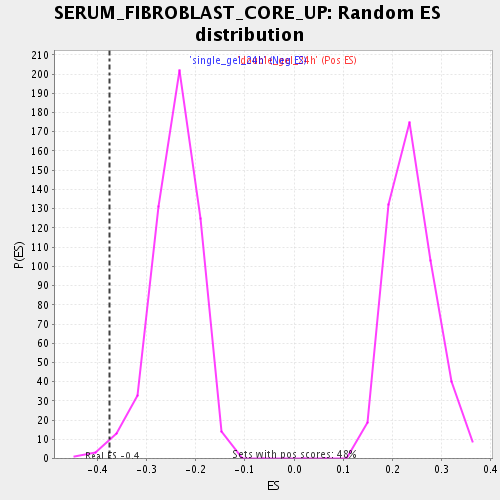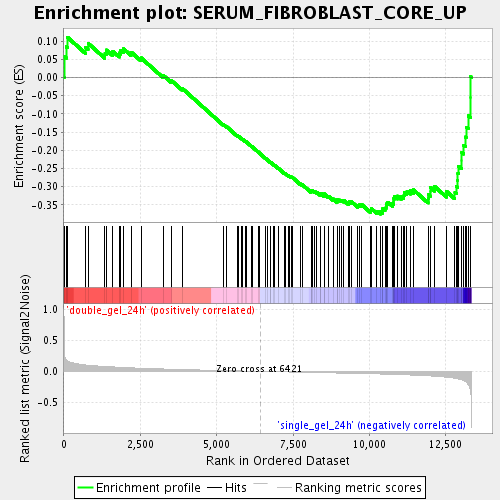
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | SERUM_FIBROBLAST_CORE_UP |
| Enrichment Score (ES) | -0.3754176 |
| Normalized Enrichment Score (NES) | -1.5526831 |
| Nominal p-value | 0.011494253 |
| FDR q-value | 0.25226828 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MAP3K8 | MAP3K8 Entrez, Source | mitogen-activated protein kinase kinase kinase 8 | 9 | 0.350 | 0.0584 | No |
| 2 | EMP2 | EMP2 Entrez, Source | epithelial membrane protein 2 | 79 | 0.187 | 0.0847 | No |
| 3 | PDIA4 | PDIA4 Entrez, Source | protein disulfide isomerase family A, member 4 | 116 | 0.170 | 0.1107 | No |
| 4 | HNRPA3 | HNRPA3 Entrez, Source | heterogeneous nuclear ribonucleoprotein A3 | 711 | 0.104 | 0.0833 | No |
| 5 | CDCA4 | CDCA4 Entrez, Source | cell division cycle associated 4 | 791 | 0.100 | 0.0942 | No |
| 6 | HAS2 | HAS2 Entrez, Source | hyaluronan synthase 2 | 1343 | 0.079 | 0.0659 | No |
| 7 | EPHB1 | EPHB1 Entrez, Source | EPH receptor B1 | 1387 | 0.078 | 0.0758 | No |
| 8 | DCK | DCK Entrez, Source | deoxycytidine kinase | 1598 | 0.072 | 0.0722 | No |
| 9 | FLNC | FLNC Entrez, Source | filamin C, gamma (actin binding protein 280) | 1809 | 0.067 | 0.0677 | No |
| 10 | MT3 | MT3 Entrez, Source | metallothionein 3 (growth inhibitory factor (neurotrophic)) | 1858 | 0.066 | 0.0752 | No |
| 11 | RNF138 | RNF138 Entrez, Source | ring finger protein 138 | 1949 | 0.064 | 0.0791 | No |
| 12 | SSR3 | SSR3 Entrez, Source | signal sequence receptor, gamma (translocon-associated protein gamma) | 2198 | 0.057 | 0.0701 | No |
| 13 | DHFR | DHFR Entrez, Source | dihydrofolate reductase | 2524 | 0.051 | 0.0542 | No |
| 14 | NUP107 | NUP107 Entrez, Source | nucleoporin 107kDa | 3252 | 0.039 | 0.0059 | No |
| 15 | NUPL1 | NUPL1 Entrez, Source | nucleoporin like 1 | 3508 | 0.035 | -0.0075 | No |
| 16 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 3869 | 0.030 | -0.0296 | No |
| 17 | BRCA2 | BRCA2 Entrez, Source | breast cancer 2, early onset | 5213 | 0.014 | -0.1288 | No |
| 18 | SFRS10 | SFRS10 Entrez, Source | splicing factor, arginine/serine-rich 10 (transformer 2 homolog, Drosophila) | 5320 | 0.012 | -0.1347 | No |
| 19 | SAR1B | SAR1B Entrez, Source | SAR1 gene homolog B (S. cerevisiae) | 5669 | 0.008 | -0.1596 | No |
| 20 | RAB3B | RAB3B Entrez, Source | RAB3B, member RAS oncogene family | 5708 | 0.008 | -0.1611 | No |
| 21 | HMGN2 | HMGN2 Entrez, Source | high-mobility group nucleosomal binding domain 2 | 5816 | 0.007 | -0.1681 | No |
| 22 | NUP93 | NUP93 Entrez, Source | nucleoporin 93kDa | 5850 | 0.006 | -0.1696 | No |
| 23 | GPLD1 | GPLD1 Entrez, Source | glycosylphosphatidylinositol specific phospholipase D1 | 5931 | 0.005 | -0.1747 | No |
| 24 | NUP35 | NUP35 Entrez, Source | nucleoporin 35kDa | 5971 | 0.005 | -0.1769 | No |
