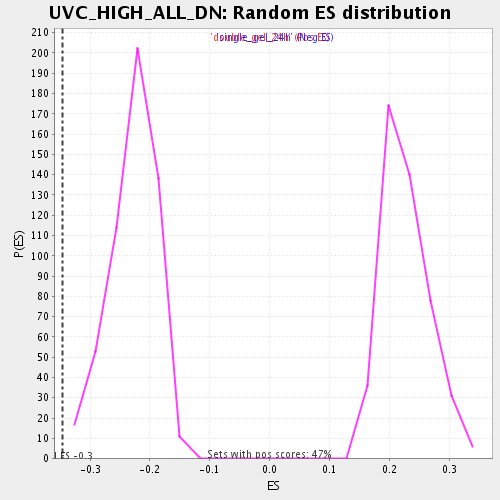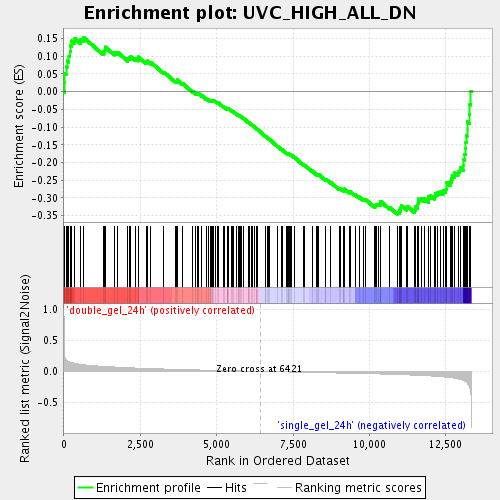
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | UVC_HIGH_ALL_DN |
| Enrichment Score (ES) | -0.3455874 |
| Normalized Enrichment Score (NES) | -1.50934 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.27325168 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 25 | 0.248 | 0.0259 | No |
| 2 | IDI1 | IDI1 Entrez, Source | isopentenyl-diphosphate delta isomerase 1 | 29 | 0.238 | 0.0523 | No |
| 3 | HERPUD1 | HERPUD1 Entrez, Source | homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 | 74 | 0.189 | 0.0701 | No |
| 4 | CXCL12 | CXCL12 Entrez, Source | chemokine (C-X-C motif) ligand 12 (stromal cell-derived factor 1) | 105 | 0.175 | 0.0875 | No |
| 5 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 164 | 0.155 | 0.1005 | No |
| 6 | IGF2BP3 | IGF2BP3 Entrez, Source | insulin-like growth factor 2 mRNA binding protein 3 | 199 | 0.149 | 0.1146 | No |
| 7 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 206 | 0.148 | 0.1307 | No |
| 8 | HMGCR | HMGCR Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A reductase | 250 | 0.141 | 0.1433 | No |
| 9 | FZD2 | FZD2 Entrez, Source | frizzled homolog 2 (Drosophila) | 360 | 0.129 | 0.1494 | No |
| 10 | IRS1 | IRS1 Entrez, Source | insulin receptor substrate 1 | 549 | 0.114 | 0.1479 | No |
| 11 | SQLE | SQLE Entrez, Source | squalene epoxidase | 634 | 0.108 | 0.1536 | No |
| 12 | NTF3 | NTF3 Entrez, Source | neurotrophin 3 | 1291 | 0.081 | 0.1129 | No |
| 13 | HAS2 | HAS2 Entrez, Source | hyaluronan synthase 2 | 1343 | 0.079 | 0.1178 | No |
| 14 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 1350 | 0.079 | 0.1262 | No |
| 15 | ZHX3 | ZHX3 Entrez, Source | zinc fingers and homeoboxes 3 | 1652 | 0.071 | 0.1113 | No |
| 16 | GATA2 | GATA2 Entrez, Source | GATA binding protein 2 | 1755 | 0.069 | 0.1113 | No |
| 17 | AGTR1 | AGTR1 Entrez, Source | angiotensin II receptor, type 1 | 2090 | 0.060 | 0.0927 | No |
| 18 | IL4R | IL4R Entrez, Source | interleukin 4 receptor | 2130 | 0.059 | 0.0964 | No |
| 19 | ADM | ADM Entrez, Source | adrenomedullin | 2189 | 0.058 | 0.0984 | No |
| 20 | CRKL | CRKL Entrez, Source | v-crk sarcoma virus CT10 oncogene homolog (avian)-like | 2336 | 0.055 | 0.0935 | No |
| 21 | YY1 | YY1 Entrez, Source | YY1 transcription factor | 2429 | 0.053 | 0.0924 | No |
| 22 | GREM1 | GREM1 Entrez, Source | gremlin 1, cysteine knot superfamily, homolog (Xenopus laevis) | 2434 | 0.053 | 0.0981 | No |
| 23 | KLF9 | KLF9 Entrez, Source | Kruppel-like factor 9 | 2689 | 0.048 | 0.0842 | No |
| 24 | TWIST1 | TWIST1 Entrez, Source | twist homolog 1 (acrocephalosyndactyly 3; Saethre-Chotzen syndrome) (Drosophila) | 2728 | 0.047 | 0.0866 | No |
| 25 | PDGFRA | PDGFRA Entrez, Source | platelet-derived growth factor receptor, alpha polypeptide | 2844 | 0.045 | 0.0830 | No |
