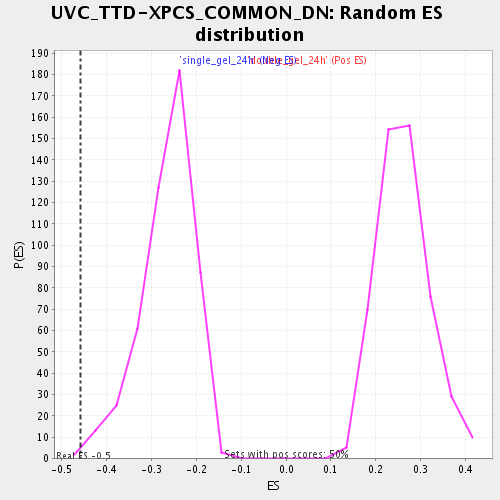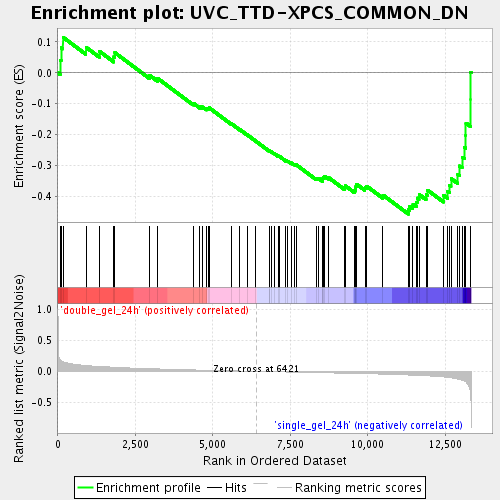
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | UVC_TTD-XPCS_COMMON_DN |
| Enrichment Score (ES) | -0.4587169 |
| Normalized Enrichment Score (NES) | -1.7359295 |
| Nominal p-value | 0.0020 |
| FDR q-value | 0.1470294 |
| FWER p-Value | 0.997 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IGF1R | IGF1R Entrez, Source | insulin-like growth factor 1 receptor | 76 | 0.189 | 0.0407 | No |
| 2 | CXCL12 | CXCL12 Entrez, Source | chemokine (C-X-C motif) ligand 12 (stromal cell-derived factor 1) | 105 | 0.175 | 0.0816 | No |
| 3 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 164 | 0.155 | 0.1154 | No |
| 4 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 906 | 0.095 | 0.0830 | No |
| 5 | CENTB2 | CENTB2 Entrez, Source | centaurin, beta 2 | 1335 | 0.079 | 0.0702 | No |
| 6 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 1805 | 0.067 | 0.0514 | No |
| 7 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 1837 | 0.067 | 0.0655 | No |
| 8 | KLF6 | KLF6 Entrez, Source | Kruppel-like factor 6 | 2960 | 0.044 | -0.0083 | No |
| 9 | MARCH7 | MARCH7 Entrez, Source | membrane-associated ring finger (C3HC4) 7 | 3217 | 0.040 | -0.0178 | No |
| 10 | E2F5 | E2F5 Entrez, Source | E2F transcription factor 5, p130-binding | 4371 | 0.024 | -0.0989 | No |
| 11 | ITSN1 | ITSN1 Entrez, Source | intersectin 1 (SH3 domain protein) | 4584 | 0.021 | -0.1097 | No |
| 12 | TIAM1 | TIAM1 Entrez, Source | T-cell lymphoma invasion and metastasis 1 | 4663 | 0.020 | -0.1107 | No |
| 13 | ORC2L | ORC2L Entrez, Source | origin recognition complex, subunit 2-like (yeast) | 4798 | 0.019 | -0.1162 | No |
| 14 | GULP1 | GULP1 Entrez, Source | GULP, engulfment adaptor PTB domain containing 1 | 4845 | 0.018 | -0.1153 | No |
| 15 | EDD1 | EDD1 Entrez, Source | E3 ubiquitin protein ligase, HECT domain containing, 1 | 4878 | 0.018 | -0.1133 | No |
| 16 | HOMER1 | HOMER1 Entrez, Source | homer homolog 1 (Drosophila) | 5598 | 0.009 | -0.1652 | No |
| 17 | GBE1 | GBE1 Entrez, Source | glucan (1,4-alpha-), branching enzyme 1 (glycogen branching enzyme, Andersen disease, glycogen storage disease type IV) | 5875 | 0.006 | -0.1846 | No |
| 18 | RBM39 | RBM39 Entrez, Source | RNA binding motif protein 39 | 6107 | 0.004 | -0.2011 | No |
| 19 | GTF2F2 | GTF2F2 Entrez, Source | general transcription factor IIF, polypeptide 2, 30kDa | 6378 | 0.000 | -0.2214 | No |
| 20 | MYO1B | MYO1B Entrez, Source | myosin IB | 6819 | -0.004 | -0.2535 | No |
| 21 | TRIM2 | TRIM2 Entrez, Source | tripartite motif-containing 2 | 6833 | -0.004 | -0.2535 | No |
| 22 | SEPT9 | SEPT9 Entrez, Source | septin 9 | 6891 | -0.005 | -0.2567 | No |
| 23 | ZNF292 | ZNF292 Entrez, Source | zinc finger protein 292 | 6983 | -0.005 | -0.2622 | No |
| 24 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 7107 | -0.007 | -0.2698 | No |
| 25 | CDC2L5 | CDC2L5 Entrez, Source | cell division cycle 2-like 5 (cholinesterase-related cell division controller) | 7141 | -0.007 | -0.2706 | No |
| 26 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 7360 | -0.009 | -0.2848 | No |
| 27 | MAP2K4 | MAP2K4 Entrez, Source | mitogen-activated protein kinase kinase 4 | 7416 | -0.010 | -0.2865 | No |
| 28 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 7532 | -0.011 | -0.2925 | No |
| 29 | ARIH1 | ARIH1 Entrez, Source | ariadne homolog, ubiquitin-conjugating enzyme E2 binding protein, 1 (Drosophila) | 7627 | -0.012 | -0.2967 | No |
| 30 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 7684 | -0.012 | -0.2979 | No |
| 31 | MTMR3 | MTMR3 Entrez, Source | myotubularin related protein 3 | 8338 | -0.019 | -0.3424 | No |
