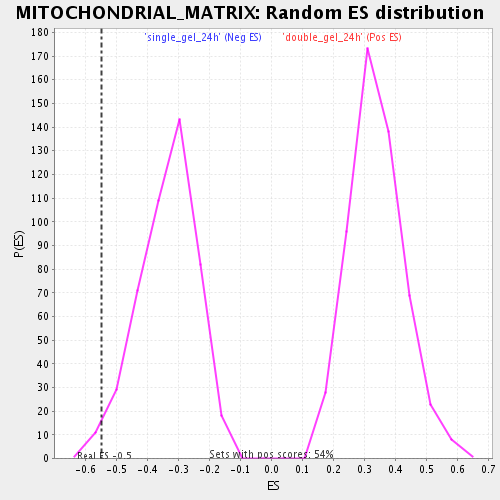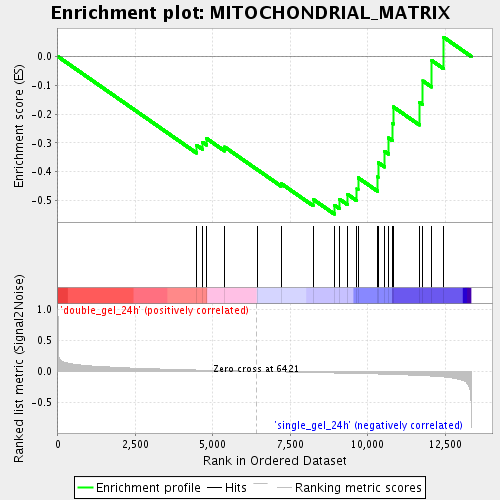
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | MITOCHONDRIAL_MATRIX |
| Enrichment Score (ES) | -0.5479772 |
| Normalized Enrichment Score (NES) | -1.6362222 |
| Nominal p-value | 0.01724138 |
| FDR q-value | 0.20460846 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NR3C1 | NR3C1 Entrez, Source | nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) | 4473 | 0.022 | -0.3083 | No |
| 2 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4664 | 0.020 | -0.2980 | No |
| 3 | ETFB | ETFB Entrez, Source | electron-transfer-flavoprotein, beta polypeptide | 4799 | 0.019 | -0.2851 | No |
| 4 | BCKDK | BCKDK Entrez, Source | branched chain ketoacid dehydrogenase kinase | 5381 | 0.011 | -0.3145 | No |
| 5 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 6435 | -0.000 | -0.3934 | No |
| 6 | ETFA | ETFA Entrez, Source | electron-transfer-flavoprotein, alpha polypeptide (glutaric aciduria II) | 7202 | -0.008 | -0.4415 | No |
| 7 | ATP5F1 | ATP5F1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit B1 | 8238 | -0.018 | -0.4968 | No |
| 8 | GRPEL1 | GRPEL1 Entrez, Source | GrpE-like 1, mitochondrial (E. coli) | 8921 | -0.025 | -0.5168 | Yes |
| 9 | CS | CS Entrez, Source | citrate synthase | 9089 | -0.027 | -0.4962 | Yes |
| 10 | DNAJA3 | DNAJA3 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 3 | 9342 | -0.029 | -0.4787 | Yes |
| 11 | MRPL40 | MRPL40 Entrez, Source | mitochondrial ribosomal protein L40 | 9651 | -0.033 | -0.4606 | Yes |
| 12 | MRPL41 | MRPL41 Entrez, Source | mitochondrial ribosomal protein L41 | 9689 | -0.034 | -0.4215 | Yes |
| 13 | MRPL23 | MRPL23 Entrez, Source | mitochondrial ribosomal protein L23 | 10323 | -0.041 | -0.4186 | Yes |
| 14 | DBT | DBT Entrez, Source | dihydrolipoamide branched chain transacylase E2 | 10335 | -0.041 | -0.3687 | Yes |
| 15 | MRPL12 | MRPL12 Entrez, Source | mitochondrial ribosomal protein L12 | 10545 | -0.044 | -0.3300 | Yes |
| 16 | NFS1 | NFS1 Entrez, Source | NFS1 nitrogen fixation 1 (S. cerevisiae) | 10672 | -0.046 | -0.2829 | Yes |
| 17 | TFB2M | TFB2M Entrez, Source | transcription factor B2, mitochondrial | 10804 | -0.048 | -0.2335 | Yes |
| 18 | MRPS15 | MRPS15 Entrez, Source | mitochondrial ribosomal protein S15 | 10823 | -0.048 | -0.1750 | Yes |
| 19 | SUPV3L1 | SUPV3L1 Entrez, Source | suppressor of var1, 3-like 1 (S. cerevisiae) | 11668 | -0.064 | -0.1591 | Yes |
| 20 | TIMM44 | TIMM44 Entrez, Source | translocase of inner mitochondrial membrane 44 homolog (yeast) | 11781 | -0.067 | -0.0849 | Yes |
| 21 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 12065 | -0.075 | -0.0138 | Yes |
| 22 | MRPS18A | MRPS18A Entrez, Source | mitochondrial ribosomal protein S18A | 12449 | -0.089 | 0.0670 | Yes |