| 26 | DUSP11 | DUSP11 Entrez, Source | dual specificity phosphatase 11 (RNA/RNP complex 1-interacting) | 3274 | 0.039 | 0.0548 | No |
| 27 | NFIL3 | NFIL3 Entrez, Source | nuclear factor, interleukin 3 regulated | 3661 | 0.033 | 0.0292 | No |
| 28 | MAP1B | MAP1B Entrez, Source | microtubule-associated protein 1B | 3700 | 0.033 | 0.0300 | No |
| 29 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 3705 | 0.032 | 0.0333 | No |
| 30 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 3887 | 0.030 | 0.0229 | No |
| 31 | CEBPD | CEBPD Entrez, Source | CCAAT/enhancer binding protein (C/EBP), delta | 4222 | 0.026 | 0.0005 | No |
| 32 | JUND | JUND Entrez, Source | jun D proto-oncogene | 4321 | 0.024 | -0.0042 | No |
| 33 | STK38 | STK38 Entrez, Source | serine/threonine kinase 38 | 4377 | 0.023 | -0.0058 | No |
| 34 | ANKRD15 | ANKRD15 Entrez, Source | ankyrin repeat domain 15 | 4389 | 0.023 | -0.0040 | No |
| 35 | EIF4E | EIF4E Entrez, Source | eukaryotic translation initiation factor 4E | 4511 | 0.022 | -0.0108 | No |
| 36 | FBXO46 | FBXO46 Entrez, Source | F-box protein 46 | 4662 | 0.020 | -0.0199 | No |
| 37 | GNE | GNE Entrez, Source | glucosamine (UDP-N-acetyl)-2-epimerase/N-acetylmannosamine kinase | 4730 | 0.019 | -0.0228 | No |
| 38 | RAF1 | RAF1 Entrez, Source | v-raf-1 murine leukemia viral oncogene homolog 1 | 4811 | 0.018 | -0.0268 | No |
| 39 | PRKCBP1 | PRKCBP1 Entrez, Source | protein kinase C binding protein 1 | 4831 | 0.018 | -0.0262 | No |
| 40 | UBE2N | UBE2N Entrez, Source | ubiquitin-conjugating enzyme E2N (UBC13 homolog, yeast) | 4864 | 0.018 | -0.0267 | No |
| 41 | EDD1 | EDD1 Entrez, Source | E3 ubiquitin protein ligase, HECT domain containing, 1 | 4878 | 0.018 | -0.0257 | No |
| 42 | TPST1 | TPST1 Entrez, Source | tyrosylprotein sulfotransferase 1 | 4909 | 0.017 | -0.0260 | No |
| 43 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 4950 | 0.017 | -0.0272 | No |
| 44 | ATP6V1B2 | ATP6V1B2 Entrez, Source | ATPase, H+ transporting, lysosomal 56/58kDa, V1 subunit B2 | 5011 | 0.016 | -0.0300 | No |
| 45 | TGIF | TGIF Entrez, Source | TGFB-induced factor (TALE family homeobox) | 5050 | 0.015 | -0.0312 | No |
| 46 | ADNP | ADNP Entrez, Source | activity-dependent neuroprotector | 5220 | 0.013 | -0.0425 | No |
| 47 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 5269 | 0.013 | -0.0447 | No |
| 48 | ZFP36L2 | ZFP36L2 Entrez, Source | zinc finger protein 36, C3H type-like 2 | 5350 | 0.012 | -0.0494 | No |
| 49 | RNF4 | RNF4 Entrez, Source | ring finger protein 4 | 5355 | 0.012 | -0.0484 | No |
| 50 | RND3 | RND3 Entrez, Source | Rho family GTPase 3 | 5357 | 0.012 | -0.0471 | No |
| 51 | BRD8 | BRD8 Entrez, Source | bromodomain containing 8 | 5377 | 0.012 | -0.0473 | No |
| 52 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 5475 | 0.010 | -0.0535 | No |
