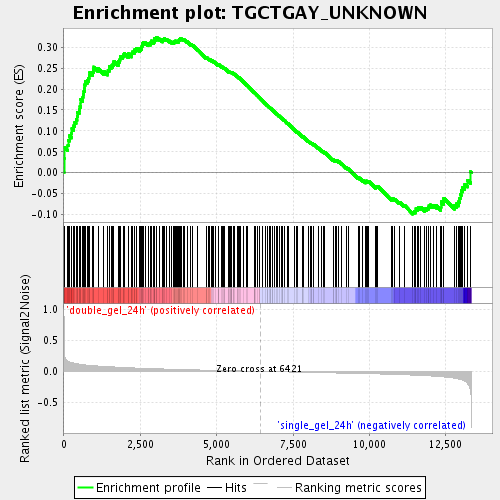
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | TGCTGAY_UNKNOWN |
| Enrichment Score (ES) | 0.32392007 |
| Normalized Enrichment Score (NES) | 1.4963647 |
| Nominal p-value | 0.0019762847 |
| FDR q-value | 0.27309686 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CSAD | CSAD Entrez, Source | cysteine sulfinic acid decarboxylase | 4 | 0.412 | 0.0340 | Yes |
| 2 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 11 | 0.331 | 0.0611 | Yes |
| 3 | GPM6B | GPM6B Entrez, Source | glycoprotein M6B | 130 | 0.167 | 0.0661 | Yes |
| 4 | PRKCQ | PRKCQ Entrez, Source | protein kinase C, theta | 147 | 0.160 | 0.0781 | Yes |
| 5 | CNTN6 | CNTN6 Entrez, Source | contactin 6 | 181 | 0.152 | 0.0882 | Yes |
| 6 | WNT7A | WNT7A Entrez, Source | wingless-type MMTV integration site family, member 7A | 251 | 0.141 | 0.0947 | Yes |
| 7 | TTN | TTN Entrez, Source | titin | 262 | 0.140 | 0.1056 | Yes |
| 8 | CACNG3 | CACNG3 Entrez, Source | calcium channel, voltage-dependent, gamma subunit 3 | 325 | 0.132 | 0.1119 | Yes |
| 9 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 356 | 0.129 | 0.1203 | Yes |
| 10 | CYP17A1 | CYP17A1 Entrez, Source | cytochrome P450, family 17, subfamily A, polypeptide 1 | 397 | 0.126 | 0.1277 | Yes |
| 11 | ENPP2 | ENPP2 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 2 (autotaxin) | 429 | 0.123 | 0.1356 | Yes |
| 12 | CKM | CKM Entrez, Source | creatine kinase, muscle | 445 | 0.122 | 0.1446 | Yes |
| 13 | SLCO2B1 | SLCO2B1 Entrez, Source | solute carrier organic anion transporter family, member 2B1 | 498 | 0.118 | 0.1505 | Yes |
| 14 | KCNK12 | KCNK12 Entrez, Source | potassium channel, subfamily K, member 12 | 524 | 0.115 | 0.1582 | Yes |
| 15 | BMP2 | BMP2 Entrez, Source | bone morphogenetic protein 2 | 547 | 0.114 | 0.1659 | Yes |
| 16 | IRS1 | IRS1 Entrez, Source | insulin receptor substrate 1 | 549 | 0.114 | 0.1753 | Yes |
| 17 | HSPA5 | HSPA5 Entrez, Source | heat shock 70kDa protein 5 (glucose-regulated protein, 78kDa) | 618 | 0.109 | 0.1792 | Yes |
| 18 | NPHS1 | NPHS1 Entrez, Source | nephrosis 1, congenital, Finnish type (nephrin) | 626 | 0.109 | 0.1877 | Yes |
| 19 | ACCN2 | ACCN2 Entrez, Source | amiloride-sensitive cation channel 2, neuronal | 653 | 0.107 | 0.1946 | Yes |
| 20 | SLC16A6 | SLC16A6 Entrez, Source | solute carrier family 16, member 6 (monocarboxylic acid transporter 7) | 674 | 0.105 | 0.2019 | Yes |
| 21 | TUB | TUB Entrez, Source | tubby homolog (mouse) | 684 | 0.105 | 0.2100 | Yes |
| 22 | CACNG2 | CACNG2 Entrez, Source | calcium channel, voltage-dependent, gamma subunit 2 | 693 | 0.104 | 0.2180 | Yes |
| 23 | SLC6A2 | SLC6A2 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, noradrenalin), member 2 | 760 | 0.101 | 0.2215 | Yes |
| 24 | SERPIND1 | SERPIND1 Entrez, Source | serpin peptidase inhibitor, clade D (heparin cofactor), member 1 | 812 | 0.099 | 0.2258 | Yes |
| 25 | IL4 | IL4 Entrez, Source | interleukin 4 | 824 | 0.098 | 0.2332 | Yes |
| 26 | SYT5 | SYT5 Entrez, Source | synaptotagmin V | 847 | 0.097 | 0.2396 | Yes |
| 27 | LRRC4 | LRRC4 Entrez, Source | leucine rich repeat containing 4 | 942 | 0.094 | 0.2402 | Yes |
