Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | NETPATH_TNF_ALPHA_PATHWAY_DOWN |
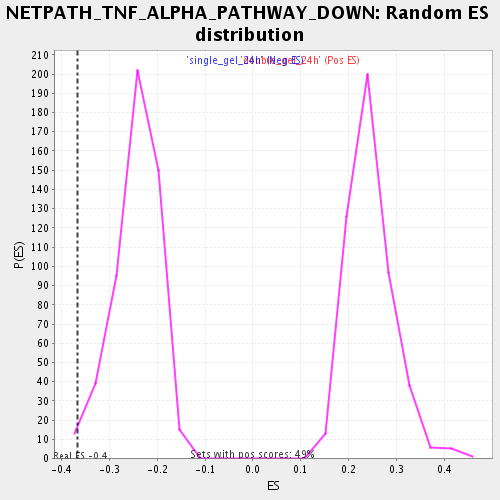
| Enrichment Score (ES) | -0.36708924 |
| Normalized Enrichment Score (NES) | -1.5023935 |
| Nominal p-value | 0.009727626 |
| FDR q-value | 0.27484292 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | G6PC | G6PC Entrez, Source | glucose-6-phosphatase, catalytic subunit | 14 | 0.295 | 0.0409 | No |
| 2 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 35 | 0.229 | 0.0719 | No |
| 3 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 164 | 0.155 | 0.0843 | No |
| 4 | ZFP36 | ZFP36 Entrez, Source | zinc finger protein 36, C3H type, homolog (mouse) | 345 | 0.130 | 0.0891 | No |
| 5 | FZD2 | FZD2 Entrez, Source | frizzled homolog 2 (Drosophila) | 360 | 0.129 | 0.1064 | No |
| 6 | APOA1 | APOA1 Entrez, Source | apolipoprotein A-I | 552 | 0.113 | 0.1080 | No |
| 7 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 584 | 0.111 | 0.1215 | No |
| 8 | LY6G6C | LY6G6C Entrez, Source | lymphocyte antigen 6 complex, locus G6C | 761 | 0.101 | 0.1226 | No |
| 9 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 816 | 0.099 | 0.1325 | No |
| 10 | IER3 | IER3 Entrez, Source | immediate early response 3 | 890 | 0.096 | 0.1406 | No |
| 11 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 906 | 0.095 | 0.1530 | No |
| 12 | INSIG1 | INSIG1 Entrez, Source | insulin induced gene 1 | 978 | 0.092 | 0.1607 | No |
| 13 | CLK1 | CLK1 Entrez, Source | CDC-like kinase 1 | 1018 | 0.091 | 0.1707 | No |
| 14 | CNR1 | CNR1 Entrez, Source | cannabinoid receptor 1 (brain) | 1085 | 0.088 | 0.1781 | No |
| 15 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 1238 | 0.082 | 0.1783 | No |
| 16 | CD80 | CD80 Entrez, Source | CD80 molecule | 1490 | 0.075 | 0.1700 | No |
| 17 | PDE8B | PDE8B Entrez, Source | phosphodiesterase 8B | 1669 | 0.071 | 0.1666 | No |
| 18 | ADD3 | ADD3 Entrez, Source | adducin 3 (gamma) | 1701 | 0.070 | 0.1742 | No |
| 19 | GATA2 | GATA2 Entrez, Source | GATA binding protein 2 | 1755 | 0.069 | 0.1799 | No |
| 20 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 1837 | 0.067 | 0.1832 | No |
| 21 | SDC4 | SDC4 Entrez, Source | syndecan 4 (amphiglycan, ryudocan) | 2339 | 0.055 | 0.1532 | No |
| 22 | ITGB4 | ITGB4 Entrez, Source | integrin, beta 4 | 2487 | 0.052 | 0.1494 | No |
| 23 | GFAP | GFAP Entrez, Source | glial fibrillary acidic protein | 2549 | 0.051 | 0.1520 | No |
| 24 | KHK | KHK Entrez, Source | ketohexokinase (fructokinase) | 2773 | 0.047 | 0.1418 | No |
| 25 | SRPR | SRPR Entrez, Source | signal recognition particle receptor ('docking protein') | 2790 | 0.046 | 0.1471 | No |
| 26 | KLF6 | KLF6 Entrez, Source | Kruppel-like factor 6 | 2960 | 0.044 | 0.1406 | No |
| 27 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 3035 | 0.043 | 0.1410 | No |
| 28 | DMN | DMN Entrez, Source | desmuslin | 3532 | 0.035 | 0.1085 | No |
| 29 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 3705 | 0.032 | 0.1001 | No |
| 30 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 3724 | 0.032 | 0.1033 | No |
| 31 | STIP1 | STIP1 Entrez, Source | stress-induced-phosphoprotein 1 (Hsp70/Hsp90-organizing protein) | 3797 | 0.031 | 0.1023 | No |
| 32 | PLK2 | PLK2 Entrez, Source | polo-like kinase 2 (Drosophila) | 3889 | 0.030 | 0.0997 | No |
| 33 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 4037 | 0.028 | 0.0925 | No |
| 34 | SF3B1 | SF3B1 Entrez, Source | splicing factor 3b, subunit 1, 155kDa | 4132 | 0.027 | 0.0892 | No |