| 53 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 5517 | 0.010 | -0.0554 | No |
| 54 | RAB1A | RAB1A Entrez, Source | RAB1A, member RAS oncogene family | 5561 | 0.009 | -0.0576 | No |
| 55 | NFYB | NFYB Entrez, Source | nuclear transcription factor Y, beta | 5644 | 0.009 | -0.0629 | No |
| 56 | TSPAN5 | TSPAN5 Entrez, Source | tetraspanin 5 | 5710 | 0.008 | -0.0670 | No |
| 57 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 5728 | 0.008 | -0.0674 | No |
| 58 | SPOP | SPOP Entrez, Source | speckle-type POZ protein | 5734 | 0.008 | -0.0669 | No |
| 59 | SERTAD2 | SERTAD2 Entrez, Source | SERTA domain containing 2 | 5784 | 0.007 | -0.0699 | No |
| 60 | NFE2L2 | NFE2L2 Entrez, Source | nuclear factor (erythroid-derived 2)-like 2 | 5811 | 0.007 | -0.0711 | No |
| 61 | GTF2E1 | GTF2E1 Entrez, Source | general transcription factor IIE, polypeptide 1, alpha 56kDa | 5878 | 0.006 | -0.0755 | No |
| 62 | TMEM97 | TMEM97 Entrez, Source | transmembrane protein 97 | 6035 | 0.004 | -0.0868 | No |
| 63 | PGRMC2 | PGRMC2 Entrez, Source | progesterone receptor membrane component 2 | 6040 | 0.004 | -0.0867 | No |
| 64 | PUM2 | PUM2 Entrez, Source | pumilio homolog 2 (Drosophila) | 6077 | 0.004 | -0.0890 | No |
| 65 | COIL | COIL Entrez, Source | coilin | 6138 | 0.003 | -0.0931 | No |
| 66 | FUS | FUS Entrez, Source | fusion (involved in t(12;16) in malignant liposarcoma) | 6187 | 0.003 | -0.0965 | No |
| 67 | PTDSR | PTDSR Entrez, Source | phosphatidylserine receptor | 6251 | 0.002 | -0.1011 | No |
| 68 | FGF5 | FGF5 Entrez, Source | fibroblast growth factor 5 | 6286 | 0.001 | -0.1035 | No |
| 69 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 6305 | 0.001 | -0.1047 | No |
| 70 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 6322 | 0.001 | -0.1058 | No |
| 71 | ARF6 | ARF6 Entrez, Source | ADP-ribosylation factor 6 | 6594 | -0.002 | -0.1262 | No |
| 72 | USP10 | USP10 Entrez, Source | ubiquitin specific peptidase 10 | 6609 | -0.002 | -0.1270 | No |
| 73 | TUBGCP3 | TUBGCP3 Entrez, Source | tubulin, gamma complex associated protein 3 | 6657 | -0.003 | -0.1303 | No |
| 74 | MAPK14 | MAPK14 Entrez, Source | mitogen-activated protein kinase 14 | 6688 | -0.003 | -0.1322 | No |
| 75 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 6719 | -0.003 | -0.1342 | No |
| 76 | BLCAP | BLCAP Entrez, Source | bladder cancer associated protein | 7002 | -0.006 | -0.1549 | No |
| 77 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 7107 | -0.007 | -0.1621 | No |
| 78 | ATP2A2 | ATP2A2 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 | 7144 | -0.007 | -0.1640 | No |
| 79 | NR2F2 | NR2F2 Entrez, Source | nuclear receptor subfamily 2, group F, member 2 | 7287 | -0.009 | -0.1738 | No |
| 80 | CASP3 | CASP3 Entrez, Source | caspase 3, apoptosis-related cysteine peptidase | 7301 | -0.009 | -0.1738 | No |