| 28 | DRP2 | DRP2 Entrez, Source | dystrophin related protein 2 | 964 | 0.093 | 0.2464 | Yes |
| 29 | MSX2 | MSX2 Entrez, Source | msh homeobox homolog 2 (Drosophila) | 981 | 0.092 | 0.2528 | Yes |
| 30 | COL1A2 | COL1A2 Entrez, Source | collagen, type I, alpha 2 | 1122 | 0.086 | 0.2493 | Yes |
| 31 | SNX15 | SNX15 Entrez, Source | sorting nexin 15 | 1311 | 0.080 | 0.2417 | Yes |
| 32 | MYOC | MYOC Entrez, Source | myocilin, trabecular meshwork inducible glucocorticoid response | 1418 | 0.077 | 0.2400 | Yes |
| 33 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 1441 | 0.077 | 0.2447 | Yes |
| 34 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 1479 | 0.075 | 0.2481 | Yes |
| 35 | RTN4R | RTN4R Entrez, Source | reticulon 4 receptor | 1487 | 0.075 | 0.2538 | Yes |
| 36 | ARMCX2 | ARMCX2 Entrez, Source | armadillo repeat containing, X-linked 2 | 1547 | 0.073 | 0.2555 | Yes |
| 37 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 1596 | 0.073 | 0.2578 | Yes |
| 38 | AXIN2 | AXIN2 Entrez, Source | axin 2 (conductin, axil) | 1617 | 0.072 | 0.2623 | Yes |
| 39 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 1635 | 0.072 | 0.2670 | Yes |
| 40 | FGF7 | FGF7 Entrez, Source | fibroblast growth factor 7 (keratinocyte growth factor) | 1778 | 0.068 | 0.2618 | Yes |
| 41 | WNT2 | WNT2 Entrez, Source | wingless-type MMTV integration site family member 2 | 1789 | 0.068 | 0.2667 | Yes |
| 42 | FILIP1 | FILIP1 Entrez, Source | filamin A interacting protein 1 | 1823 | 0.067 | 0.2697 | Yes |
| 43 | GSTA2 | GSTA2 Entrez, Source | glutathione S-transferase A2 | 1836 | 0.067 | 0.2743 | Yes |
| 44 | PDE1C | PDE1C Entrez, Source | phosphodiesterase 1C, calmodulin-dependent 70kDa | 1849 | 0.066 | 0.2789 | Yes |
| 45 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 1933 | 0.064 | 0.2779 | Yes |
| 46 | AMELX | AMELX Entrez, Source | amelogenin (amelogenesis imperfecta 1, X-linked) | 1951 | 0.064 | 0.2819 | Yes |
| 47 | FGF6 | FGF6 Entrez, Source | fibroblast growth factor 6 | 1981 | 0.063 | 0.2850 | Yes |
| 48 | GALNT13 | GALNT13 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 13 (GalNAc-T13) | 2108 | 0.060 | 0.2803 | Yes |
| 49 | SLC26A3 | SLC26A3 Entrez, Source | solute carrier family 26, member 3 | 2119 | 0.060 | 0.2845 | Yes |
| 50 | SNCB | SNCB Entrez, Source | synuclein, beta | 2223 | 0.057 | 0.2814 | Yes |
| 51 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 2224 | 0.057 | 0.2861 | Yes |
| 52 | ITPR3 | ITPR3 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 3 | 2251 | 0.056 | 0.2888 | Yes |
| 53 | USP2 | USP2 Entrez, Source | ubiquitin specific peptidase 2 | 2300 | 0.055 | 0.2898 | Yes |
| 54 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 2310 | 0.055 | 0.2937 | Yes |
| 55 | OPN1SW | OPN1SW Entrez, Source | opsin 1 (cone pigments), short-wave-sensitive (color blindness, tritan) | 2360 | 0.054 | 0.2945 | Yes |
| 56 | CRYBA1 | CRYBA1 Entrez, Source | crystallin, beta A1 | 2380 | 0.054 | 0.2975 | Yes |
| 57 | RNF12 | RNF12 Entrez, Source | ring finger protein 12 | 2482 | 0.052 | 0.2942 | Yes |
| 58 | MAG | MAG Entrez, Source | myelin associated glycoprotein | 2504 | 0.051 | 0.2968 | Yes |
