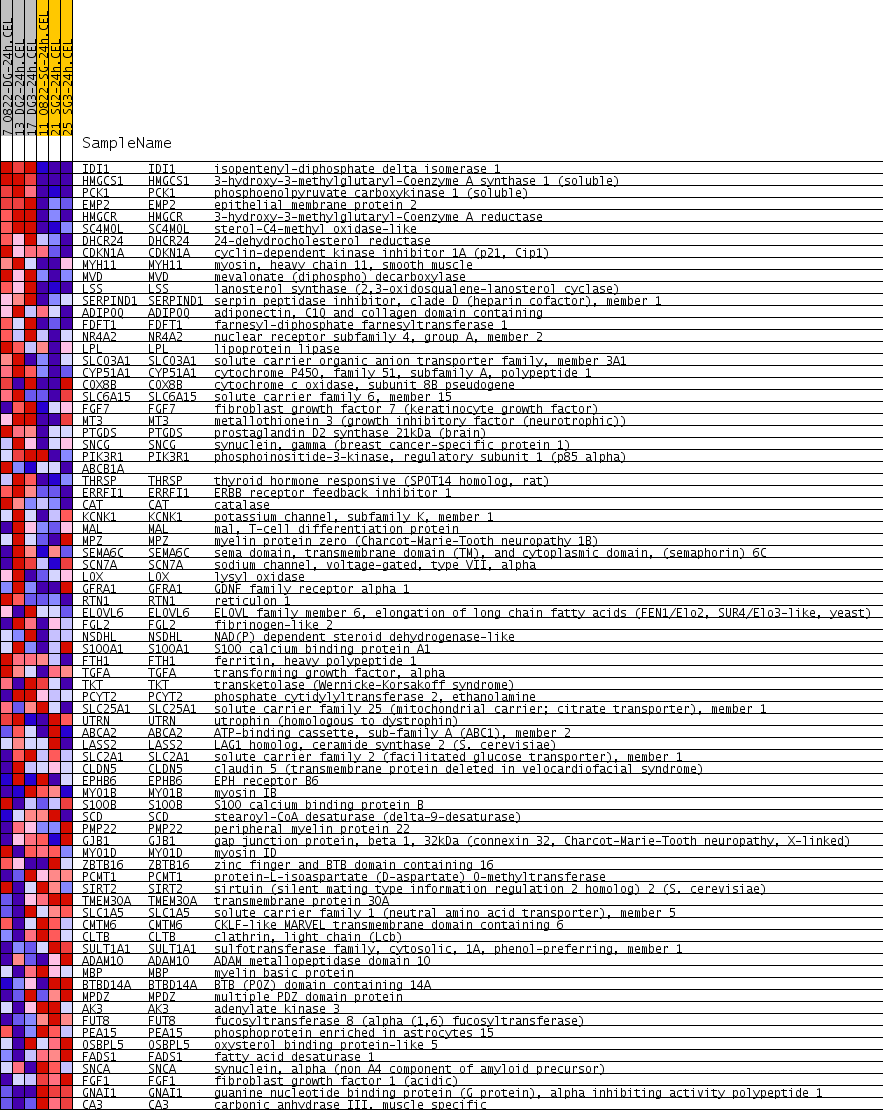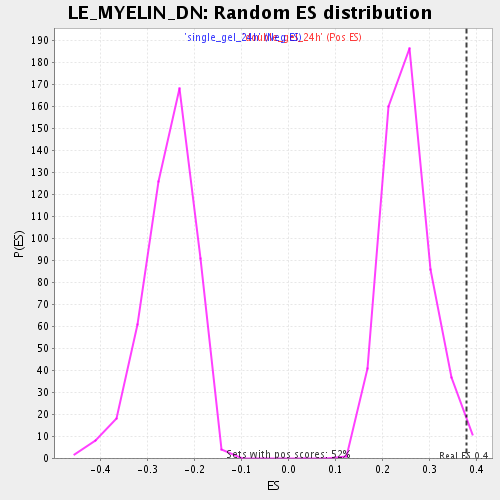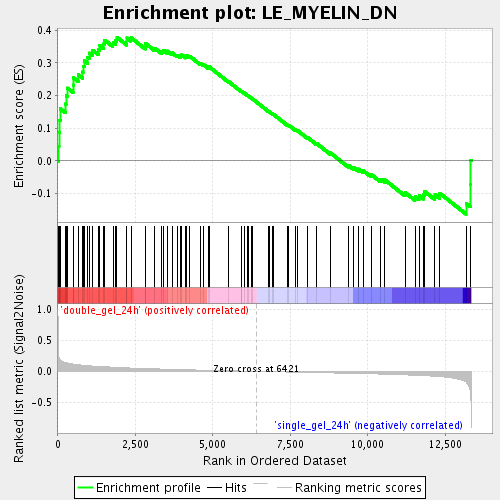
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | LE_MYELIN_DN |
| Enrichment Score (ES) | 0.37812412 |
| Normalized Enrichment Score (NES) | 1.5046377 |
| Nominal p-value | 0.009578544 |
| FDR q-value | 0.27054846 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IDI1 | IDI1 Entrez, Source | isopentenyl-diphosphate delta isomerase 1 | 29 | 0.238 | 0.0439 | Yes |
| 2 | HMGCS1 | HMGCS1 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 1 (soluble) | 36 | 0.229 | 0.0877 | Yes |
| 3 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 59 | 0.201 | 0.1250 | Yes |
| 4 | EMP2 | EMP2 Entrez, Source | epithelial membrane protein 2 | 79 | 0.187 | 0.1597 | Yes |
| 5 | HMGCR | HMGCR Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A reductase | 250 | 0.141 | 0.1742 | Yes |
| 6 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 266 | 0.140 | 0.2001 | Yes |
| 7 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 308 | 0.134 | 0.2230 | Yes |
| 8 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 491 | 0.118 | 0.2322 | Yes |
| 9 | MYH11 | MYH11 Entrez, Source | myosin, heavy chain 11, smooth muscle | 500 | 0.118 | 0.2544 | Yes |
| 10 | MVD | MVD Entrez, Source | mevalonate (diphospho) decarboxylase | 658 | 0.107 | 0.2632 | Yes |
| 11 | LSS | LSS Entrez, Source | lanosterol synthase (2,3-oxidosqualene-lanosterol cyclase) | 797 | 0.100 | 0.2721 | Yes |
| 12 | SERPIND1 | SERPIND1 Entrez, Source | serpin peptidase inhibitor, clade D (heparin cofactor), member 1 | 812 | 0.099 | 0.2902 | Yes |
| 13 | ADIPOQ | ADIPOQ Entrez, Source | adiponectin, C1Q and collagen domain containing | 852 | 0.097 | 0.3061 | Yes |
| 14 | FDFT1 | FDFT1 Entrez, Source | farnesyl-diphosphate farnesyltransferase 1 | 952 | 0.094 | 0.3167 | Yes |
| 15 | NR4A2 | NR4A2 Entrez, Source | nuclear receptor subfamily 4, group A, member 2 | 1011 | 0.091 | 0.3300 | Yes |
| 16 | LPL | LPL Entrez, Source | lipoprotein lipase | 1116 | 0.087 | 0.3389 | Yes |
| 17 | SLCO3A1 | SLCO3A1 Entrez, Source | solute carrier organic anion transporter family, member 3A1 | 1303 | 0.080 | 0.3404 | Yes |
