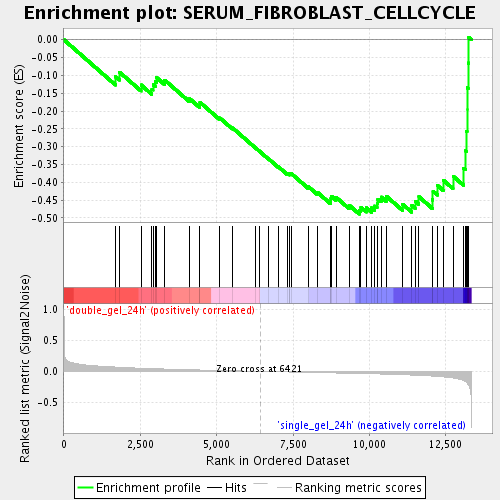
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | SERUM_FIBROBLAST_CELLCYCLE |
| Enrichment Score (ES) | -0.48938286 |
| Normalized Enrichment Score (NES) | -1.8136655 |
| Nominal p-value | 0.003929273 |
| FDR q-value | 0.11624012 |
| FWER p-Value | 0.961 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MAPK13 | MAPK13 Entrez, Source | mitogen-activated protein kinase 13 | 1690 | 0.070 | -0.1037 | No |
| 2 | WSB1 | WSB1 Entrez, Source | WD repeat and SOCS box-containing 1 | 1817 | 0.067 | -0.0907 | No |
| 3 | DHFR | DHFR Entrez, Source | dihydrofolate reductase | 2524 | 0.051 | -0.1267 | No |
| 4 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 2878 | 0.045 | -0.1383 | No |
| 5 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 2916 | 0.044 | -0.1262 | No |
| 6 | CIITA | CIITA Entrez, Source | class II, major histocompatibility complex, transactivator | 2991 | 0.043 | -0.1173 | No |
| 7 | RANGAP1 | RANGAP1 Entrez, Source | Ran GTPase activating protein 1 | 3033 | 0.043 | -0.1061 | No |
| 8 | CDKN3 | CDKN3 Entrez, Source | cyclin-dependent kinase inhibitor 3 (CDK2-associated dual specificity phosphatase) | 3296 | 0.038 | -0.1130 | No |
| 9 | CCNF | CCNF Entrez, Source | cyclin F | 4114 | 0.027 | -0.1655 | No |
| 10 | SPAG5 | SPAG5 Entrez, Source | sperm associated antigen 5 | 4449 | 0.023 | -0.1830 | No |
| 11 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 4450 | 0.023 | -0.1755 | No |
| 12 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 5100 | 0.015 | -0.2193 | No |
| 13 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 5517 | 0.010 | -0.2473 | No |
| 14 | FBXL20 | FBXL20 Entrez, Source | F-box and leucine-rich repeat protein 20 | 6285 | 0.001 | -0.3045 | No |
| 15 | TUBB2C | TUBB2C Entrez, Source | tubulin, beta 2C | 6401 | 0.000 | -0.3130 | No |
| 16 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 6701 | -0.003 | -0.3346 | No |
| 17 | ANP32E | ANP32E Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member E | 7007 | -0.006 | -0.3556 | No |
| 18 | CASP3 | CASP3 Entrez, Source | caspase 3, apoptosis-related cysteine peptidase | 7301 | -0.009 | -0.3748 | No |
| 19 | CDCA1 | CDCA1 Entrez, Source | cell division cycle associated 1 | 7368 | -0.009 | -0.3767 | No |
| 20 | SDC1 | SDC1 Entrez, Source | syndecan 1 | 7387 | -0.009 | -0.3749 | No |
| 21 | LYAR | LYAR Entrez, Source | - | 7434 | -0.010 | -0.3751 | No |
| 22 | PAQR4 | PAQR4 Entrez, Source | progestin and adipoQ receptor family member IV | 7999 | -0.016 | -0.4123 | No |
| 23 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 8303 | -0.019 | -0.4288 | No |
| 24 | TACC3 | TACC3 Entrez, Source | transforming, acidic coiled-coil containing protein 3 | 8727 | -0.023 | -0.4529 | No |
| 25 | ABCC5 | ABCC5 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 5 | 8737 | -0.023 | -0.4458 | No |
| 26 | HN1 | HN1 Entrez, Source | hematological and neurological expressed 1 | 8753 | -0.023 | -0.4391 | No |
| 27 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 8905 | -0.025 | -0.4421 | No |
| 28 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9337 | -0.029 | -0.4646 | No |
| 29 | AURKA | AURKA Entrez, Source | aurora kinase A | 9667 | -0.034 | -0.4781 | Yes |
| 30 | PRIM2A | PRIM2A Entrez, Source | primase, polypeptide 2A, 58kDa | 9718 | -0.034 | -0.4704 | Yes |
| 31 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 9894 | -0.036 | -0.4715 | Yes |
| 32 | CDCA8 | CDCA8 Entrez, Source | cell division cycle associated 8 | 10068 | -0.038 | -0.4718 | Yes |
| 33 | DONSON | DONSON Entrez, Source | downstream neighbor of SON | 10174 | -0.039 | -0.4666 | Yes |
| 34 | RFC2 | RFC2 Entrez, Source | replication factor C (activator 1) 2, 40kDa | 10249 | -0.040 | -0.4588 | Yes |
| 35 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 10273 | -0.040 | -0.4470 | Yes |
| 36 | YWHAH | YWHAH Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide | 10384 | -0.042 | -0.4414 | Yes |
| 37 | ANKRD10 | ANKRD10 Entrez, Source | ankyrin repeat domain 10 | 10548 | -0.044 | -0.4389 | Yes |
| 38 | BARD1 | BARD1 Entrez, Source | BRCA1 associated RING domain 1 | 11085 | -0.053 | -0.4616 | Yes |
| 39 | TUBB | TUBB Entrez, Source | tubulin, beta | 11384 | -0.058 | -0.4645 | Yes |
| 40 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 11497 | -0.060 | -0.4527 | Yes |
| 41 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 11618 | -0.063 | -0.4407 | Yes |
| 42 | KIFC1 | KIFC1 Entrez, Source | kinesin family member C1 | 12059 | -0.075 | -0.4488 | Yes |
| 43 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 12078 | -0.075 | -0.4249 | Yes |
| 44 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 12226 | -0.080 | -0.4091 | Yes |
| 45 | TIMP1 | TIMP1 Entrez, Source | TIMP metallopeptidase inhibitor 1 | 12415 | -0.087 | -0.3940 | Yes |
| 46 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 12735 | -0.105 | -0.3828 | Yes |
| 47 | TRIP13 | TRIP13 Entrez, Source | thyroid hormone receptor interactor 13 | 13078 | -0.144 | -0.3602 | Yes |
| 48 | MET | MET Entrez, Source | met proto-oncogene (hepatocyte growth factor receptor) | 13148 | -0.161 | -0.3116 | Yes |
| 49 | ADAMTS1 | ADAMTS1 Entrez, Source | ADAM metallopeptidase with thrombospondin type 1 motif, 1 | 13169 | -0.170 | -0.2563 | Yes |
| 50 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13194 | -0.185 | -0.1961 | Yes |
| 51 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13196 | -0.186 | -0.1338 | Yes |
| 52 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 13232 | -0.213 | -0.0650 | Yes |
| 53 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 13248 | -0.219 | 0.0071 | Yes |