| 59 | CACNA1E | CACNA1E Entrez, Source | calcium channel, voltage-dependent, alpha 1E subunit | 2545 | 0.051 | 0.2980 | Yes |
| 60 | POU3F4 | POU3F4 Entrez, Source | POU domain, class 3, transcription factor 4 | 2551 | 0.051 | 0.3018 | Yes |
| 61 | KCNH3 | KCNH3 Entrez, Source | potassium voltage-gated channel, subfamily H (eag-related), member 3 | 2561 | 0.050 | 0.3054 | Yes |
| 62 | HSPB7 | HSPB7 Entrez, Source | heat shock 27kDa protein family, member 7 (cardiovascular) | 2572 | 0.050 | 0.3088 | Yes |
| 63 | STMN4 | STMN4 Entrez, Source | stathmin-like 4 | 2594 | 0.050 | 0.3113 | Yes |
| 64 | AMY2A | AMY2A Entrez, Source | amylase, alpha 2A; pancreatic | 2654 | 0.049 | 0.3109 | Yes |
| 65 | PTP4A1 | PTP4A1 Entrez, Source | protein tyrosine phosphatase type IVA, member 1 | 2756 | 0.047 | 0.3071 | Yes |
| 66 | LRRN1 | LRRN1 Entrez, Source | leucine rich repeat neuronal 1 | 2774 | 0.047 | 0.3097 | Yes |
| 67 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 2834 | 0.046 | 0.3090 | Yes |
| 68 | TCF4 | TCF4 Entrez, Source | transcription factor 4 | 2845 | 0.045 | 0.3120 | Yes |
| 69 | GALNT10 | GALNT10 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 10 (GalNAc-T10) | 2854 | 0.045 | 0.3151 | Yes |
| 70 | EIF4A2 | EIF4A2 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 2 | 2944 | 0.044 | 0.3120 | Yes |
| 71 | RHOBTB2 | RHOBTB2 Entrez, Source | Rho-related BTB domain containing 2 | 2953 | 0.044 | 0.3151 | Yes |
| 72 | JPH2 | JPH2 Entrez, Source | junctophilin 2 | 2958 | 0.044 | 0.3184 | Yes |
| 73 | SNAP25 | SNAP25 Entrez, Source | synaptosomal-associated protein, 25kDa | 2966 | 0.044 | 0.3215 | Yes |
| 74 | CDH10 | CDH10 Entrez, Source | cadherin 10, type 2 (T2-cadherin) | 3027 | 0.043 | 0.3205 | Yes |
| 75 | ARFIP1 | ARFIP1 Entrez, Source | ADP-ribosylation factor interacting protein 1 (arfaptin 1) | 3029 | 0.043 | 0.3239 | Yes |
| 76 | TTR | TTR Entrez, Source | transthyretin (prealbumin, amyloidosis type I) | 3128 | 0.041 | 0.3199 | No |
| 77 | NRP2 | NRP2 Entrez, Source | neuropilin 2 | 3225 | 0.040 | 0.3159 | No |
| 78 | SHANK1 | SHANK1 Entrez, Source | SH3 and multiple ankyrin repeat domains 1 | 3234 | 0.040 | 0.3185 | No |
| 79 | TITF1 | TITF1 Entrez, Source | thyroid transcription factor 1 | 3273 | 0.039 | 0.3189 | No |
| 80 | PFKFB3 | PFKFB3 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3 | 3285 | 0.039 | 0.3213 | No |
| 81 | TEAD1 | TEAD1 Entrez, Source | TEA domain family member 1 (SV40 transcriptional enhancer factor) | 3364 | 0.037 | 0.3184 | No |
| 82 | OTX2 | OTX2 Entrez, Source | orthodenticle homolog 2 (Drosophila) | 3447 | 0.036 | 0.3152 | No |
| 83 | SIX1 | SIX1 Entrez, Source | sine oculis homeobox homolog 1 (Drosophila) | 3519 | 0.035 | 0.3127 | No |
| 84 | DTNB | DTNB Entrez, Source | dystrobrevin, beta | 3578 | 0.034 | 0.3111 | No |
| 85 | SEZ6 | SEZ6 Entrez, Source | seizure related 6 homolog (mouse) | 3603 | 0.034 | 0.3121 | No |
| 86 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 3623 | 0.033 | 0.3134 | No |
| 87 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 3637 | 0.033 | 0.3152 | No |