| 35 | CD28 | CD28 Entrez, Source | CD28 molecule | 4201 | 0.026 | 0.0877 | No |
| 36 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 4407 | 0.023 | 0.0755 | No |
| 37 | NR3C1 | NR3C1 Entrez, Source | nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) | 4473 | 0.022 | 0.0738 | No |
| 38 | JUN | JUN Entrez, Source | jun oncogene | 4841 | 0.018 | 0.0486 | No |
| 39 | C1QB | C1QB Entrez, Source | complement component 1, q subcomponent, B chain | 4891 | 0.018 | 0.0474 | No |
| 40 | UGP2 | UGP2 Entrez, Source | UDP-glucose pyrophosphorylase 2 | 5449 | 0.011 | 0.0069 | No |
| 41 | DDX5 | DDX5 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 5 | 5457 | 0.011 | 0.0079 | No |
| 42 | TKT | TKT Entrez, Source | transketolase (Wernicke-Korsakoff syndrome) | 5509 | 0.010 | 0.0055 | No |
| 43 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 5728 | 0.008 | -0.0099 | No |
| 44 | SF3A3 | SF3A3 Entrez, Source | splicing factor 3a, subunit 3, 60kDa | 5760 | 0.007 | -0.0112 | No |
| 45 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 5790 | 0.007 | -0.0124 | No |
| 46 | NUP93 | NUP93 Entrez, Source | nucleoporin 93kDa | 5850 | 0.006 | -0.0160 | No |
| 47 | FN1 | FN1 Entrez, Source | fibronectin 1 | 6064 | 0.004 | -0.0316 | No |
| 48 | METAP2 | METAP2 Entrez, Source | methionyl aminopeptidase 2 | 6178 | 0.003 | -0.0397 | No |
| 49 | ATP6V1C1 | ATP6V1C1 Entrez, Source | ATPase, H+ transporting, lysosomal 42kDa, V1 subunit C1 | 6343 | 0.001 | -0.0520 | No |
| 50 | RPL8 | RPL8 Entrez, Source | ribosomal protein L8 | 6427 | -0.000 | -0.0582 | No |
| 51 | PPIE | PPIE Entrez, Source | peptidylprolyl isomerase E (cyclophilin E) | 6591 | -0.002 | -0.0703 | No |
| 52 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 6743 | -0.003 | -0.0812 | No |
| 53 | THOC1 | THOC1 Entrez, Source | THO complex 1 | 7179 | -0.007 | -0.1130 | No |
| 54 | NFKB1 | NFKB1 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 (p105) | 7260 | -0.008 | -0.1179 | No |
| 55 | SDC1 | SDC1 Entrez, Source | syndecan 1 | 7387 | -0.009 | -0.1261 | No |
| 56 | DDAH2 | DDAH2 Entrez, Source | dimethylarginine dimethylaminohydrolase 2 | 7463 | -0.010 | -0.1303 | No |
| 57 | EIF3S6 | EIF3S6 Entrez, Source | eukaryotic translation initiation factor 3, subunit 6 48kDa | 7516 | -0.011 | -0.1327 | No |
| 58 | TFF3 | TFF3 Entrez, Source | trefoil factor 3 (intestinal) | 7570 | -0.011 | -0.1352 | No |
| 59 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 7608 | -0.012 | -0.1363 | No |
| 60 | DDX1 | DDX1 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 1 | 7656 | -0.012 | -0.1381 | No |
| 61 | RPL32 | RPL32 Entrez, Source | ribosomal protein L32 | 7742 | -0.013 | -0.1427 | No |
| 62 | NDUFV1 | NDUFV1 Entrez, Source | NADH dehydrogenase (ubiquinone) flavoprotein 1, 51kDa | 8014 | -0.016 | -0.1609 | No |
| 63 | HSPE1 | HSPE1 Entrez, Source | heat shock 10kDa protein 1 (chaperonin 10) | 8023 | -0.016 | -0.1593 | No |
| 64 | NME2 | NME2 Entrez, Source | non-metastatic cells 2, protein (NM23B) expressed in | 8212 | -0.018 | -0.1710 | No |
| 65 | MARK3 | MARK3 Entrez, Source | MAP/microtubule affinity-regulating kinase 3 | 8547 | -0.021 | -0.1932 | No |
| 66 | TARS | TARS Entrez, Source | threonyl-tRNA synthetase | 8662 | -0.022 | -0.1986 | No |
| 67 | ABCG1 | ABCG1 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 1 | 8730 | -0.023 | -0.2004 | No |
| 68 | ITGB4BP | ITGB4BP Entrez, Source | integrin beta 4 binding protein | 8810 | -0.024 | -0.2029 | No |
