Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
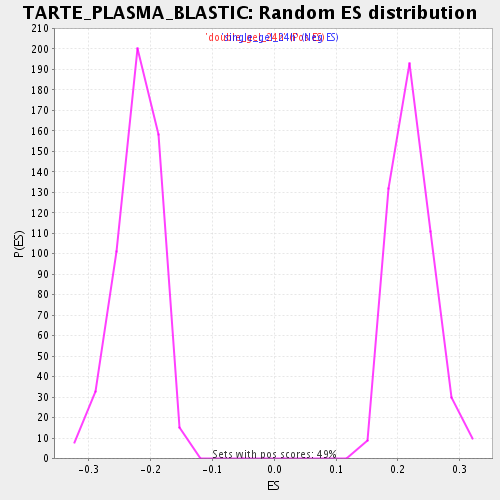
| GeneSet | TARTE_PLASMA_BLASTIC |
| Enrichment Score (ES) | -0.38076103 |
| Normalized Enrichment Score (NES) | -1.7217606 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.1493625 |
| FWER p-Value | 0.999 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 308 | 0.134 | -0.0081 | No |
| 2 | SELL | SELL Entrez, Source | selectin L (lymphocyte adhesion molecule 1) | 417 | 0.125 | -0.0022 | No |
| 3 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 491 | 0.118 | 0.0058 | No |
| 4 | FASN | FASN Entrez, Source | fatty acid synthase | 607 | 0.110 | 0.0095 | No |
| 5 | FDFT1 | FDFT1 Entrez, Source | farnesyl-diphosphate farnesyltransferase 1 | 952 | 0.094 | -0.0060 | No |
| 6 | SLC16A1 | SLC16A1 Entrez, Source | solute carrier family 16, member 1 (monocarboxylic acid transporter 1) | 1229 | 0.083 | -0.0175 | No |
| 7 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 1348 | 0.079 | -0.0175 | No |
| 8 | GZMB | GZMB Entrez, Source | granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) | 1567 | 0.073 | -0.0258 | No |
| 9 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 1596 | 0.073 | -0.0197 | No |
| 10 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 1609 | 0.072 | -0.0123 | No |
| 11 | ENTPD1 | ENTPD1 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 1 | 1610 | 0.072 | -0.0041 | No |
| 12 | FDPS | FDPS Entrez, Source | farnesyl diphosphate synthase (farnesyl pyrophosphate synthetase, dimethylallyltranstransferase, geranyltranstransferase) | 1791 | 0.068 | -0.0101 | No |
| 13 | LGALS1 | LGALS1 Entrez, Source | lectin, galactoside-binding, soluble, 1 (galectin 1) | 1865 | 0.066 | -0.0081 | No |
| 14 | PRKCB1 | PRKCB1 Entrez, Source | protein kinase C, beta 1 | 2052 | 0.061 | -0.0153 | No |
| 15 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 2110 | 0.060 | -0.0128 | No |
| 16 | GALNT3 | GALNT3 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 3 (GalNAc-T3) | 2156 | 0.059 | -0.0096 | No |
| 17 | BYSL | BYSL Entrez, Source | bystin-like | 2678 | 0.048 | -0.0437 | No |
| 18 | GNL2 | GNL2 Entrez, Source | guanine nucleotide binding protein-like 2 (nucleolar) | 2742 | 0.047 | -0.0431 | No |
| 19 | ARHGDIB | ARHGDIB Entrez, Source | Rho GDP dissociation inhibitor (GDI) beta | 2879 | 0.045 | -0.0483 | No |
| 20 | IL2RB | IL2RB Entrez, Source | interleukin 2 receptor, beta | 3050 | 0.042 | -0.0564 | No |
| 21 | LDLR | LDLR Entrez, Source | low density lipoprotein receptor (familial hypercholesterolemia) | 3110 | 0.041 | -0.0562 | No |
| 22 | CDKN3 | CDKN3 Entrez, Source | cyclin-dependent kinase inhibitor 3 (CDK2-associated dual specificity phosphatase) | 3296 | 0.038 | -0.0659 | No |
| 23 | NFIL3 | NFIL3 Entrez, Source | nuclear factor, interleukin 3 regulated | 3661 | 0.033 | -0.0898 | No |
| 24 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 3705 | 0.032 | -0.0894 | No |
| 25 | PPIF | PPIF Entrez, Source | peptidylprolyl isomerase F (cyclophilin F) | 3868 | 0.030 | -0.0983 | No |
| 26 | GLIPR1 | GLIPR1 Entrez, Source | GLI pathogenesis-related 1 (glioma) | 3896 | 0.030 | -0.0970 | No |