| 88 | HTR2B | HTR2B Entrez, Source | 5-hydroxytryptamine (serotonin) receptor 2B | 3684 | 0.033 | 0.3144 | No |
| 89 | GPR85 | GPR85 Entrez, Source | G protein-coupled receptor 85 | 3730 | 0.032 | 0.3136 | No |
| 90 | RARB | RARB Entrez, Source | retinoic acid receptor, beta | 3752 | 0.032 | 0.3147 | No |
| 91 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 3760 | 0.032 | 0.3168 | No |
| 92 | PADI4 | PADI4 Entrez, Source | peptidyl arginine deiminase, type IV | 3776 | 0.031 | 0.3183 | No |
| 93 | YWHAG | YWHAG Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide | 3798 | 0.031 | 0.3193 | No |
| 94 | KCND3 | KCND3 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 3 | 3813 | 0.031 | 0.3208 | No |
| 95 | KCNA1 | KCNA1 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 1 (episodic ataxia with myokymia) | 3853 | 0.030 | 0.3203 | No |
| 96 | PTPRN | PTPRN Entrez, Source | protein tyrosine phosphatase, receptor type, N | 3909 | 0.030 | 0.3186 | No |
| 97 | CEACAM1 | CEACAM1 Entrez, Source | carcinoembryonic antigen-related cell adhesion molecule 1 (biliary glycoprotein) | 3951 | 0.029 | 0.3179 | No |
| 98 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 4037 | 0.028 | 0.3137 | No |
| 99 | SEMA4B | SEMA4B Entrez, Source | sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4B | 4154 | 0.026 | 0.3070 | No |
| 100 | GRIN2B | GRIN2B Entrez, Source | glutamate receptor, ionotropic, N-methyl D-aspartate 2B | 4206 | 0.026 | 0.3053 | No |
| 101 | GBA2 | GBA2 Entrez, Source | glucosidase, beta (bile acid) 2 | 4384 | 0.023 | 0.2937 | No |
| 102 | TBXAS1 | TBXAS1 Entrez, Source | thromboxane A synthase 1 (platelet, cytochrome P450, family 5, subfamily A) | 4650 | 0.020 | 0.2752 | No |
| 103 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 4673 | 0.020 | 0.2752 | No |
| 104 | DNMT3A | DNMT3A Entrez, Source | DNA (cytosine-5-)-methyltransferase 3 alpha | 4742 | 0.019 | 0.2716 | No |
| 105 | DDIT4L | DDIT4L Entrez, Source | DNA-damage-inducible transcript 4-like | 4776 | 0.019 | 0.2706 | No |
| 106 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 4824 | 0.018 | 0.2686 | No |
| 107 | TCF21 | TCF21 Entrez, Source | transcription factor 21 | 4856 | 0.018 | 0.2677 | No |
| 108 | GJA1 | GJA1 Entrez, Source | gap junction protein, alpha 1, 43kDa (connexin 43) | 4902 | 0.017 | 0.2657 | No |
| 109 | RAB30 | RAB30 Entrez, Source | RAB30, member RAS oncogene family | 4956 | 0.016 | 0.2630 | No |
| 110 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 5047 | 0.015 | 0.2574 | No |
| 111 | SLIT3 | SLIT3 Entrez, Source | slit homolog 3 (Drosophila) | 5054 | 0.015 | 0.2583 | No |
| 112 | GABRA6 | GABRA6 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 6 | 5056 | 0.015 | 0.2595 | No |
| 113 | PPP1R10 | PPP1R10 Entrez, Source | protein phosphatase 1, regulatory subunit 10 | 5145 | 0.014 | 0.2539 | No |
| 114 | ADARB2 | ADARB2 Entrez, Source | adenosine deaminase, RNA-specific, B2 (RED2 homolog rat) | 5173 | 0.014 | 0.2531 | No |
| 115 | ADNP | ADNP Entrez, Source | activity-dependent neuroprotector | 5220 | 0.013 | 0.2507 | No |
| 116 | SLC22A17 | SLC22A17 Entrez, Source | solute carrier family 22 (organic cation transporter), member 17 | 5240 | 0.013 | 0.2503 | No |