| 18 | CYP51A1 | CYP51A1 Entrez, Source | cytochrome P450, family 51, subfamily A, polypeptide 1 | 1345 | 0.079 | 0.3526 | Yes |
| 19 | COX8B | COX8B Entrez, Source | cytochrome c oxidase, subunit 8B pseudogene | 1472 | 0.075 | 0.3577 | Yes |
| 20 | SLC6A15 | SLC6A15 Entrez, Source | solute carrier family 6, member 15 | 1509 | 0.074 | 0.3694 | Yes |
| 21 | FGF7 | FGF7 Entrez, Source | fibroblast growth factor 7 (keratinocyte growth factor) | 1778 | 0.068 | 0.3623 | Yes |
| 22 | MT3 | MT3 Entrez, Source | metallothionein 3 (growth inhibitory factor (neurotrophic)) | 1858 | 0.066 | 0.3692 | Yes |
| 23 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 1906 | 0.065 | 0.3781 | Yes |
| 24 | SNCG | SNCG Entrez, Source | synuclein, gamma (breast cancer-specific protein 1) | 2226 | 0.057 | 0.3651 | No |
| 25 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 2228 | 0.057 | 0.3760 | No |
| 26 | ABCB1A | 2358 | 0.054 | 0.3768 | No | ||
| 27 | THRSP | THRSP Entrez, Source | thyroid hormone responsive (SPOT14 homolog, rat) | 2819 | 0.046 | 0.3510 | No |
| 28 | ERRFI1 | ERRFI1 Entrez, Source | ERBB receptor feedback inhibitor 1 | 2825 | 0.046 | 0.3595 | No |
| 29 | CAT | CAT Entrez, Source | catalase | 3129 | 0.041 | 0.3446 | No |
| 30 | KCNK1 | KCNK1 Entrez, Source | potassium channel, subfamily K, member 1 | 3345 | 0.037 | 0.3357 | No |
| 31 | MAL | MAL Entrez, Source | mal, T-cell differentiation protein | 3407 | 0.037 | 0.3381 | No |
| 32 | MPZ | MPZ Entrez, Source | myelin protein zero (Charcot-Marie-Tooth neuropathy 1B) | 3523 | 0.035 | 0.3362 | No |
| 33 | SEMA6C | SEMA6C Entrez, Source | sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6C | 3686 | 0.033 | 0.3303 | No |
| 34 | SCN7A | SCN7A Entrez, Source | sodium channel, voltage-gated, type VII, alpha | 3851 | 0.030 | 0.3238 | No |
| 35 | LOX | LOX Entrez, Source | lysyl oxidase | 3959 | 0.029 | 0.3214 | No |
| 36 | GFRA1 | GFRA1 Entrez, Source | GDNF family receptor alpha 1 | 3997 | 0.028 | 0.3241 | No |
| 37 | RTN1 | RTN1 Entrez, Source | reticulon 1 | 4123 | 0.027 | 0.3199 | No |
| 38 | ELOVL6 | ELOVL6 Entrez, Source | ELOVL family member 6, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 4158 | 0.026 | 0.3224 | No |