| 27 | CCNF | CCNF Entrez, Source | cyclin F | 4114 | 0.027 | -0.1104 | No |
| 28 | EBP | EBP Entrez, Source | emopamil binding protein (sterol isomerase) | 4120 | 0.027 | -0.1077 | No |
| 29 | CORO1A | CORO1A Entrez, Source | coronin, actin binding protein, 1A | 4124 | 0.027 | -0.1049 | No |
| 30 | SSR1 | SSR1 Entrez, Source | signal sequence receptor, alpha (translocon-associated protein alpha) | 4186 | 0.026 | -0.1066 | No |
| 31 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 4287 | 0.025 | -0.1114 | No |
| 32 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 4601 | 0.021 | -0.1328 | No |
| 33 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 4803 | 0.018 | -0.1460 | No |
| 34 | EIF5 | EIF5 Entrez, Source | eukaryotic translation initiation factor 5 | 4823 | 0.018 | -0.1454 | No |
| 35 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 5083 | 0.015 | -0.1634 | No |
| 36 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 5100 | 0.015 | -0.1629 | No |
| 37 | SFRS10 | SFRS10 Entrez, Source | splicing factor, arginine/serine-rich 10 (transformer 2 homolog, Drosophila) | 5320 | 0.012 | -0.1781 | No |
| 38 | BRD8 | BRD8 Entrez, Source | bromodomain containing 8 | 5377 | 0.012 | -0.1811 | No |
| 39 | DBI | DBI Entrez, Source | diazepam binding inhibitor (GABA receptor modulator, acyl-Coenzyme A binding protein) | 5459 | 0.011 | -0.1860 | No |
| 40 | TKT | TKT Entrez, Source | transketolase (Wernicke-Korsakoff syndrome) | 5509 | 0.010 | -0.1886 | No |
| 41 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 5560 | 0.009 | -0.1913 | No |
| 42 | PGK1 | PGK1 Entrez, Source | phosphoglycerate kinase 1 | 5631 | 0.009 | -0.1957 | No |
| 43 | EIF3S10 | EIF3S10 Entrez, Source | eukaryotic translation initiation factor 3, subunit 10 theta, 150/170kDa | 5672 | 0.008 | -0.1978 | No |
| 44 | ETF1 | ETF1 Entrez, Source | eukaryotic translation termination factor 1 | 5691 | 0.008 | -0.1982 | No |
| 45 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 5696 | 0.008 | -0.1976 | No |
| 46 | HNRPC | HNRPC Entrez, Source | heterogeneous nuclear ribonucleoprotein C (C1/C2) | 5697 | 0.008 | -0.1967 | No |
| 47 | TIMM17A | TIMM17A Entrez, Source | translocase of inner mitochondrial membrane 17 homolog A (yeast) | 5700 | 0.008 | -0.1960 | No |
| 48 | ACOT7 | ACOT7 Entrez, Source | acyl-CoA thioesterase 7 | 5733 | 0.008 | -0.1975 | No |
| 49 | EWSR1 | EWSR1 Entrez, Source | Ewing sarcoma breakpoint region 1 | 5987 | 0.005 | -0.2162 | No |
| 50 | TMEM97 | TMEM97 Entrez, Source | transmembrane protein 97 | 6035 | 0.004 | -0.2193 | No |
| 51 | NARS | NARS Entrez, Source | asparaginyl-tRNA synthetase | 6132 | 0.003 | -0.2263 | No |
| 52 | UFD1L | UFD1L Entrez, Source | ubiquitin fusion degradation 1 like (yeast) | 6141 | 0.003 | -0.2265 | No |
| 53 | ATP1B3 | ATP1B3 Entrez, Source | ATPase, Na+/K+ transporting, beta 3 polypeptide | 6238 | 0.002 | -0.2336 | No |
| 54 | SUMO1 | SUMO1 Entrez, Source | SMT3 suppressor of mif two 3 homolog 1 (S. cerevisiae) | 6246 | 0.002 | -0.2339 | No |
| 55 | AHCY | AHCY Entrez, Source | S-adenosylhomocysteine hydrolase | 6399 | 0.000 | -0.2454 | No |