| 117 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 5269 | 0.013 | 0.2493 | No |
| 118 | APOH | APOH Entrez, Source | apolipoprotein H (beta-2-glycoprotein I) | 5386 | 0.011 | 0.2414 | No |
| 119 | NAPA | NAPA Entrez, Source | N-ethylmaleimide-sensitive factor attachment protein, alpha | 5398 | 0.011 | 0.2415 | No |
| 120 | PLAU | PLAU Entrez, Source | plasminogen activator, urokinase | 5422 | 0.011 | 0.2406 | No |
| 121 | MYH8 | MYH8 Entrez, Source | myosin, heavy chain 8, skeletal muscle, perinatal | 5444 | 0.011 | 0.2400 | No |
| 122 | SLCO1C1 | SLCO1C1 Entrez, Source | solute carrier organic anion transporter family, member 1C1 | 5456 | 0.011 | 0.2400 | No |
| 123 | P2RY6 | P2RY6 Entrez, Source | pyrimidinergic receptor P2Y, G-protein coupled, 6 | 5479 | 0.010 | 0.2392 | No |
| 124 | MEPE | MEPE Entrez, Source | matrix, extracellular phosphoglycoprotein with ASARM motif (bone) | 5482 | 0.010 | 0.2399 | No |
| 125 | ALDOA | ALDOA Entrez, Source | aldolase A, fructose-bisphosphate | 5499 | 0.010 | 0.2395 | No |
| 126 | RRAGA | RRAGA Entrez, Source | Ras-related GTP binding A | 5543 | 0.010 | 0.2371 | No |
| 127 | RAB1A | RAB1A Entrez, Source | RAB1A, member RAS oncogene family | 5561 | 0.009 | 0.2366 | No |
| 128 | RYR1 | RYR1 Entrez, Source | ryanodine receptor 1 (skeletal) | 5589 | 0.009 | 0.2353 | No |
| 129 | KCNIP4 | KCNIP4 Entrez, Source | Kv channel interacting protein 4 | 5671 | 0.008 | 0.2298 | No |
| 130 | ACR | ACR Entrez, Source | acrosin | 5683 | 0.008 | 0.2296 | No |
| 131 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 5728 | 0.008 | 0.2269 | No |
| 132 | SPOP | SPOP Entrez, Source | speckle-type POZ protein | 5734 | 0.008 | 0.2271 | No |
| 133 | RARA | RARA Entrez, Source | retinoic acid receptor, alpha | 5779 | 0.007 | 0.2244 | No |
| 134 | SLC39A13 | SLC39A13 Entrez, Source | solute carrier family 39 (zinc transporter), member 13 | 5883 | 0.006 | 0.2170 | No |
| 135 | EWSR1 | EWSR1 Entrez, Source | Ewing sarcoma breakpoint region 1 | 5987 | 0.005 | 0.2095 | No |
| 136 | PREB | PREB Entrez, Source | prolactin regulatory element binding | 5999 | 0.005 | 0.2091 | No |
| 137 | FXYD6 | FXYD6 Entrez, Source | FXYD domain containing ion transport regulator 6 | 6228 | 0.002 | 0.1918 | No |
| 138 | NAV2 | NAV2 Entrez, Source | neuron navigator 2 | 6229 | 0.002 | 0.1920 | No |
| 139 | MAP3K7IP2 | MAP3K7IP2 Entrez, Source | mitogen-activated protein kinase kinase kinase 7 interacting protein 2 | 6244 | 0.002 | 0.1911 | No |
| 140 | P2RXL1 | P2RXL1 Entrez, Source | purinergic receptor P2X-like 1, orphan receptor | 6278 | 0.002 | 0.1887 | No |
| 141 | MAPK8IP2 | MAPK8IP2 Entrez, Source | mitogen-activated protein kinase 8 interacting protein 2 | 6319 | 0.001 | 0.1857 | No |
| 142 | NUMBL | NUMBL Entrez, Source | numb homolog (Drosophila)-like | 6411 | 0.000 | 0.1788 | No |
| 143 | SYT4 | SYT4 Entrez, Source | synaptotagmin IV | 6412 | 0.000 | 0.1788 | No |
| 144 | INPP4B | INPP4B Entrez, Source | inositol polyphosphate-4-phosphatase, type II, 105kDa | 6486 | -0.001 | 0.1733 | No |
| 145 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 6595 | -0.002 | 0.1652 | No |