| 39 | FGL2 | FGL2 Entrez, Source | fibrinogen-like 2 | 4253 | 0.025 | 0.3202 | No |
| 40 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 4607 | 0.021 | 0.2976 | No |
| 41 | S100A1 | S100A1 Entrez, Source | S100 calcium binding protein A1 | 4694 | 0.020 | 0.2949 | No |
| 42 | FTH1 | FTH1 Entrez, Source | ferritin, heavy polypeptide 1 | 4858 | 0.018 | 0.2860 | No |
| 43 | TGFA | TGFA Entrez, Source | transforming growth factor, alpha | 4880 | 0.018 | 0.2879 | No |
| 44 | TKT | TKT Entrez, Source | transketolase (Wernicke-Korsakoff syndrome) | 5509 | 0.010 | 0.2425 | No |
| 45 | PCYT2 | PCYT2 Entrez, Source | phosphate cytidylyltransferase 2, ethanolamine | 5921 | 0.005 | 0.2125 | No |
| 46 | SLC25A1 | SLC25A1 Entrez, Source | solute carrier family 25 (mitochondrial carrier; citrate transporter), member 1 | 5938 | 0.005 | 0.2123 | No |
| 47 | UTRN | UTRN Entrez, Source | utrophin (homologous to dystrophin) | 6005 | 0.005 | 0.2082 | No |
| 48 | ABCA2 | ABCA2 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 2 | 6113 | 0.004 | 0.2008 | No |
| 49 | LASS2 | LASS2 Entrez, Source | LAG1 homolog, ceramide synthase 2 (S. cerevisiae) | 6148 | 0.003 | 0.1989 | No |
| 50 | SLC2A1 | SLC2A1 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 1 | 6242 | 0.002 | 0.1922 | No |
| 51 | CLDN5 | CLDN5 Entrez, Source | claudin 5 (transmembrane protein deleted in velocardiofacial syndrome) | 6273 | 0.002 | 0.1903 | No |
| 52 | EPHB6 | EPHB6 Entrez, Source | EPH receptor B6 | 6786 | -0.004 | 0.1524 | No |
| 53 | MYO1B | MYO1B Entrez, Source | myosin IB | 6819 | -0.004 | 0.1507 | No |
| 54 | S100B | S100B Entrez, Source | S100 calcium binding protein B | 6931 | -0.005 | 0.1433 | No |
| 55 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 6967 | -0.005 | 0.1417 | No |
| 56 | PMP22 | PMP22 Entrez, Source | peripheral myelin protein 22 | 7414 | -0.010 | 0.1099 | No |
| 57 | GJB1 | GJB1 Entrez, Source | gap junction protein, beta 1, 32kDa (connexin 32, Charcot-Marie-Tooth neuropathy, X-linked) | 7457 | -0.010 | 0.1087 | No |
| 58 | MYO1D | MYO1D Entrez, Source | myosin ID | 7665 | -0.012 | 0.0955 | No |
| 59 | ZBTB16 | ZBTB16 Entrez, Source | zinc finger and BTB domain containing 16 | 7722 | -0.013 | 0.0937 | No |