| 56 | GPI | GPI Entrez, Source | glucose phosphate isomerase | 6497 | -0.001 | -0.2527 | No |
| 57 | HADH2 | HADH2 Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase, type II | 6668 | -0.003 | -0.2654 | No |
| 58 | PSMD4 | PSMD4 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 4 | 6720 | -0.003 | -0.2689 | No |
| 59 | COX7B | COX7B Entrez, Source | cytochrome c oxidase subunit VIIb | 6745 | -0.003 | -0.2703 | No |
| 60 | HPRT1 | HPRT1 Entrez, Source | hypoxanthine phosphoribosyltransferase 1 (Lesch-Nyhan syndrome) | 6770 | -0.004 | -0.2718 | No |
| 61 | MAPK1 | MAPK1 Entrez, Source | mitogen-activated protein kinase 1 | 6842 | -0.004 | -0.2767 | No |
| 62 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 6879 | -0.004 | -0.2789 | No |
| 63 | PPP1R2 | PPP1R2 Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 2 | 6926 | -0.005 | -0.2819 | No |
| 64 | GALNT1 | GALNT1 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 1 (GalNAc-T1) | 6935 | -0.005 | -0.2819 | No |
| 65 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 6975 | -0.005 | -0.2843 | No |
| 66 | SLA | SLA Entrez, Source | Src-like-adaptor | 6999 | -0.006 | -0.2854 | No |
| 67 | CPOX | CPOX Entrez, Source | coproporphyrinogen oxidase | 7027 | -0.006 | -0.2868 | No |
| 68 | CCNB1 | CCNB1 Entrez, Source | cyclin B1 | 7114 | -0.007 | -0.2926 | No |
| 69 | GARS | GARS Entrez, Source | glycyl-tRNA synthetase | 7116 | -0.007 | -0.2919 | No |
| 70 | UBE2D3 | UBE2D3 Entrez, Source | ubiquitin-conjugating enzyme E2D 3 (UBC4/5 homolog, yeast) | 7172 | -0.007 | -0.2952 | No |
| 71 | ITGB7 | ITGB7 Entrez, Source | integrin, beta 7 | 7193 | -0.007 | -0.2959 | No |
| 72 | CCT4 | CCT4 Entrez, Source | chaperonin containing TCP1, subunit 4 (delta) | 7230 | -0.008 | -0.2977 | No |
| 73 | EBNA1BP2 | EBNA1BP2 Entrez, Source | EBNA1 binding protein 2 | 7242 | -0.008 | -0.2976 | No |
| 74 | PSMA5 | PSMA5 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 5 | 7274 | -0.008 | -0.2990 | No |
| 75 | COX6C | COX6C Entrez, Source | cytochrome c oxidase subunit VIc | 7285 | -0.008 | -0.2988 | No |
| 76 | SUMO2 | SUMO2 Entrez, Source | SMT3 suppressor of mif two 3 homolog 2 (S. cerevisiae) | 7313 | -0.009 | -0.2999 | No |
| 77 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 7360 | -0.009 | -0.3024 | No |
| 78 | PPP4C | PPP4C Entrez, Source | protein phosphatase 4 (formerly X), catalytic subunit | 7406 | -0.010 | -0.3047 | No |
| 79 | ABCE1 | ABCE1 Entrez, Source | ATP-binding cassette, sub-family E (OABP), member 1 | 7480 | -0.010 | -0.3091 | No |
| 80 | U2AF2 | U2AF2 Entrez, Source | U2 small nuclear RNA auxiliary factor 2 | 7481 | -0.010 | -0.3079 | No |
| 81 | KIF22 | KIF22 Entrez, Source | kinesin family member 22 | 7498 | -0.010 | -0.3079 | No |
| 82 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 7500 | -0.010 | -0.3068 | No |
| 83 | DNAJC7 | DNAJC7 Entrez, Source | DnaJ (Hsp40) homolog, subfamily C, member 7 | 7554 | -0.011 | -0.3096 | No |
| 84 | ATIC | ATIC Entrez, Source | 5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase | 7625 | -0.012 | -0.3136 | No |