| 146 | FGF9 | FGF9 Entrez, Source | fibroblast growth factor 9 (glia-activating factor) | 6646 | -0.002 | 0.1616 | No |
| 147 | CACNA1B | CACNA1B Entrez, Source | calcium channel, voltage-dependent, L type, alpha 1B subunit | 6647 | -0.002 | 0.1618 | No |
| 148 | WBP4 | WBP4 Entrez, Source | WW domain binding protein 4 (formin binding protein 21) | 6729 | -0.003 | 0.1559 | No |
| 149 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 6754 | -0.003 | 0.1544 | No |
| 150 | ABCF3 | ABCF3 Entrez, Source | ATP-binding cassette, sub-family F (GCN20), member 3 | 6773 | -0.004 | 0.1533 | No |
| 151 | UBE1C | UBE1C Entrez, Source | ubiquitin-activating enzyme E1C (UBA3 homolog, yeast) | 6831 | -0.004 | 0.1493 | No |
| 152 | TRIM2 | TRIM2 Entrez, Source | tripartite motif-containing 2 | 6833 | -0.004 | 0.1495 | No |
| 153 | PITX2 | PITX2 Entrez, Source | paired-like homeodomain transcription factor 2 | 6901 | -0.005 | 0.1448 | No |
| 154 | NEXN | NEXN Entrez, Source | nexilin (F actin binding protein) | 6958 | -0.005 | 0.1410 | No |
| 155 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 6990 | -0.006 | 0.1391 | No |
| 156 | MRPS18B | MRPS18B Entrez, Source | mitochondrial ribosomal protein S18B | 7041 | -0.006 | 0.1358 | No |
| 157 | ARL2BP | ARL2BP Entrez, Source | ADP-ribosylation factor-like 2 binding protein | 7112 | -0.007 | 0.1310 | No |
| 158 | ANGPT1 | ANGPT1 Entrez, Source | angiopoietin 1 | 7146 | -0.007 | 0.1290 | No |
| 159 | AVP | AVP Entrez, Source | arginine vasopressin (neurophysin II, antidiuretic hormone, diabetes insipidus, neurohypophyseal) | 7157 | -0.007 | 0.1289 | No |
| 160 | CHRNB4 | CHRNB4 Entrez, Source | cholinergic receptor, nicotinic, beta 4 | 7214 | -0.008 | 0.1253 | No |
| 161 | SNX10 | SNX10 Entrez, Source | sorting nexin 10 | 7325 | -0.009 | 0.1176 | No |
| 162 | UCN2 | UCN2 Entrez, Source | urocortin 2 | 7333 | -0.009 | 0.1178 | No |
| 163 | GNG10 | GNG10 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 10 | 7545 | -0.011 | 0.1026 | No |
| 164 | ARIH1 | ARIH1 Entrez, Source | ariadne homolog, ubiquitin-conjugating enzyme E2 binding protein, 1 (Drosophila) | 7627 | -0.012 | 0.0974 | No |
| 165 | SND1 | SND1 Entrez, Source | - | 7653 | -0.012 | 0.0965 | No |
| 166 | RPL28 | RPL28 Entrez, Source | ribosomal protein L28 | 7793 | -0.013 | 0.0870 | No |
| 167 | ARF1 | ARF1 Entrez, Source | ADP-ribosylation factor 1 | 7820 | -0.014 | 0.0862 | No |
| 168 | SON | SON Entrez, Source | SON DNA binding protein | 7838 | -0.014 | 0.0861 | No |
| 169 | PAQR4 | PAQR4 Entrez, Source | progestin and adipoQ receptor family member IV | 7999 | -0.016 | 0.0752 | No |
| 170 | PCSK1 | PCSK1 Entrez, Source | proprotein convertase subtilisin/kexin type 1 | 8055 | -0.016 | 0.0723 | No |
| 171 | DIO3 | DIO3 Entrez, Source | deiodinase, iodothyronine, type III | 8094 | -0.017 | 0.0708 | No |
| 172 | CAMK2A | CAMK2A Entrez, Source | calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha | 8111 | -0.017 | 0.0710 | No |
| 173 | THRAP3 | THRAP3 Entrez, Source | thyroid hormone receptor associated protein 3 | 8178 | -0.017 | 0.0674 | No |
| 174 | PPP2CA | PPP2CA Entrez, Source | protein phosphatase 2 (formerly 2A), catalytic subunit, alpha isoform | 8183 | -0.017 | 0.0685 | No |
