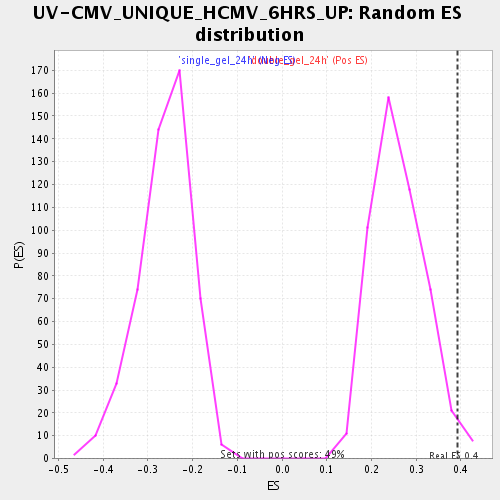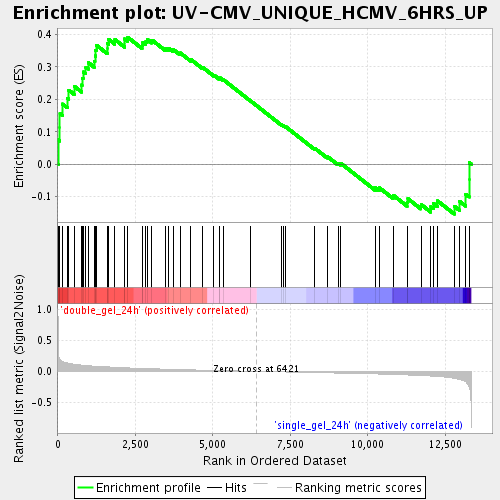
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | UV-CMV_UNIQUE_HCMV_6HRS_UP |
| Enrichment Score (ES) | 0.3916191 |
| Normalized Enrichment Score (NES) | 1.5024252 |
| Nominal p-value | 0.020366598 |
| FDR q-value | 0.2689236 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 10 | 0.347 | 0.0752 | Yes |
| 2 | MX2 | MX2 Entrez, Source | myxovirus (influenza virus) resistance 2 (mouse) | 64 | 0.196 | 0.1141 | Yes |
| 3 | DNAJB9 | DNAJB9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 9 | 66 | 0.194 | 0.1565 | Yes |
| 4 | TFEC | TFEC Entrez, Source | transcription factor EC | 144 | 0.160 | 0.1859 | Yes |
| 5 | TNFSF9 | TNFSF9 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 9 | 304 | 0.135 | 0.2034 | Yes |
| 6 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 347 | 0.130 | 0.2287 | Yes |
| 7 | IER2 | IER2 Entrez, Source | immediate early response 2 | 528 | 0.115 | 0.2403 | Yes |
| 8 | CX3CR1 | CX3CR1 Entrez, Source | chemokine (C-X3-C motif) receptor 1 | 768 | 0.101 | 0.2444 | Yes |
| 9 | CRYGD | CRYGD Entrez, Source | crystallin, gamma D | 781 | 0.100 | 0.2655 | Yes |
| 10 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 816 | 0.099 | 0.2846 | Yes |
| 11 | ELN | ELN Entrez, Source | elastin (supravalvular aortic stenosis, Williams-Beuren syndrome) | 900 | 0.095 | 0.2993 | Yes |
| 12 | OAS1 | OAS1 Entrez, Source | 2',5'-oligoadenylate synthetase 1, 40/46kDa | 980 | 0.092 | 0.3135 | Yes |
| 13 | TAC1 | TAC1 Entrez, Source | tachykinin, precursor 1 (substance K, substance P, neurokinin 1, neurokinin 2, neuromedin L, neurokinin alpha, neuropeptide K, neuropeptide gamma) | 1171 | 0.085 | 0.3177 | Yes |
| 14 | RGS6 | RGS6 Entrez, Source | regulator of G-protein signalling 6 | 1198 | 0.084 | 0.3341 | Yes |
| 15 | AREG | AREG Entrez, Source | amphiregulin (schwannoma-derived growth factor) | 1215 | 0.083 | 0.3511 | Yes |
| 16 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 1238 | 0.082 | 0.3675 | Yes |
| 17 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 1593 | 0.073 | 0.3567 | Yes |
| 18 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 1596 | 0.073 | 0.3724 | Yes |
| 19 | CD38 | CD38 Entrez, Source | CD38 molecule | 1632 | 0.072 | 0.3855 | Yes |
| 20 | COL15A1 | COL15A1 Entrez, Source | collagen, type XV, alpha 1 | 1838 | 0.067 | 0.3846 | Yes |
| 21 | GALNT3 | GALNT3 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 3 (GalNAc-T3) | 2156 | 0.059 | 0.3736 | Yes |
| 22 | PFKFB2 | PFKFB2 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2 | 2157 | 0.059 | 0.3865 | Yes |
| 23 | FGF2 | FGF2 Entrez, Source | fibroblast growth factor 2 (basic) | 2253 | 0.056 | 0.3916 | Yes |
| 24 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 2716 | 0.048 | 0.3673 | No |
| 25 | CRY1 | CRY1 Entrez, Source | cryptochrome 1 (photolyase-like) | 2740 | 0.047 | 0.3759 | No |
| 26 | PDE1A | PDE1A Entrez, Source | phosphodiesterase 1A, calmodulin-dependent | 2839 | 0.045 | 0.3785 | No |
| 27 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 2878 | 0.045 | 0.3855 | No |
| 28 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 3035 | 0.043 | 0.3830 | No |
| 29 | SOCS1 | SOCS1 Entrez, Source | suppressor of cytokine signaling 1 | 3470 | 0.036 | 0.3582 | No |