| 85 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 7635 | -0.012 | -0.3129 | No |
| 86 | EIF2S1 | EIF2S1 Entrez, Source | eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa | 7641 | -0.012 | -0.3119 | No |
| 87 | TCEB1 | TCEB1 Entrez, Source | transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) | 7679 | -0.012 | -0.3133 | No |
| 88 | GSPT1 | GSPT1 Entrez, Source | G1 to S phase transition 1 | 7697 | -0.012 | -0.3132 | No |
| 89 | TCP1 | TCP1 Entrez, Source | t-complex 1 | 7728 | -0.013 | -0.3140 | No |
| 90 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 7800 | -0.014 | -0.3179 | No |
| 91 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 7827 | -0.014 | -0.3183 | No |
| 92 | CCT6A | CCT6A Entrez, Source | chaperonin containing TCP1, subunit 6A (zeta 1) | 7874 | -0.014 | -0.3202 | No |
| 93 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 7897 | -0.014 | -0.3202 | No |
| 94 | OAT | OAT Entrez, Source | ornithine aminotransferase (gyrate atrophy) | 7916 | -0.015 | -0.3199 | No |
| 95 | HSPE1 | HSPE1 Entrez, Source | heat shock 10kDa protein 1 (chaperonin 10) | 8023 | -0.016 | -0.3261 | No |
| 96 | PSMA2 | PSMA2 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 2 | 8089 | -0.017 | -0.3292 | No |
| 97 | ATP5F1 | ATP5F1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit B1 | 8238 | -0.018 | -0.3384 | No |
| 98 | COPS5 | COPS5 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 5 (Arabidopsis) | 8255 | -0.018 | -0.3375 | No |
| 99 | ABCD3 | ABCD3 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 3 | 8286 | -0.019 | -0.3377 | No |
| 100 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 8303 | -0.019 | -0.3368 | No |
| 101 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 8374 | -0.019 | -0.3399 | No |
| 102 | PSMB2 | PSMB2 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 2 | 8376 | -0.019 | -0.3377 | No |
| 103 | VDAC1 | VDAC1 Entrez, Source | voltage-dependent anion channel 1 | 8435 | -0.020 | -0.3399 | No |
| 104 | CCT2 | CCT2 Entrez, Source | chaperonin containing TCP1, subunit 2 (beta) | 8544 | -0.021 | -0.3457 | No |
| 105 | TARS | TARS Entrez, Source | threonyl-tRNA synthetase | 8662 | -0.022 | -0.3520 | No |
| 106 | UQCRH | UQCRH Entrez, Source | ubiquinol-cytochrome c reductase hinge protein | 8723 | -0.023 | -0.3539 | No |
| 107 | SLC19A1 | SLC19A1 Entrez, Source | solute carrier family 19 (folate transporter), member 1 | 8805 | -0.024 | -0.3574 | No |
| 108 | PSMC5 | PSMC5 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 5 | 8846 | -0.024 | -0.3576 | No |
| 109 | APPBP1 | APPBP1 Entrez, Source | amyloid beta precursor protein binding protein 1 | 8865 | -0.025 | -0.3562 | No |
| 110 | CCT5 | CCT5 Entrez, Source | chaperonin containing TCP1, subunit 5 (epsilon) | 8949 | -0.025 | -0.3596 | No |
| 111 | PDAP1 | PDAP1 Entrez, Source | PDGFA associated protein 1 | 8993 | -0.026 | -0.3599 | No |
| 112 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 9143 | -0.027 | -0.3681 | No |
| 113 | OSTF1 | OSTF1 Entrez, Source | osteoclast stimulating factor 1 | 9146 | -0.027 | -0.3651 | No |