| 175 | FKBP1A | FKBP1A Entrez, Source | FK506 binding protein 1A, 12kDa | 8328 | -0.019 | 0.0592 | No |
| 176 | OMG | OMG Entrez, Source | oligodendrocyte myelin glycoprotein | 8432 | -0.020 | 0.0530 | No |
| 177 | DGKZ | DGKZ Entrez, Source | diacylglycerol kinase, zeta 104kDa | 8481 | -0.020 | 0.0510 | No |
| 178 | HHEX | HHEX Entrez, Source | homeobox, hematopoietically expressed | 8540 | -0.021 | 0.0483 | No |
| 179 | BUB3 | BUB3 Entrez, Source | BUB3 budding uninhibited by benzimidazoles 3 homolog (yeast) | 8807 | -0.024 | 0.0301 | No |
| 180 | BLNK | BLNK Entrez, Source | B-cell linker | 8820 | -0.024 | 0.0311 | No |
| 181 | CMAS | CMAS Entrez, Source | cytidine monophosphate N-acetylneuraminic acid synthetase | 8876 | -0.025 | 0.0290 | No |
| 182 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 8916 | -0.025 | 0.0281 | No |
| 183 | MRPL24 | MRPL24 Entrez, Source | mitochondrial ribosomal protein L24 | 8919 | -0.025 | 0.0301 | No |
| 184 | AIF1 | AIF1 Entrez, Source | allograft inflammatory factor 1 | 9000 | -0.026 | 0.0261 | No |
| 185 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 9093 | -0.027 | 0.0214 | No |
| 186 | SGTB | SGTB Entrez, Source | small glutamine-rich tetratricopeptide repeat (TPR)-containing, beta | 9240 | -0.028 | 0.0126 | No |
| 187 | VPS16 | VPS16 Entrez, Source | vacuolar protein sorting 16 (yeast) | 9308 | -0.029 | 0.0099 | No |
| 188 | SLC7A2 | SLC7A2 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 | 9632 | -0.033 | -0.0120 | No |
| 189 | IGFBP6 | IGFBP6 Entrez, Source | insulin-like growth factor binding protein 6 | 9677 | -0.034 | -0.0125 | No |
| 190 | ADCYAP1 | ADCYAP1 Entrez, Source | adenylate cyclase activating polypeptide 1 (pituitary) | 9783 | -0.035 | -0.0176 | No |
| 191 | PRKAA2 | PRKAA2 Entrez, Source | protein kinase, AMP-activated, alpha 2 catalytic subunit | 9876 | -0.036 | -0.0217 | No |
| 192 | NR3C2 | NR3C2 Entrez, Source | nuclear receptor subfamily 3, group C, member 2 | 9890 | -0.036 | -0.0196 | No |
| 193 | PSMD3 | PSMD3 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 3 | 9942 | -0.037 | -0.0205 | No |
| 194 | FGF13 | FGF13 Entrez, Source | fibroblast growth factor 13 | 9981 | -0.037 | -0.0203 | No |
| 195 | RGS8 | RGS8 Entrez, Source | regulator of G-protein signalling 8 | 10208 | -0.040 | -0.0343 | No |
| 196 | TOMM22 | TOMM22 Entrez, Source | translocase of outer mitochondrial membrane 22 homolog (yeast) | 10227 | -0.040 | -0.0323 | No |
| 197 | RHOG | RHOG Entrez, Source | ras homolog gene family, member G (rho G) | 10269 | -0.040 | -0.0321 | No |
| 198 | PLEC1 | PLEC1 Entrez, Source | plectin 1, intermediate filament binding protein 500kDa | 10726 | -0.047 | -0.0630 | No |
| 199 | PCTK3 | PCTK3 Entrez, Source | PCTAIRE protein kinase 3 | 10763 | -0.047 | -0.0618 | No |
| 200 | CLSTN3 | CLSTN3 Entrez, Source | calsyntenin 3 | 10833 | -0.049 | -0.0630 | No |
| 201 | PPP2R2B | PPP2R2B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), beta isoform | 10989 | -0.051 | -0.0706 | No |
| 202 | NCDN | NCDN Entrez, Source | neurochondrin | 11136 | -0.054 | -0.0773 | No |