| 25 | CCND3 | CCND3 Entrez, Source | cyclin D3 | 6145 | 0.003 | -0.1894 | No |
| 26 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 6161 | 0.003 | -0.1901 | No |
| 27 | ACTL6A | ACTL6A Entrez, Source | actin-like 6A | 6365 | 0.001 | -0.2053 | No |
| 28 | LYPLA2 | LYPLA2 Entrez, Source | lysophospholipase II | 6392 | 0.000 | -0.2072 | No |
| 29 | TNFRSF12A | TNFRSF12A Entrez, Source | tumor necrosis factor receptor superfamily, member 12A | 6604 | -0.002 | -0.2228 | No |
| 30 | SLC25A5 | SLC25A5 Entrez, Source | solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 5 | 6656 | -0.003 | -0.2262 | No |
| 31 | SNRPB | SNRPB Entrez, Source | small nuclear ribonucleoprotein polypeptides B and B1 | 6753 | -0.003 | -0.2329 | No |
| 32 | HNRPAB | HNRPAB Entrez, Source | heterogeneous nuclear ribonucleoprotein A/B | 6758 | -0.003 | -0.2326 | No |
| 33 | SNRPA | SNRPA Entrez, Source | small nuclear ribonucleoprotein polypeptide A | 6866 | -0.004 | -0.2400 | No |
| 34 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 6886 | -0.005 | -0.2406 | No |
| 35 | WDR77 | WDR77 Entrez, Source | WD repeat domain 77 | 7028 | -0.006 | -0.2503 | No |
| 36 | PSMD12 | PSMD12 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 12 | 7220 | -0.008 | -0.2634 | No |
| 37 | EBNA1BP2 | EBNA1BP2 Entrez, Source | EBNA1 binding protein 2 | 7242 | -0.008 | -0.2636 | No |
| 38 | EIF4EBP1 | EIF4EBP1 Entrez, Source | eukaryotic translation initiation factor 4E binding protein 1 | 7338 | -0.009 | -0.2693 | No |
| 39 | SDC1 | SDC1 Entrez, Source | syndecan 1 | 7387 | -0.009 | -0.2714 | No |
| 40 | LYAR | LYAR Entrez, Source | - | 7434 | -0.010 | -0.2732 | No |
| 41 | CENPJ | CENPJ Entrez, Source | centromere protein J | 7469 | -0.010 | -0.2740 | No |
| 42 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 7738 | -0.013 | -0.2921 | No |
| 43 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 7800 | -0.014 | -0.2944 | No |
| 44 | TFPI2 | TFPI2 Entrez, Source | tissue factor pathway inhibitor 2 | 8088 | -0.017 | -0.3133 | No |
| 45 | FARSLB | FARSLB Entrez, Source | phenylalanine-tRNA synthetase-like, beta subunit | 8103 | -0.017 | -0.3115 | No |
| 46 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 8126 | -0.017 | -0.3103 | No |
| 47 | RPN1 | RPN1 Entrez, Source | ribophorin I | 8214 | -0.018 | -0.3139 | No |
| 48 | RUVBL1 | RUVBL1 Entrez, Source | RuvB-like 1 (E. coli) | 8268 | -0.018 | -0.3148 | No |
| 49 | UMPS | UMPS Entrez, Source | uridine monophosphate synthetase (orotate phosphoribosyl transferase and orotidine-5'-decarboxylase) | 8391 | -0.020 | -0.3207 | No |
| 50 | PFN1 | PFN1 Entrez, Source | profilin 1 | 8402 | -0.020 | -0.3182 | No |
| 51 | PSMD2 | PSMD2 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 2 | 8512 | -0.021 | -0.3229 | No |
| 52 | MAPRE1 | MAPRE1 Entrez, Source | microtubule-associated protein, RP/EB family, member 1 | 8516 | -0.021 | -0.3196 | No |
