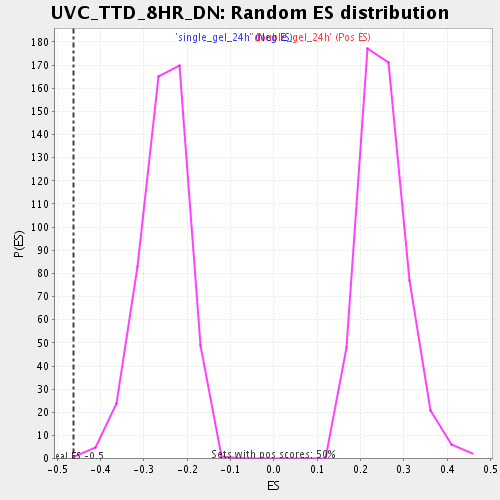
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | UVC_TTD_8HR_DN |
| Enrichment Score (ES) | -0.4622043 |
| Normalized Enrichment Score (NES) | -1.8222574 |
| Nominal p-value | 0.002008032 |
| FDR q-value | 0.11816355 |
| FWER p-Value | 0.945 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CXCL12 | CXCL12 Entrez, Source | chemokine (C-X-C motif) ligand 12 (stromal cell-derived factor 1) | 105 | 0.175 | 0.0253 | No |
| 2 | IGF2BP3 | IGF2BP3 Entrez, Source | insulin-like growth factor 2 mRNA binding protein 3 | 199 | 0.149 | 0.0466 | No |
| 3 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 263 | 0.140 | 0.0685 | No |
| 4 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 731 | 0.103 | 0.0528 | No |
| 5 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 1411 | 0.077 | 0.0163 | No |
| 6 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 1805 | 0.067 | -0.0006 | No |
| 7 | AMPH | AMPH Entrez, Source | amphiphysin (Stiff-Man syndrome with breast cancer 128kDa autoantigen) | 1857 | 0.066 | 0.0081 | No |
| 8 | FGF2 | FGF2 Entrez, Source | fibroblast growth factor 2 (basic) | 2253 | 0.056 | -0.0110 | No |
| 9 | GPR176 | GPR176 Entrez, Source | G protein-coupled receptor 176 | 2354 | 0.054 | -0.0082 | No |
| 10 | PARD3 | PARD3 Entrez, Source | par-3 partitioning defective 3 homolog (C. elegans) | 2421 | 0.053 | -0.0030 | No |
| 11 | CTBP2 | CTBP2 Entrez, Source | C-terminal binding protein 2 | 2635 | 0.049 | -0.0097 | No |
| 12 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 2716 | 0.048 | -0.0067 | No |
| 13 | COL5A2 | COL5A2 Entrez, Source | collagen, type V, alpha 2 | 3169 | 0.041 | -0.0331 | No |
| 14 | ENC1 | ENC1 Entrez, Source | ectodermal-neural cortex (with BTB-like domain) | 3196 | 0.040 | -0.0274 | No |
| 15 | CAP2 | CAP2 Entrez, Source | CAP, adenylate cyclase-associated protein, 2 (yeast) | 3198 | 0.040 | -0.0199 | No |
| 16 | RGS7 | RGS7 Entrez, Source | regulator of G-protein signalling 7 | 3598 | 0.034 | -0.0435 | No |
| 17 | CUTL1 | CUTL1 Entrez, Source | cut-like 1, CCAAT displacement protein (Drosophila) | 4235 | 0.025 | -0.0866 | No |
| 18 | MDN1 | MDN1 Entrez, Source | MDN1, midasin homolog (yeast) | 4316 | 0.024 | -0.0880 | No |
| 19 | EIF4E | EIF4E Entrez, Source | eukaryotic translation initiation factor 4E | 4511 | 0.022 | -0.0985 | No |
| 20 | CBLB | CBLB Entrez, Source | Cas-Br-M (murine) ecotropic retroviral transforming sequence b | 4550 | 0.021 | -0.0974 | No |
| 21 | GULP1 | GULP1 Entrez, Source | GULP, engulfment adaptor PTB domain containing 1 | 4845 | 0.018 | -0.1161 | No |
| 22 | PRKCA | PRKCA Entrez, Source | protein kinase C, alpha | 4883 | 0.018 | -0.1155 | No |
| 23 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 5269 | 0.013 | -0.1421 | No |
| 24 | TSPAN5 | TSPAN5 Entrez, Source | tetraspanin 5 | 5710 | 0.008 | -0.1738 | No |
| 25 | PTPRM | PTPRM Entrez, Source | protein tyrosine phosphatase, receptor type, M | 5768 | 0.007 | -0.1767 | No |