| 69 | TRADD | TRADD Entrez, Source | TNFRSF1A-associated via death domain | 9040 | -0.026 | -0.2165 | No |
| 70 | EEF1D | EEF1D Entrez, Source | eukaryotic translation elongation factor 1 delta (guanine nucleotide exchange protein) | 9064 | -0.027 | -0.2144 | No |
| 71 | COPB2 | COPB2 Entrez, Source | coatomer protein complex, subunit beta 2 (beta prime) | 9086 | -0.027 | -0.2122 | No |
| 72 | GPX1 | GPX1 Entrez, Source | glutathione peroxidase 1 | 9784 | -0.035 | -0.2599 | No |
| 73 | GSK3B | GSK3B Entrez, Source | glycogen synthase kinase 3 beta | 10151 | -0.039 | -0.2820 | No |
| 74 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 10560 | -0.044 | -0.3065 | No |
| 75 | ANK1 | ANK1 Entrez, Source | ankyrin 1, erythrocytic | 10948 | -0.050 | -0.3286 | No |
| 76 | PHB | PHB Entrez, Source | prohibitin | 10993 | -0.051 | -0.3247 | No |
| 77 | CBR1 | CBR1 Entrez, Source | carbonyl reductase 1 | 10998 | -0.051 | -0.3178 | No |
| 78 | WASF1 | WASF1 Entrez, Source | WAS protein family, member 1 | 11176 | -0.054 | -0.3234 | No |
| 79 | MRPL2 | MRPL2 Entrez, Source | mitochondrial ribosomal protein L2 | 11209 | -0.055 | -0.3180 | No |
| 80 | FASTK | FASTK Entrez, Source | Fas-activated serine/threonine kinase | 11479 | -0.060 | -0.3298 | No |
| 81 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 11568 | -0.062 | -0.3277 | No |
| 82 | CKB | CKB Entrez, Source | creatine kinase, brain | 11647 | -0.064 | -0.3245 | No |
| 83 | VLDLR | VLDLR Entrez, Source | very low density lipoprotein receptor | 11859 | -0.069 | -0.3307 | No |
| 84 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 12342 | -0.085 | -0.3550 | Yes |
| 85 | PTGS2 | PTGS2 Entrez, Source | prostaglandin-endoperoxide synthase 2 (prostaglandin G/H synthase and cyclooxygenase) | 12422 | -0.088 | -0.3486 | Yes |
| 86 | GNAS | GNAS Entrez, Source | GNAS complex locus | 12462 | -0.089 | -0.3388 | Yes |
| 87 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 12511 | -0.092 | -0.3293 | Yes |
| 88 | SH3BP4 | SH3BP4 Entrez, Source | SH3-domain binding protein 4 | 12793 | -0.110 | -0.3349 | Yes |
| 89 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 12901 | -0.121 | -0.3258 | Yes |
| 90 | AKAP9 | AKAP9 Entrez, Source | A kinase (PRKA) anchor protein (yotiao) 9 | 12906 | -0.122 | -0.3088 | Yes |
| 91 | SIPA1L1 | SIPA1L1 Entrez, Source | signal-induced proliferation-associated 1 like 1 | 12930 | -0.125 | -0.2929 | Yes |
| 92 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 12964 | -0.127 | -0.2773 | Yes |
| 93 | VIL2 | VIL2 Entrez, Source | villin 2 (ezrin) | 13021 | -0.135 | -0.2624 | Yes |
| 94 | GSTM5 | GSTM5 Entrez, Source | glutathione S-transferase M5 | 13155 | -0.164 | -0.2491 | Yes |
| 95 | SYNJ2 | SYNJ2 Entrez, Source | synaptojanin 2 | 13158 | -0.165 | -0.2258 | Yes |
| 96 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 13185 | -0.179 | -0.2024 | Yes |
| 97 | CLIC1 | CLIC1 Entrez, Source | chloride intracellular channel 1 | 13242 | -0.218 | -0.1757 | Yes |
| 98 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 13262 | -0.231 | -0.1444 | Yes |
| 99 | IRF1 | IRF1 Entrez, Source | interferon regulatory factor 1 | 13267 | -0.235 | -0.1113 | Yes |
| 100 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13284 | -0.263 | -0.0752 | Yes |
| 101 | SERPINH1 | SERPINH1 Entrez, Source | serpin peptidase inhibitor, clade H (heat shock protein 47), member 1, (collagen binding protein 1) | 13285 | -0.265 | -0.0376 | Yes |
| 102 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 13302 | -0.295 | 0.0030 | Yes |