| 114 | NDUFV2 | NDUFV2 Entrez, Source | NADH dehydrogenase (ubiquinone) flavoprotein 2, 24kDa | 9181 | -0.028 | -0.3646 | No |
| 115 | DDX39 | DDX39 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 39 | 9306 | -0.029 | -0.3707 | No |
| 116 | MTHFD1 | MTHFD1 Entrez, Source | methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1, methenyltetrahydrofolate cyclohydrolase, formyltetrahydrofolate synthetase | 9334 | -0.029 | -0.3694 | No |
| 117 | SMC1A | SMC1A Entrez, Source | structural maintenance of chromosomes 1A | 9336 | -0.029 | -0.3661 | No |
| 118 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9337 | -0.029 | -0.3627 | No |
| 119 | C10ORF7 | C10ORF7 Entrez, Source | chromosome 10 open reading frame 7 | 9370 | -0.030 | -0.3618 | No |
| 120 | CCNE1 | CCNE1 Entrez, Source | cyclin E1 | 9404 | -0.030 | -0.3608 | No |
| 121 | DUT | DUT Entrez, Source | dUTP pyrophosphatase | 9409 | -0.030 | -0.3577 | No |
| 122 | GPIAP1 | GPIAP1 Entrez, Source | GPI-anchored membrane protein 1 | 9675 | -0.034 | -0.3740 | No |
| 123 | NDUFS1 | NDUFS1 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) | 9714 | -0.034 | -0.3730 | No |
| 124 | RNASE6 | RNASE6 Entrez, Source | ribonuclease, RNase A family, k6 | 9735 | -0.034 | -0.3706 | No |
| 125 | ATP5J | ATP5J Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit F6 | 9825 | -0.035 | -0.3733 | No |
| 126 | POLD2 | POLD2 Entrez, Source | polymerase (DNA directed), delta 2, regulatory subunit 50kDa | 9913 | -0.036 | -0.3758 | No |
| 127 | ASNS | ASNS Entrez, Source | asparagine synthetase | 9972 | -0.037 | -0.3760 | No |
| 128 | KLHDC3 | KLHDC3 Entrez, Source | kelch domain containing 3 | 10035 | -0.038 | -0.3765 | Yes |
| 129 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 10041 | -0.038 | -0.3726 | Yes |
| 130 | ANXA6 | ANXA6 Entrez, Source | annexin A6 | 10067 | -0.038 | -0.3701 | Yes |
| 131 | TDG | TDG Entrez, Source | thymine-DNA glycosylase | 10070 | -0.038 | -0.3660 | Yes |
| 132 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 10085 | -0.038 | -0.3627 | Yes |
| 133 | ATP5O | ATP5O Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, O subunit (oligomycin sensitivity conferring protein) | 10128 | -0.039 | -0.3615 | Yes |
| 134 | PSMB8 | PSMB8 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 8 (large multifunctional peptidase 7) | 10133 | -0.039 | -0.3573 | Yes |
| 135 | ASNA1 | ASNA1 Entrez, Source | arsA arsenite transporter, ATP-binding, homolog 1 (bacterial) | 10147 | -0.039 | -0.3539 | Yes |
| 136 | PTPN1 | PTPN1 Entrez, Source | protein tyrosine phosphatase, non-receptor type 1 | 10210 | -0.040 | -0.3541 | Yes |
| 137 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 10273 | -0.040 | -0.3542 | Yes |
| 138 | MRPL23 | MRPL23 Entrez, Source | mitochondrial ribosomal protein L23 | 10323 | -0.041 | -0.3533 | Yes |
| 139 | MAPKAPK3 | MAPKAPK3 Entrez, Source | mitogen-activated protein kinase-activated protein kinase 3 | 10382 | -0.042 | -0.3530 | Yes |
| 140 | YWHAH | YWHAH Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide | 10384 | -0.042 | -0.3483 | Yes |