| 53 | RNF41 | RNF41 Entrez, Source | ring finger protein 41 | 8665 | -0.022 | -0.3270 | No |
| 54 | CEP78 | CEP78 Entrez, Source | centrosomal protein 78kDa | 8809 | -0.024 | -0.3337 | No |
| 55 | TOMM40 | TOMM40 Entrez, Source | translocase of outer mitochondrial membrane 40 homolog (yeast) | 8940 | -0.025 | -0.3393 | No |
| 56 | CCT5 | CCT5 Entrez, Source | chaperonin containing TCP1, subunit 5 (epsilon) | 8949 | -0.025 | -0.3356 | No |
| 57 | TXNL2 | TXNL2 Entrez, Source | thioredoxin-like 2 | 9021 | -0.026 | -0.3365 | No |
| 58 | COPB2 | COPB2 Entrez, Source | coatomer protein complex, subunit beta 2 (beta prime) | 9086 | -0.027 | -0.3368 | No |
| 59 | STX3 | STX3 Entrez, Source | syntaxin 3 | 9160 | -0.028 | -0.3377 | No |
| 60 | EIF4G1 | EIF4G1 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 1 | 9329 | -0.029 | -0.3454 | No |
| 61 | MTHFD1 | MTHFD1 Entrez, Source | methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1, methenyltetrahydrofolate cyclohydrolase, formyltetrahydrofolate synthetase | 9334 | -0.029 | -0.3408 | No |
| 62 | DUT | DUT Entrez, Source | dUTP pyrophosphatase | 9409 | -0.030 | -0.3412 | No |
| 63 | MNAT1 | MNAT1 Entrez, Source | menage a trois homolog 1, cyclin H assembly factor (Xenopus laevis) | 9616 | -0.033 | -0.3512 | No |
| 64 | CLCF1 | CLCF1 Entrez, Source | cardiotrophin-like cytokine factor 1 | 9674 | -0.034 | -0.3499 | No |
| 65 | PSMA7 | PSMA7 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 7 | 9751 | -0.034 | -0.3498 | No |
| 66 | RNASEH2A | RNASEH2A Entrez, Source | ribonuclease H2, subunit A | 10032 | -0.037 | -0.3646 | No |
| 67 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 10063 | -0.038 | -0.3605 | No |
| 68 | PSMC3 | PSMC3 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 3 | 10244 | -0.040 | -0.3673 | No |
| 69 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 10352 | -0.041 | -0.3685 | Yes |
| 70 | MRPL37 | MRPL37 Entrez, Source | mitochondrial ribosomal protein L37 | 10420 | -0.042 | -0.3664 | Yes |
| 71 | POLE3 | POLE3 Entrez, Source | polymerase (DNA directed), epsilon 3 (p17 subunit) | 10436 | -0.042 | -0.3604 | Yes |
| 72 | PDLIM7 | PDLIM7 Entrez, Source | PDZ and LIM domain 7 (enigma) | 10511 | -0.044 | -0.3586 | Yes |
| 73 | MRPL12 | MRPL12 Entrez, Source | mitochondrial ribosomal protein L12 | 10545 | -0.044 | -0.3537 | Yes |
| 74 | NCLN | NCLN Entrez, Source | nicalin homolog (zebrafish) | 10562 | -0.044 | -0.3474 | Yes |
| 75 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 10597 | -0.045 | -0.3425 | Yes |
| 76 | NPTN | NPTN Entrez, Source | neuroplastin | 10756 | -0.047 | -0.3464 | Yes |
| 77 | MKKS | MKKS Entrez, Source | McKusick-Kaufman syndrome | 10791 | -0.048 | -0.3409 | Yes |
| 78 | PCSK7 | PCSK7 Entrez, Source | proprotein convertase subtilisin/kexin type 7 | 10794 | -0.048 | -0.3330 | Yes |