| 26 | GBE1 | GBE1 Entrez, Source | glucan (1,4-alpha-), branching enzyme 1 (glycogen branching enzyme, Andersen disease, glycogen storage disease type IV) | 5875 | 0.006 | -0.1836 | No |
| 27 | FGF5 | FGF5 Entrez, Source | fibroblast growth factor 5 | 6286 | 0.001 | -0.2143 | No |
| 28 | GTF2F2 | GTF2F2 Entrez, Source | general transcription factor IIF, polypeptide 2, 30kDa | 6378 | 0.000 | -0.2210 | No |
| 29 | ZNF148 | ZNF148 Entrez, Source | zinc finger protein 148 | 6724 | -0.003 | -0.2465 | No |
| 30 | MYO1B | MYO1B Entrez, Source | myosin IB | 6819 | -0.004 | -0.2528 | No |
| 31 | SEPT9 | SEPT9 Entrez, Source | septin 9 | 6891 | -0.005 | -0.2573 | No |
| 32 | AFAP | AFAP Entrez, Source | - | 7125 | -0.007 | -0.2736 | No |
| 33 | TDRD3 | TDRD3 Entrez, Source | tudor domain containing 3 | 7165 | -0.007 | -0.2751 | No |
| 34 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 7336 | -0.009 | -0.2863 | No |
| 35 | CDC42BPA | CDC42BPA Entrez, Source | CDC42 binding protein kinase alpha (DMPK-like) | 7499 | -0.010 | -0.2965 | No |
| 36 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 7532 | -0.011 | -0.2968 | No |
| 37 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 7684 | -0.012 | -0.3059 | No |
| 38 | XRCC4 | XRCC4 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 4 | 7798 | -0.014 | -0.3119 | No |
| 39 | FLNB | FLNB Entrez, Source | filamin B, beta (actin binding protein 278) | 7935 | -0.015 | -0.3193 | No |
| 40 | STK39 | STK39 Entrez, Source | serine threonine kinase 39 (STE20/SPS1 homolog, yeast) | 8189 | -0.018 | -0.3350 | No |
| 41 | MGLL | MGLL Entrez, Source | monoglyceride lipase | 8545 | -0.021 | -0.3577 | No |
| 42 | EDG2 | EDG2 Entrez, Source | endothelial differentiation, lysophosphatidic acid G-protein-coupled receptor, 2 | 8571 | -0.022 | -0.3555 | No |
| 43 | ZMYND11 | ZMYND11 Entrez, Source | zinc finger, MYND domain containing 11 | 8725 | -0.023 | -0.3627 | No |
| 44 | ATP2B1 | ATP2B1 Entrez, Source | ATPase, Ca++ transporting, plasma membrane 1 | 8980 | -0.026 | -0.3769 | No |
| 45 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 9251 | -0.029 | -0.3919 | No |
| 46 | PKN2 | PKN2 Entrez, Source | protein kinase N2 | 9258 | -0.029 | -0.3869 | No |
| 47 | DPYD | DPYD Entrez, Source | dihydropyrimidine dehydrogenase | 9279 | -0.029 | -0.3829 | No |
| 48 | NFIB | NFIB Entrez, Source | nuclear factor I/B | 9610 | -0.033 | -0.4015 | No |
| 49 | MNAT1 | MNAT1 Entrez, Source | menage a trois homolog 1, cyclin H assembly factor (Xenopus laevis) | 9616 | -0.033 | -0.3957 | No |
| 50 | PRIM2A | PRIM2A Entrez, Source | primase, polypeptide 2A, 58kDa | 9718 | -0.034 | -0.3968 | No |
| 51 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 9911 | -0.036 | -0.4044 | No |
| 52 | WDR7 | WDR7 Entrez, Source | WD repeat domain 7 | 9968 | -0.037 | -0.4016 | No |
| 53 | GSK3B | GSK3B Entrez, Source | glycogen synthase kinase 3 beta | 10151 | -0.039 | -0.4079 | No |
