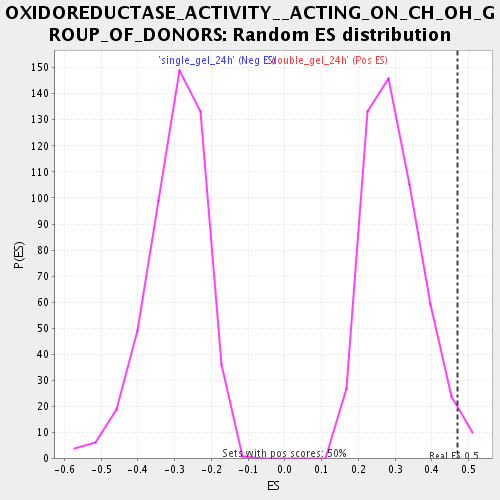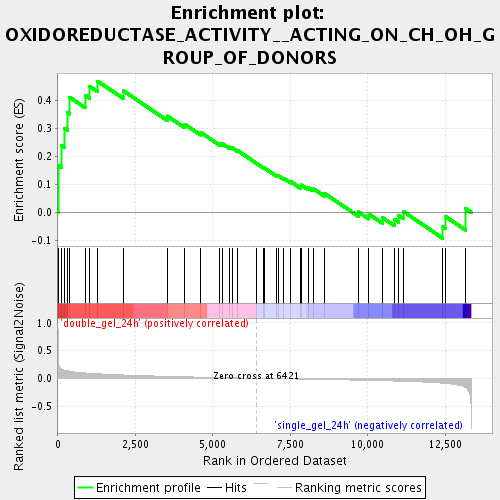
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | OXIDOREDUCTASE_ACTIVITY__ACTING_ON_CH_OH_GROUP_OF_DONORS |
| Enrichment Score (ES) | 0.4694601 |
| Normalized Enrichment Score (NES) | 1.5717043 |
| Nominal p-value | 0.029761905 |
| FDR q-value | 0.23321298 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 7 | 0.362 | 0.1690 | Yes |
| 2 | BDH1 | BDH1 Entrez, Source | 3-hydroxybutyrate dehydrogenase, type 1 | 121 | 0.169 | 0.2395 | Yes |
| 3 | ADH7 | ADH7 Entrez, Source | alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide | 214 | 0.147 | 0.3013 | Yes |
| 4 | HSD17B7 | HSD17B7 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 7 | 313 | 0.134 | 0.3566 | Yes |
| 5 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 379 | 0.127 | 0.4114 | Yes |
| 6 | MIOX | MIOX Entrez, Source | myo-inositol oxygenase | 881 | 0.096 | 0.4188 | Yes |
| 7 | HSD3B1 | HSD3B1 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 | 1022 | 0.091 | 0.4507 | Yes |
| 8 | AKR1B10 | AKR1B10 Entrez, Source | aldo-keto reductase family 1, member B10 (aldose reductase) | 1279 | 0.081 | 0.4695 | Yes |
| 9 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 2110 | 0.060 | 0.4351 | No |
| 10 | LDHA | LDHA Entrez, Source | lactate dehydrogenase A | 3529 | 0.035 | 0.3448 | No |
| 11 | GPD2 | GPD2 Entrez, Source | glycerol-3-phosphate dehydrogenase 2 (mitochondrial) | 4091 | 0.027 | 0.3154 | No |
| 12 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 4607 | 0.021 | 0.2863 | No |
| 13 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 5228 | 0.013 | 0.2460 | No |
| 14 | MDH1 | MDH1 Entrez, Source | malate dehydrogenase 1, NAD (soluble) | 5310 | 0.012 | 0.2458 | No |
| 15 | ADH6 | ADH6 Entrez, Source | alcohol dehydrogenase 6 (class V) | 5527 | 0.010 | 0.2342 | No |
| 16 | DHRS9 | DHRS9 Entrez, Source | dehydrogenase/reductase (SDR family) member 9 | 5622 | 0.009 | 0.2313 | No |
| 17 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 5790 | 0.007 | 0.2219 | No |
| 18 | SPR | SPR Entrez, Source | sepiapterin reductase (7,8-dihydrobiopterin:NADP+ oxidoreductase) | 6410 | 0.000 | 0.1755 | No |
| 19 | ADH4 | ADH4 Entrez, Source | alcohol dehydrogenase 4 (class II), pi polypeptide | 6634 | -0.002 | 0.1598 | No |
| 20 | HSD17B4 | HSD17B4 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 4 | 6653 | -0.002 | 0.1596 | No |
| 21 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 7066 | -0.006 | 0.1316 | No |
| 22 | AKR7A2 | AKR7A2 Entrez, Source | aldo-keto reductase family 7, member A2 (aflatoxin aldehyde reductase) | 7115 | -0.007 | 0.1311 | No |
| 23 | IMPDH2 | IMPDH2 Entrez, Source | IMP (inosine monophosphate) dehydrogenase 2 | 7288 | -0.009 | 0.1222 | No |
| 24 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 7513 | -0.011 | 0.1103 | No |
| 25 | RDH10 | RDH10 Entrez, Source | retinol dehydrogenase 10 (all-trans) | 7839 | -0.014 | 0.0924 | No |
| 26 | LDHC | LDHC Entrez, Source | lactate dehydrogenase C | 7848 | -0.014 | 0.0984 | No |
| 27 | RDH11 | RDH11 Entrez, Source | retinol dehydrogenase 11 (all-trans/9-cis/11-cis) | 8080 | -0.017 | 0.0888 | No |
| 28 | AKR1A1 | AKR1A1 Entrez, Source | aldo-keto reductase family 1, member A1 (aldehyde reductase) | 8240 | -0.018 | 0.0853 | No |
| 29 | IDH1 | IDH1 Entrez, Source | isocitrate dehydrogenase 1 (NADP+), soluble | 8609 | -0.022 | 0.0680 | No |
| 30 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 9698 | -0.034 | 0.0021 | No |
| 31 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 10037 | -0.038 | -0.0057 | No |
| 32 | IDH3G | IDH3G Entrez, Source | isocitrate dehydrogenase 3 (NAD+) gamma | 10481 | -0.043 | -0.0188 | No |
| 33 | IDH3B | IDH3B Entrez, Source | isocitrate dehydrogenase 3 (NAD+) beta | 10868 | -0.049 | -0.0248 | No |
| 34 | CBR1 | CBR1 Entrez, Source | carbonyl reductase 1 | 10998 | -0.051 | -0.0106 | No |
| 35 | AKR7A3 | AKR7A3 Entrez, Source | aldo-keto reductase family 7, member A3 (aflatoxin aldehyde reductase) | 11159 | -0.054 | 0.0027 | No |
| 36 | PHGDH | PHGDH Entrez, Source | phosphoglycerate dehydrogenase | 12426 | -0.088 | -0.0513 | No |
| 37 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 12511 | -0.092 | -0.0145 | No |
| 38 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 13156 | -0.164 | 0.0140 | No |