| 32 | NUP98 | NUP98 Entrez, Source | nucleoporin 98kDa | 8404 | -0.020 | -0.3424 | No |
| 33 | MGLL | MGLL Entrez, Source | monoglyceride lipase | 8545 | -0.021 | -0.3478 | No |
| 34 | MARK3 | MARK3 Entrez, Source | MAP/microtubule affinity-regulating kinase 3 | 8547 | -0.021 | -0.3426 | No |
| 35 | EDG2 | EDG2 Entrez, Source | endothelial differentiation, lysophosphatidic acid G-protein-coupled receptor, 2 | 8571 | -0.022 | -0.3391 | No |
| 36 | GLRB | GLRB Entrez, Source | glycine receptor, beta | 8597 | -0.022 | -0.3356 | No |
| 37 | ZMYND11 | ZMYND11 Entrez, Source | zinc finger, MYND domain containing 11 | 8725 | -0.023 | -0.3395 | No |
| 38 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 9251 | -0.029 | -0.3720 | No |
| 39 | DPYD | DPYD Entrez, Source | dihydropyrimidine dehydrogenase | 9279 | -0.029 | -0.3670 | No |
| 40 | PTPN2 | PTPN2 Entrez, Source | protein tyrosine phosphatase, non-receptor type 2 | 9568 | -0.032 | -0.3807 | No |
| 41 | NFIB | NFIB Entrez, Source | nuclear factor I/B | 9610 | -0.033 | -0.3757 | No |
| 42 | MNAT1 | MNAT1 Entrez, Source | menage a trois homolog 1, cyclin H assembly factor (Xenopus laevis) | 9616 | -0.033 | -0.3680 | No |
| 43 | OGT | OGT Entrez, Source | O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) | 9640 | -0.033 | -0.3616 | No |
| 44 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 9911 | -0.036 | -0.3730 | No |
| 45 | APPBP2 | APPBP2 Entrez, Source | amyloid beta precursor protein (cytoplasmic tail) binding protein 2 | 9965 | -0.037 | -0.3680 | No |
| 46 | FARS2 | FARS2 Entrez, Source | phenylalanine-tRNA synthetase 2 (mitochondrial) | 10489 | -0.043 | -0.3967 | No |
| 47 | MTF2 | MTF2 Entrez, Source | metal response element binding transcription factor 2 | 11313 | -0.057 | -0.4447 | Yes |
| 48 | RASA1 | RASA1 Entrez, Source | RAS p21 protein activator (GTPase activating protein) 1 | 11353 | -0.058 | -0.4334 | Yes |
| 49 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 11458 | -0.060 | -0.4266 | Yes |
| 50 | CNOT4 | CNOT4 Entrez, Source | CCR4-NOT transcription complex, subunit 4 | 11569 | -0.062 | -0.4197 | Yes |
| 51 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 11598 | -0.062 | -0.4064 | Yes |
| 52 | FUT8 | FUT8 Entrez, Source | fucosyltransferase 8 (alpha (1,6) fucosyltransferase) | 11656 | -0.064 | -0.3950 | Yes |
| 53 | PCTK2 | PCTK2 Entrez, Source | PCTAIRE protein kinase 2 | 11883 | -0.070 | -0.3949 | Yes |
| 54 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 11928 | -0.071 | -0.3809 | Yes |
| 55 | CASK | CASK Entrez, Source | calcium/calmodulin-dependent serine protein kinase (MAGUK family) | 12444 | -0.088 | -0.3979 | Yes |
| 56 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 12577 | -0.095 | -0.3844 | Yes |
| 57 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 12641 | -0.099 | -0.3648 | Yes |
| 58 | BMP2K | BMP2K Entrez, Source | BMP2 inducible kinase | 12687 | -0.102 | -0.3431 | Yes |
| 59 | PDLIM5 | PDLIM5 Entrez, Source | PDZ and LIM domain 5 | 12904 | -0.122 | -0.3294 | Yes |
| 60 | KIF2A | KIF2A Entrez, Source | kinesin heavy chain member 2A | 12962 | -0.127 | -0.3024 | Yes |
| 61 | NRG1 | NRG1 Entrez, Source | neuregulin 1 | 13047 | -0.140 | -0.2744 | Yes |
| 62 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 13119 | -0.153 | -0.2422 | Yes |
| 63 | SOS2 | SOS2 Entrez, Source | son of sevenless homolog 2 (Drosophila) | 13162 | -0.167 | -0.2044 | Yes |
| 64 | CLASP1 | CLASP1 Entrez, Source | cytoplasmic linker associated protein 1 | 13168 | -0.170 | -0.1631 | Yes |
| 65 | GNAI1 | GNAI1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | 13318 | -0.354 | -0.0872 | Yes |
| 66 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 13320 | -0.362 | 0.0017 | Yes |