| 81 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 7336 | -0.009 | -0.1754 | No |
| 82 | SS18 | SS18 Entrez, Source | synovial sarcoma translocation, chromosome 18 | 7348 | -0.009 | -0.1753 | No |
| 83 | RAB11A | RAB11A Entrez, Source | RAB11A, member RAS oncogene family | 7378 | -0.009 | -0.1764 | No |
| 84 | PMP22 | PMP22 Entrez, Source | peripheral myelin protein 22 | 7414 | -0.010 | -0.1780 | No |
| 85 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 7439 | -0.010 | -0.1787 | No |
| 86 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 7532 | -0.011 | -0.1845 | No |
| 87 | PITPNB | PITPNB Entrez, Source | phosphatidylinositol transfer protein, beta | 7840 | -0.014 | -0.2062 | No |
| 88 | ARHGEF7 | ARHGEF7 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 7 | 7877 | -0.014 | -0.2073 | No |
| 89 | CLINT1 | CLINT1 Entrez, Source | clathrin interactor 1 | 8119 | -0.017 | -0.2237 | No |
| 90 | RAB14 | RAB14 Entrez, Source | RAB14, member RAS oncogene family | 8275 | -0.018 | -0.2334 | No |
| 91 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 8310 | -0.019 | -0.2339 | No |
| 92 | MTMR3 | MTMR3 Entrez, Source | myotubularin related protein 3 | 8338 | -0.019 | -0.2338 | No |
| 93 | EDG2 | EDG2 Entrez, Source | endothelial differentiation, lysophosphatidic acid G-protein-coupled receptor, 2 | 8571 | -0.022 | -0.2490 | No |
| 94 | THUMPD1 | THUMPD1 Entrez, Source | THUMP domain containing 1 | 8574 | -0.022 | -0.2467 | No |
| 95 | UBE2G1 | UBE2G1 Entrez, Source | ubiquitin-conjugating enzyme E2G 1 (UBC7 homolog, yeast) | 8739 | -0.023 | -0.2565 | No |
| 96 | PPP2R1B | PPP2R1B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), beta isoform | 9012 | -0.026 | -0.2742 | No |
| 97 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 9041 | -0.026 | -0.2734 | No |
| 98 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 9143 | -0.027 | -0.2780 | No |
| 99 | STX3 | STX3 Entrez, Source | syntaxin 3 | 9160 | -0.028 | -0.2761 | No |
| 100 | NDUFV2 | NDUFV2 Entrez, Source | NADH dehydrogenase (ubiquinone) flavoprotein 2, 24kDa | 9181 | -0.028 | -0.2745 | No |
| 101 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9337 | -0.029 | -0.2830 | No |
| 102 | RBM5 | RBM5 Entrez, Source | RNA binding motif protein 5 | 9365 | -0.030 | -0.2817 | No |
| 103 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 9527 | -0.032 | -0.2904 | No |
| 104 | AURKA | AURKA Entrez, Source | aurora kinase A | 9667 | -0.034 | -0.2971 | No |
| 105 | TRIP10 | TRIP10 Entrez, Source | thyroid hormone receptor interactor 10 | 9804 | -0.035 | -0.3035 | No |
| 106 | NCOA3 | NCOA3 Entrez, Source | nuclear receptor coactivator 3 | 9861 | -0.036 | -0.3038 | No |
| 107 | CDK7 | CDK7 Entrez, Source | cyclin-dependent kinase 7 (MO15 homolog, Xenopus laevis, cdk-activating kinase) | 10156 | -0.039 | -0.3217 | No |