| 60 | PCMT1 | PCMT1 Entrez, Source | protein-L-isoaspartate (D-aspartate) O-methyltransferase | 8056 | -0.016 | 0.0718 | No |
| 61 | SIRT2 | SIRT2 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 2 (S. cerevisiae) | 8333 | -0.019 | 0.0546 | No |
| 62 | TMEM30A | TMEM30A Entrez, Source | transmembrane protein 30A | 8787 | -0.024 | 0.0251 | No |
| 63 | SLC1A5 | SLC1A5 Entrez, Source | solute carrier family 1 (neutral amino acid transporter), member 5 | 9379 | -0.030 | -0.0137 | No |
| 64 | CMTM6 | CMTM6 Entrez, Source | CKLF-like MARVEL transmembrane domain containing 6 | 9535 | -0.032 | -0.0192 | No |
| 65 | CLTB | CLTB Entrez, Source | clathrin, light chain (Lcb) | 9694 | -0.034 | -0.0246 | No |
| 66 | SULT1A1 | SULT1A1 Entrez, Source | sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 | 9854 | -0.036 | -0.0296 | No |
| 67 | ADAM10 | ADAM10 Entrez, Source | ADAM metallopeptidase domain 10 | 10107 | -0.038 | -0.0412 | No |
| 68 | MBP | MBP Entrez, Source | myelin basic protein | 10406 | -0.042 | -0.0556 | No |
| 69 | BTBD14A | BTBD14A Entrez, Source | BTB (POZ) domain containing 14A | 10531 | -0.044 | -0.0564 | No |
| 70 | MPDZ | MPDZ Entrez, Source | multiple PDZ domain protein | 11206 | -0.055 | -0.0966 | No |
| 71 | AK3 | AK3 Entrez, Source | adenylate kinase 3 | 11525 | -0.061 | -0.1088 | No |
| 72 | FUT8 | FUT8 Entrez, Source | fucosyltransferase 8 (alpha (1,6) fucosyltransferase) | 11656 | -0.064 | -0.1062 | No |
| 73 | PEA15 | PEA15 Entrez, Source | phosphoprotein enriched in astrocytes 15 | 11797 | -0.067 | -0.1038 | No |
| 74 | OSBPL5 | OSBPL5 Entrez, Source | oxysterol binding protein-like 5 | 11832 | -0.068 | -0.0931 | No |
| 75 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 12169 | -0.078 | -0.1033 | No |
| 76 | SNCA | SNCA Entrez, Source | synuclein, alpha (non A4 component of amyloid precursor) | 12324 | -0.084 | -0.0987 | No |
| 77 | FGF1 | FGF1 Entrez, Source | fibroblast growth factor 1 (acidic) | 13172 | -0.171 | -0.1295 | No |
| 78 | GNAI1 | GNAI1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | 13318 | -0.354 | -0.0719 | No |
| 79 | CA3 | CA3 Entrez, Source | carbonic anhydrase III, muscle specific | 13324 | -0.380 | 0.0014 | No |