| 54 | SKAP2 | SKAP2 Entrez, Source | src kinase associated phosphoprotein 2 | 10200 | -0.039 | -0.4040 | No |
| 55 | FARS2 | FARS2 Entrez, Source | phenylalanine-tRNA synthetase 2 (mitochondrial) | 10489 | -0.043 | -0.4176 | No |
| 56 | GNAQ | GNAQ Entrez, Source | guanine nucleotide binding protein (G protein), q polypeptide | 10825 | -0.048 | -0.4336 | No |
| 57 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 10869 | -0.049 | -0.4275 | No |
| 58 | PPP1R12A | PPP1R12A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 12A | 11026 | -0.051 | -0.4295 | No |
| 59 | SERPINB2 | SERPINB2 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 2 | 11162 | -0.054 | -0.4294 | No |
| 60 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 11598 | -0.062 | -0.4503 | Yes |
| 61 | PTK2 | PTK2 Entrez, Source | PTK2 protein tyrosine kinase 2 | 11607 | -0.063 | -0.4390 | Yes |
| 62 | FUT8 | FUT8 Entrez, Source | fucosyltransferase 8 (alpha (1,6) fucosyltransferase) | 11656 | -0.064 | -0.4305 | Yes |
| 63 | UBL3 | UBL3 Entrez, Source | ubiquitin-like 3 | 11808 | -0.068 | -0.4291 | Yes |
| 64 | SLC4A7 | SLC4A7 Entrez, Source | solute carrier family 4, sodium bicarbonate cotransporter, member 7 | 11826 | -0.068 | -0.4174 | Yes |
| 65 | CLASP2 | CLASP2 Entrez, Source | cytoplasmic linker associated protein 2 | 11968 | -0.072 | -0.4144 | Yes |
| 66 | PTPRK | PTPRK Entrez, Source | protein tyrosine phosphatase, receptor type, K | 11989 | -0.073 | -0.4021 | Yes |
| 67 | VEGFC | VEGFC Entrez, Source | vascular endothelial growth factor C | 12416 | -0.088 | -0.4176 | Yes |
| 68 | CASK | CASK Entrez, Source | calcium/calmodulin-dependent serine protein kinase (MAGUK family) | 12444 | -0.088 | -0.4028 | Yes |
| 69 | TRIO | TRIO Entrez, Source | triple functional domain (PTPRF interacting) | 12578 | -0.095 | -0.3947 | Yes |
| 70 | STK3 | STK3 Entrez, Source | serine/threonine kinase 3 (STE20 homolog, yeast) | 12679 | -0.101 | -0.3830 | Yes |
| 71 | BMP2K | BMP2K Entrez, Source | BMP2 inducible kinase | 12687 | -0.102 | -0.3641 | Yes |
| 72 | PDLIM5 | PDLIM5 Entrez, Source | PDZ and LIM domain 5 | 12904 | -0.122 | -0.3573 | Yes |
| 73 | NRG1 | NRG1 Entrez, Source | neuregulin 1 | 13047 | -0.140 | -0.3414 | Yes |
| 74 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 13119 | -0.153 | -0.3178 | Yes |
| 75 | ROCK2 | ROCK2 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 2 | 13146 | -0.160 | -0.2893 | Yes |
| 76 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 13153 | -0.163 | -0.2588 | Yes |
| 77 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 13195 | -0.185 | -0.2267 | Yes |
| 78 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13284 | -0.263 | -0.1834 | Yes |
| 79 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 13302 | -0.295 | -0.1287 | Yes |
| 80 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 13312 | -0.331 | -0.0666 | Yes |
| 81 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 13320 | -0.362 | 0.0017 | Yes |