| 79 | HNRPR | HNRPR Entrez, Source | heterogeneous nuclear ribonucleoprotein R | 10811 | -0.048 | -0.3261 | Yes |
| 80 | DCLRE1B | DCLRE1B Entrez, Source | DNA cross-link repair 1B (PSO2 homolog, S. cerevisiae) | 10917 | -0.050 | -0.3256 | Yes |
| 81 | FABP5 | FABP5 Entrez, Source | fatty acid binding protein 5 (psoriasis-associated) | 11036 | -0.052 | -0.3258 | Yes |
| 82 | RPP40 | RPP40 Entrez, Source | ribonuclease P 40kDa subunit | 11128 | -0.053 | -0.3237 | Yes |
| 83 | GNG11 | GNG11 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 11 | 11142 | -0.054 | -0.3156 | Yes |
| 84 | MCTS1 | MCTS1 Entrez, Source | malignant T cell amplified sequence 1 | 11227 | -0.056 | -0.3126 | Yes |
| 85 | TDP1 | TDP1 Entrez, Source | tyrosyl-DNA phosphodiesterase 1 | 11332 | -0.058 | -0.3107 | Yes |
| 86 | ST3GAL6 | ST3GAL6 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 6 | 11435 | -0.059 | -0.3084 | Yes |
| 87 | NLN | NLN Entrez, Source | neurolysin (metallopeptidase M3 family) | 11920 | -0.070 | -0.3331 | Yes |
| 88 | ACTC1 | ACTC1 Entrez, Source | actin, alpha, cardiac muscle 1 | 11932 | -0.071 | -0.3220 | Yes |
| 89 | DCBLD2 | DCBLD2 Entrez, Source | discoidin, CUB and LCCL domain containing 2 | 11988 | -0.072 | -0.3139 | Yes |
| 90 | NUDT1 | NUDT1 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 1 | 11990 | -0.073 | -0.3018 | Yes |
| 91 | GGH | GGH Entrez, Source | gamma-glutamyl hydrolase (conjugase, folylpolygammaglutamyl hydrolase) | 12131 | -0.077 | -0.2994 | Yes |
| 92 | CCDC5 | CCDC5 Entrez, Source | coiled-coil domain containing 5 (spindle associated) | 12521 | -0.093 | -0.3131 | Yes |
| 93 | WSB2 | WSB2 Entrez, Source | WD repeat and SOCS box-containing 2 | 12797 | -0.110 | -0.3152 | Yes |
| 94 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 12847 | -0.115 | -0.2996 | Yes |
| 95 | SMS | SMS Entrez, Source | spermine synthase | 12884 | -0.119 | -0.2822 | Yes |
| 96 | PTS | PTS Entrez, Source | 6-pyruvoyltetrahydropterin synthase | 12892 | -0.120 | -0.2625 | Yes |
| 97 | RFC3 | RFC3 Entrez, Source | replication factor C (activator 1) 3, 38kDa | 12915 | -0.123 | -0.2434 | Yes |
| 98 | TPRKB | TPRKB Entrez, Source | TP53RK binding protein | 13020 | -0.135 | -0.2285 | Yes |
| 99 | VIL2 | VIL2 Entrez, Source | villin 2 (ezrin) | 13021 | -0.135 | -0.2057 | Yes |
| 100 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 13084 | -0.146 | -0.1857 | Yes |
| 101 | MET | MET Entrez, Source | met proto-oncogene (hepatocyte growth factor receptor) | 13148 | -0.161 | -0.1634 | Yes |
| 102 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 13167 | -0.169 | -0.1362 | Yes |
| 103 | MSN | MSN Entrez, Source | moesin | 13233 | -0.213 | -0.1051 | Yes |
| 104 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 13312 | -0.331 | -0.0553 | Yes |
| 105 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 13314 | -0.341 | 0.0021 | Yes |