| 141 | UQCRC2 | UQCRC2 Entrez, Source | ubiquinol-cytochrome c reductase core protein II | 10520 | -0.044 | -0.3536 | Yes |
| 142 | PSMA1 | PSMA1 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 1 | 10534 | -0.044 | -0.3496 | Yes |
| 143 | GNB1 | GNB1 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 1 | 10542 | -0.044 | -0.3451 | Yes |
| 144 | MRPL12 | MRPL12 Entrez, Source | mitochondrial ribosomal protein L12 | 10545 | -0.044 | -0.3402 | Yes |
| 145 | CASP6 | CASP6 Entrez, Source | caspase 6, apoptosis-related cysteine peptidase | 10649 | -0.045 | -0.3428 | Yes |
| 146 | CLCN3 | CLCN3 Entrez, Source | chloride channel 3 | 10689 | -0.046 | -0.3406 | Yes |
| 147 | COASY | COASY Entrez, Source | Coenzyme A synthase | 10703 | -0.046 | -0.3363 | Yes |
| 148 | IFI30 | IFI30 Entrez, Source | interferon, gamma-inducible protein 30 | 10707 | -0.046 | -0.3312 | Yes |
| 149 | NME1 | NME1 Entrez, Source | non-metastatic cells 1, protein (NM23A) expressed in | 10783 | -0.048 | -0.3315 | Yes |
| 150 | GIT2 | GIT2 Entrez, Source | G protein-coupled receptor kinase interactor 2 | 10815 | -0.048 | -0.3283 | Yes |
| 151 | BLMH | BLMH Entrez, Source | bleomycin hydrolase | 10846 | -0.049 | -0.3250 | Yes |
| 152 | PDLIM1 | PDLIM1 Entrez, Source | PDZ and LIM domain 1 (elfin) | 10847 | -0.049 | -0.3195 | Yes |
| 153 | CD52 | CD52 Entrez, Source | CD52 molecule | 10867 | -0.049 | -0.3153 | Yes |
| 154 | PHB | PHB Entrez, Source | prohibitin | 10993 | -0.051 | -0.3190 | Yes |
| 155 | ITGB1 | ITGB1 Entrez, Source | integrin, beta 1 (fibronectin receptor, beta polypeptide, antigen CD29 includes MDF2, MSK12) | 11016 | -0.051 | -0.3149 | Yes |
| 156 | PSME4 | PSME4 Entrez, Source | proteasome (prosome, macropain) activator subunit 4 | 11061 | -0.052 | -0.3123 | Yes |
| 157 | TRAP1 | TRAP1 Entrez, Source | TNF receptor-associated protein 1 | 11230 | -0.056 | -0.3187 | Yes |
| 158 | USP14 | USP14 Entrez, Source | ubiquitin specific peptidase 14 (tRNA-guanine transglycosylase) | 11235 | -0.056 | -0.3127 | Yes |
| 159 | RECQL | RECQL Entrez, Source | RecQ protein-like (DNA helicase Q1-like) | 11262 | -0.056 | -0.3082 | Yes |
| 160 | TAX1BP3 | TAX1BP3 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 3 | 11278 | -0.056 | -0.3029 | Yes |
| 161 | PAICS | PAICS Entrez, Source | phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoimidazole succinocarboxamide synthetase | 11374 | -0.058 | -0.3035 | Yes |
| 162 | PSME2 | PSME2 Entrez, Source | proteasome (prosome, macropain) activator subunit 2 (PA28 beta) | 11462 | -0.060 | -0.3033 | Yes |
| 163 | LBR | LBR Entrez, Source | lamin B receptor | 11583 | -0.062 | -0.3054 | Yes |
| 164 | CDC27 | CDC27 Entrez, Source | cell division cycle 27 | 11606 | -0.063 | -0.2999 | Yes |
| 165 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 11618 | -0.063 | -0.2936 | Yes |
| 166 | S100A10 | S100A10 Entrez, Source | S100 calcium binding protein A10 | 11713 | -0.065 | -0.2933 | Yes |
| 167 | ANP32A | ANP32A Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member A | 11774 | -0.067 | -0.2903 | Yes |
| 168 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 11987 | -0.072 | -0.2982 | Yes |