| 108 | PTPN1 | PTPN1 Entrez, Source | protein tyrosine phosphatase, non-receptor type 1 | 10210 | -0.040 | -0.3213 | No |
| 109 | UBTF | UBTF Entrez, Source | upstream binding transcription factor, RNA polymerase I | 10220 | -0.040 | -0.3175 | No |
| 110 | SLC20A2 | SLC20A2 Entrez, Source | solute carrier family 20 (phosphate transporter), member 2 | 10283 | -0.040 | -0.3177 | No |
| 111 | MLX | MLX Entrez, Source | MAX-like protein X | 10350 | -0.041 | -0.3181 | No |
| 112 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 10367 | -0.041 | -0.3147 | No |
| 113 | CASP8 | CASP8 Entrez, Source | caspase 8, apoptosis-related cysteine peptidase | 10368 | -0.041 | -0.3100 | No |
| 114 | DR1 | DR1 Entrez, Source | down-regulator of transcription 1, TBP-binding (negative cofactor 2) | 10653 | -0.046 | -0.3265 | No |
| 115 | CDR2 | CDR2 Entrez, Source | cerebellar degeneration-related protein 2, 62kDa | 10906 | -0.050 | -0.3400 | Yes |
| 116 | PLEKHC1 | PLEKHC1 Entrez, Source | pleckstrin homology domain containing, family C (with FERM domain) member 1 | 10973 | -0.051 | -0.3394 | Yes |
| 117 | SPRED2 | SPRED2 Entrez, Source | sprouty-related, EVH1 domain containing 2 | 10977 | -0.051 | -0.3339 | Yes |
| 118 | STRN3 | STRN3 Entrez, Source | striatin, calmodulin binding protein 3 | 11023 | -0.051 | -0.3316 | Yes |
| 119 | PPP1R12A | PPP1R12A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 12A | 11026 | -0.051 | -0.3260 | Yes |
| 120 | SIAH2 | SIAH2 Entrez, Source | seven in absentia homolog 2 (Drosophila) | 11053 | -0.052 | -0.3221 | Yes |
| 121 | CNOT2 | CNOT2 Entrez, Source | CCR4-NOT transcription complex, subunit 2 | 11214 | -0.055 | -0.3281 | Yes |
| 122 | TAF11 | TAF11 Entrez, Source | TAF11 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 28kDa | 11247 | -0.056 | -0.3243 | Yes |
| 123 | PRKCD | PRKCD Entrez, Source | protein kinase C, delta | 11467 | -0.060 | -0.3342 | Yes |
| 124 | MARCKS | MARCKS Entrez, Source | myristoylated alanine-rich protein kinase C substrate | 11501 | -0.060 | -0.3299 | Yes |
| 125 | REV3L | REV3L Entrez, Source | REV3-like, catalytic subunit of DNA polymerase zeta (yeast) | 11512 | -0.061 | -0.3239 | Yes |
| 126 | LBR | LBR Entrez, Source | lamin B receptor | 11583 | -0.062 | -0.3222 | Yes |
| 127 | DICER1 | DICER1 Entrez, Source | Dicer1, Dcr-1 homolog (Drosophila) | 11587 | -0.062 | -0.3155 | Yes |
| 128 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 11598 | -0.062 | -0.3093 | Yes |
| 129 | PTK2 | PTK2 Entrez, Source | PTK2 protein tyrosine kinase 2 | 11607 | -0.063 | -0.3028 | Yes |
| 130 | CBX3 | CBX3 Entrez, Source | chromobox homolog 3 (HP1 gamma homolog, Drosophila) | 11686 | -0.064 | -0.3016 | Yes |
| 131 | STXBP3 | STXBP3 Entrez, Source | syntaxin binding protein 3 | 11792 | -0.067 | -0.3020 | Yes |
| 132 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 11928 | -0.071 | -0.3043 | Yes |