| 203 | MAPK12 | MAPK12 Entrez, Source | mitogen-activated protein kinase 12 | 11422 | -0.059 | -0.0941 | No |
| 204 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 11458 | -0.060 | -0.0918 | No |
| 205 | MAOB | MAOB Entrez, Source | monoamine oxidase B | 11520 | -0.061 | -0.0914 | No |
| 206 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 11521 | -0.061 | -0.0863 | No |
| 207 | DICER1 | DICER1 Entrez, Source | Dicer1, Dcr-1 homolog (Drosophila) | 11587 | -0.062 | -0.0861 | No |
| 208 | MC4R | MC4R Entrez, Source | melanocortin 4 receptor | 11608 | -0.063 | -0.0824 | No |
| 209 | PTOV1 | PTOV1 Entrez, Source | prostate tumor overexpressed gene 1 | 11677 | -0.064 | -0.0822 | No |
| 210 | MCART1 | MCART1 Entrez, Source | mitochondrial carrier triple repeat 1 | 11809 | -0.068 | -0.0866 | No |
| 211 | GABBR1 | GABBR1 Entrez, Source | gamma-aminobutyric acid (GABA) B receptor, 1 | 11865 | -0.069 | -0.0851 | No |
| 212 | NLN | NLN Entrez, Source | neurolysin (metallopeptidase M3 family) | 11920 | -0.070 | -0.0833 | No |
| 213 | COL5A3 | COL5A3 Entrez, Source | collagen, type V, alpha 3 | 11947 | -0.071 | -0.0794 | No |
| 214 | PPP2R5B | PPP2R5B Entrez, Source | protein phosphatase 2, regulatory subunit B (B56), beta isoform | 11996 | -0.073 | -0.0770 | No |
| 215 | AP1G1 | AP1G1 Entrez, Source | adaptor-related protein complex 1, gamma 1 subunit | 12092 | -0.076 | -0.0779 | No |
| 216 | VGLL4 | VGLL4 Entrez, Source | vestigial like 4 (Drosophila) | 12183 | -0.079 | -0.0782 | No |
| 217 | FUT11 | FUT11 Entrez, Source | fucosyltransferase 11 (alpha (1,3) fucosyltransferase) | 12339 | -0.085 | -0.0830 | No |
| 218 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 12342 | -0.085 | -0.0761 | No |
| 219 | GART | GART Entrez, Source | phosphoribosylglycinamide formyltransferase, phosphoribosylglycinamide synthetase, phosphoribosylaminoimidazole synthetase | 12346 | -0.085 | -0.0692 | No |
| 220 | RHOQ | RHOQ Entrez, Source | ras homolog gene family, member Q | 12429 | -0.088 | -0.0682 | No |
| 221 | SLC5A3 | SLC5A3 Entrez, Source | solute carrier family 5 (inositol transporters), member 3 | 12431 | -0.088 | -0.0609 | No |
| 222 | FBXO11 | FBXO11 Entrez, Source | F-box protein 11 | 12776 | -0.109 | -0.0781 | No |
| 223 | CALM3 | CALM3 Entrez, Source | calmodulin 3 (phosphorylase kinase, delta) | 12855 | -0.115 | -0.0744 | No |
| 224 | RAB3C | RAB3C Entrez, Source | RAB3C, member RAS oncogene family | 12925 | -0.124 | -0.0694 | No |
| 225 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 12953 | -0.126 | -0.0609 | No |
| 226 | TPM4 | TPM4 Entrez, Source | tropomyosin 4 | 12972 | -0.128 | -0.0516 | No |
| 227 | DTX2 | DTX2 Entrez, Source | deltex homolog 2 (Drosophila) | 13002 | -0.132 | -0.0428 | No |
| 228 | NRG1 | NRG1 Entrez, Source | neuregulin 1 | 13047 | -0.140 | -0.0346 | No |
| 229 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 13119 | -0.153 | -0.0273 | No |
| 230 | SLC31A2 | SLC31A2 Entrez, Source | solute carrier family 31 (copper transporters), member 2 | 13212 | -0.199 | -0.0177 | No |
| 231 | ARHGAP24 | ARHGAP24 Entrez, Source | Rho GTPase activating protein 24 | 13313 | -0.331 | 0.0022 | No |