| 169 | MRPL19 | MRPL19 Entrez, Source | mitochondrial ribosomal protein L19 | 12043 | -0.074 | -0.2939 | Yes |
| 170 | ADD1 | ADD1 Entrez, Source | adducin 1 (alpha) | 12055 | -0.074 | -0.2862 | Yes |
| 171 | KIFC1 | KIFC1 Entrez, Source | kinesin family member C1 | 12059 | -0.075 | -0.2780 | Yes |
| 172 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 12078 | -0.075 | -0.2708 | Yes |
| 173 | GGH | GGH Entrez, Source | gamma-glutamyl hydrolase (conjugase, folylpolygammaglutamyl hydrolase) | 12131 | -0.077 | -0.2659 | Yes |
| 174 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 12283 | -0.082 | -0.2681 | Yes |
| 175 | GART | GART Entrez, Source | phosphoribosylglycinamide formyltransferase, phosphoribosylglycinamide synthetase, phosphoribosylaminoimidazole synthetase | 12346 | -0.085 | -0.2631 | Yes |
| 176 | PAK2 | PAK2 Entrez, Source | p21 (CDKN1A)-activated kinase 2 | 12379 | -0.086 | -0.2557 | Yes |
| 177 | TUBG1 | TUBG1 Entrez, Source | tubulin, gamma 1 | 12486 | -0.090 | -0.2535 | Yes |
| 178 | MYO5A | MYO5A Entrez, Source | myosin VA (heavy chain 12, myoxin) | 12504 | -0.092 | -0.2443 | Yes |
| 179 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 12511 | -0.092 | -0.2343 | Yes |
| 180 | PSMC3IP | PSMC3IP Entrez, Source | PSMC3 interacting protein | 12571 | -0.095 | -0.2279 | Yes |
| 181 | ORC1L | ORC1L Entrez, Source | origin recognition complex, subunit 1-like (yeast) | 12603 | -0.097 | -0.2192 | Yes |
| 182 | RB1 | RB1 Entrez, Source | retinoblastoma 1 (including osteosarcoma) | 12607 | -0.097 | -0.2084 | Yes |
| 183 | HSPA4 | HSPA4 Entrez, Source | heat shock 70kDa protein 4 | 12729 | -0.105 | -0.2056 | Yes |
| 184 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 12735 | -0.105 | -0.1940 | Yes |
| 185 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 12799 | -0.110 | -0.1862 | Yes |
| 186 | TMPO | TMPO Entrez, Source | thymopoietin | 12812 | -0.111 | -0.1744 | Yes |
| 187 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 12841 | -0.114 | -0.1636 | Yes |
| 188 | MVP | MVP Entrez, Source | major vault protein | 12888 | -0.120 | -0.1534 | Yes |
| 189 | TMSB10 | TMSB10 Entrez, Source | thymosin, beta 10 | 12891 | -0.120 | -0.1399 | Yes |
| 190 | RFC3 | RFC3 Entrez, Source | replication factor C (activator 1) 3, 38kDa | 12915 | -0.123 | -0.1276 | Yes |
| 191 | KIF2A | KIF2A Entrez, Source | kinesin heavy chain member 2A | 12962 | -0.127 | -0.1166 | Yes |
| 192 | TRIP13 | TRIP13 Entrez, Source | thyroid hormone receptor interactor 13 | 13078 | -0.144 | -0.1089 | Yes |
| 193 | RAB27A | RAB27A Entrez, Source | RAB27A, member RAS oncogene family | 13087 | -0.147 | -0.0928 | Yes |
| 194 | PRPS2 | PRPS2 Entrez, Source | phosphoribosyl pyrophosphate synthetase 2 | 13107 | -0.151 | -0.0770 | Yes |
| 195 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13194 | -0.185 | -0.0624 | Yes |
| 196 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 13232 | -0.213 | -0.0409 | Yes |
| 197 | MSN | MSN Entrez, Source | moesin | 13233 | -0.213 | -0.0166 | Yes |
| 198 | CLIC1 | CLIC1 Entrez, Source | chloride intracellular channel 1 | 13242 | -0.218 | 0.0076 | Yes |