| 133 | NAP1L1 | NAP1L1 Entrez, Source | nucleosome assembly protein 1-like 1 | 11929 | -0.071 | -0.2964 | Yes |
| 134 | TRIM32 | TRIM32 Entrez, Source | tripartite motif-containing 32 | 12008 | -0.073 | -0.2942 | Yes |
| 135 | BMPR1A | BMPR1A Entrez, Source | bone morphogenetic protein receptor, type IA | 12130 | -0.077 | -0.2947 | Yes |
| 136 | FADD | FADD Entrez, Source | Fas (TNFRSF6)-associated via death domain | 12156 | -0.078 | -0.2879 | Yes |
| 137 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 12226 | -0.080 | -0.2842 | Yes |
| 138 | PARP1 | PARP1 Entrez, Source | poly (ADP-ribose) polymerase family, member 1 | 12319 | -0.083 | -0.2818 | Yes |
| 139 | VEGFC | VEGFC Entrez, Source | vascular endothelial growth factor C | 12416 | -0.088 | -0.2793 | Yes |
| 140 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 12503 | -0.092 | -0.2755 | Yes |
| 141 | PHC2 | PHC2 Entrez, Source | polyhomeotic homolog 2 (Drosophila) | 12525 | -0.093 | -0.2667 | Yes |
| 142 | STX7 | STX7 Entrez, Source | syntaxin 7 | 12526 | -0.093 | -0.2563 | Yes |
| 143 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 12641 | -0.099 | -0.2539 | Yes |
| 144 | MAT2A | MAT2A Entrez, Source | methionine adenosyltransferase II, alpha | 12685 | -0.102 | -0.2457 | Yes |
| 145 | PODXL | PODXL Entrez, Source | podocalyxin-like | 12713 | -0.104 | -0.2361 | Yes |
| 146 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 12787 | -0.110 | -0.2293 | Yes |
| 147 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 12899 | -0.121 | -0.2242 | Yes |
| 148 | KIF2A | KIF2A Entrez, Source | kinesin heavy chain member 2A | 12962 | -0.127 | -0.2147 | Yes |
| 149 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 13084 | -0.146 | -0.2074 | Yes |
| 150 | RRAS2 | RRAS2 Entrez, Source | related RAS viral (r-ras) oncogene homolog 2 | 13088 | -0.147 | -0.1912 | Yes |
| 151 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 13119 | -0.153 | -0.1764 | Yes |
| 152 | MET | MET Entrez, Source | met proto-oncogene (hepatocyte growth factor receptor) | 13148 | -0.161 | -0.1605 | Yes |
| 153 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 13153 | -0.163 | -0.1426 | Yes |
| 154 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 13167 | -0.169 | -0.1246 | Yes |
| 155 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13194 | -0.185 | -0.1059 | Yes |
| 156 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 13195 | -0.185 | -0.0851 | Yes |
| 157 | PDE4B | PDE4B Entrez, Source | phosphodiesterase 4B, cAMP-specific (phosphodiesterase E4 dunce homolog, Drosophila) | 13271 | -0.241 | -0.0638 | Yes |
| 158 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13284 | -0.263 | -0.0353 | Yes |
| 159 | GNAI1 | GNAI1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | 13318 | -0.354 | 0.0018 | Yes |