| 30 | MSX1 | MSX1 Entrez, Source | msh homeobox homolog 1 (Drosophila) | 3576 | 0.034 | 0.3577 | No |
| 31 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 3724 | 0.032 | 0.3537 | No |
| 32 | EPHA5 | EPHA5 Entrez, Source | EPH receptor A5 | 3947 | 0.029 | 0.3434 | No |
| 33 | PLA2G4A | PLA2G4A Entrez, Source | phospholipase A2, group IVA (cytosolic, calcium-dependent) | 4288 | 0.025 | 0.3232 | No |
| 34 | IRF7 | IRF7 Entrez, Source | interferon regulatory factor 7 | 4680 | 0.020 | 0.2981 | No |
| 35 | IGFBP2 | IGFBP2 Entrez, Source | insulin-like growth factor binding protein 2, 36kDa | 5034 | 0.016 | 0.2750 | No |
| 36 | BRCA2 | BRCA2 Entrez, Source | breast cancer 2, early onset | 5213 | 0.014 | 0.2646 | No |
| 37 | TREX1 | TREX1 Entrez, Source | three prime repair exonuclease 1 | 5230 | 0.013 | 0.2663 | No |
| 38 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 5344 | 0.012 | 0.2604 | No |
| 39 | HBEGF | HBEGF Entrez, Source | heparin-binding EGF-like growth factor | 6200 | 0.002 | 0.1966 | No |
| 40 | BAMBI | BAMBI Entrez, Source | BMP and activin membrane-bound inhibitor homolog (Xenopus laevis) | 7218 | -0.008 | 0.1217 | No |
| 41 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 7292 | -0.009 | 0.1181 | No |
| 42 | PIK3R3 | PIK3R3 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 3 (p55, gamma) | 7359 | -0.009 | 0.1151 | No |
| 43 | GTF2B | GTF2B Entrez, Source | general transcription factor IIB | 8289 | -0.019 | 0.0493 | No |
| 44 | MYD88 | MYD88 Entrez, Source | myeloid differentiation primary response gene (88) | 8694 | -0.023 | 0.0238 | No |
| 45 | TRADD | TRADD Entrez, Source | TNFRSF1A-associated via death domain | 9040 | -0.026 | 0.0037 | No |
| 46 | CHEK2 | CHEK2 Entrez, Source | CHK2 checkpoint homolog (S. pombe) | 9129 | -0.027 | 0.0030 | No |
| 47 | ING1 | ING1 Entrez, Source | inhibitor of growth family, member 1 | 10251 | -0.040 | -0.0726 | No |
| 48 | CCRK | CCRK Entrez, Source | cell cycle related kinase | 10374 | -0.042 | -0.0727 | No |
| 49 | RBBP6 | RBBP6 Entrez, Source | retinoblastoma binding protein 6 | 10819 | -0.048 | -0.0955 | No |
| 50 | TDRD7 | TDRD7 Entrez, Source | tudor domain containing 7 | 11269 | -0.056 | -0.1170 | No |
| 51 | RQCD1 | RQCD1 Entrez, Source | RCD1 required for cell differentiation1 homolog (S. pombe) | 11296 | -0.057 | -0.1065 | No |
| 52 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 11718 | -0.065 | -0.1240 | No |
| 53 | RRAD | RRAD Entrez, Source | Ras-related associated with diabetes | 12036 | -0.074 | -0.1316 | No |
| 54 | IL1RAP | IL1RAP Entrez, Source | interleukin 1 receptor accessory protein | 12123 | -0.077 | -0.1213 | No |
| 55 | ELF1 | ELF1 Entrez, Source | E74-like factor 1 (ets domain transcription factor) | 12250 | -0.081 | -0.1130 | No |
| 56 | NFKB2 | NFKB2 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) | 12794 | -0.110 | -0.1298 | No |
| 57 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 12964 | -0.127 | -0.1147 | No |
| 58 | CH25H | CH25H Entrez, Source | cholesterol 25-hydroxylase | 13161 | -0.166 | -0.0930 | No |
| 59 | STX11 | STX11 Entrez, Source | syntaxin 11 | 13272 | -0.242 | -0.0483 | No |
| 60 | FGFR3 | FGFR3 Entrez, Source | fibroblast growth factor receptor 3 (achondroplasia, thanatophoric dwarfism) | 13275 | -0.244 | 0.0050 | No |