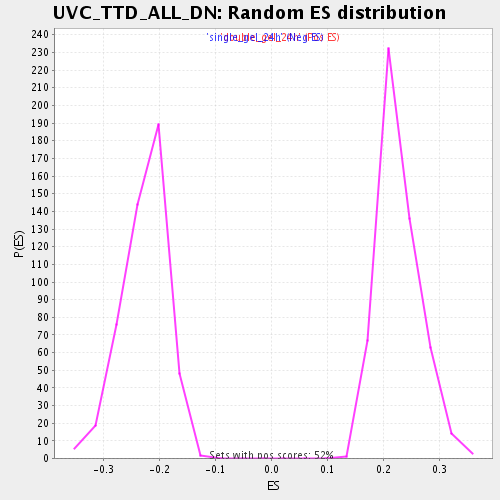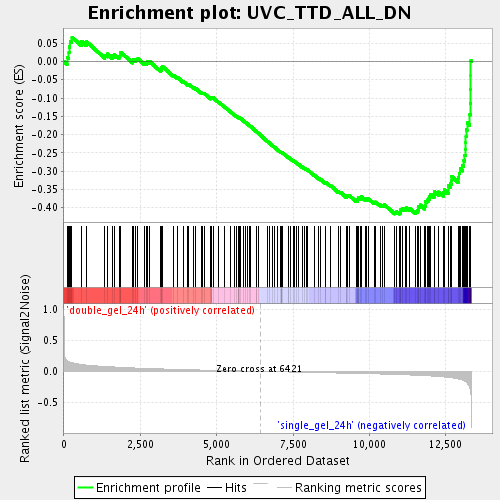
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | UVC_TTD_ALL_DN |
| Enrichment Score (ES) | -0.41741383 |
| Normalized Enrichment Score (NES) | -1.8396299 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.12163367 |
| FWER p-Value | 0.917 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CXCL12 | CXCL12 Entrez, Source | chemokine (C-X-C motif) ligand 12 (stromal cell-derived factor 1) | 105 | 0.175 | 0.0115 | No |
| 2 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 164 | 0.155 | 0.0244 | No |
| 3 | CREBBP | CREBBP Entrez, Source | CREB binding protein (Rubinstein-Taybi syndrome) | 179 | 0.152 | 0.0403 | No |
| 4 | IGF2BP3 | IGF2BP3 Entrez, Source | insulin-like growth factor 2 mRNA binding protein 3 | 199 | 0.149 | 0.0554 | No |
| 5 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 263 | 0.140 | 0.0662 | No |
| 6 | SLIT1 | SLIT1 Entrez, Source | slit homolog 1 (Drosophila) | 571 | 0.112 | 0.0554 | No |
| 7 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 731 | 0.103 | 0.0548 | No |
| 8 | HAS2 | HAS2 Entrez, Source | hyaluronan synthase 2 | 1343 | 0.079 | 0.0172 | No |
| 9 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 1411 | 0.077 | 0.0208 | No |
| 10 | HNRPDL | HNRPDL Entrez, Source | heterogeneous nuclear ribonucleoprotein D-like | 1582 | 0.073 | 0.0160 | No |
| 11 | ZHX3 | ZHX3 Entrez, Source | zinc fingers and homeoboxes 3 | 1652 | 0.071 | 0.0186 | No |
| 12 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 1805 | 0.067 | 0.0146 | No |
| 13 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 1837 | 0.067 | 0.0197 | No |
| 14 | AMPH | AMPH Entrez, Source | amphiphysin (Stiff-Man syndrome with breast cancer 128kDa autoantigen) | 1857 | 0.066 | 0.0256 | No |
| 15 | FGF2 | FGF2 Entrez, Source | fibroblast growth factor 2 (basic) | 2253 | 0.056 | 0.0019 | No |
| 16 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 2282 | 0.056 | 0.0060 | No |
| 17 | GPR176 | GPR176 Entrez, Source | G protein-coupled receptor 176 | 2354 | 0.054 | 0.0066 | No |
| 18 | PARD3 | PARD3 Entrez, Source | par-3 partitioning defective 3 homolog (C. elegans) | 2421 | 0.053 | 0.0076 | No |
| 19 | CTBP2 | CTBP2 Entrez, Source | C-terminal binding protein 2 | 2635 | 0.049 | -0.0031 | No |
| 20 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 2716 | 0.048 | -0.0039 | No |
| 21 | CRY1 | CRY1 Entrez, Source | cryptochrome 1 (photolyase-like) | 2740 | 0.047 | -0.0004 | No |
| 22 | MGAT5 | MGAT5 Entrez, Source | mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase | 2803 | 0.046 | 0.0001 | No |
| 23 | COL5A2 | COL5A2 Entrez, Source | collagen, type V, alpha 2 | 3169 | 0.041 | -0.0231 | No |
| 24 | ENC1 | ENC1 Entrez, Source | ectodermal-neural cortex (with BTB-like domain) | 3196 | 0.040 | -0.0206 | No |
| 25 | CAP2 | CAP2 Entrez, Source | CAP, adenylate cyclase-associated protein, 2 (yeast) | 3198 | 0.040 | -0.0162 | No |
| 26 | MARCH7 | MARCH7 Entrez, Source | membrane-associated ring finger (C3HC4) 7 | 3217 | 0.040 | -0.0132 | No |
| 27 | RGS7 | RGS7 Entrez, Source | regulator of G-protein signalling 7 | 3598 | 0.034 | -0.0382 | No |
| 28 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 3705 | 0.032 | -0.0426 | No |
| 29 | PEX14 | PEX14 Entrez, Source | peroxisomal biogenesis factor 14 | 3901 | 0.030 | -0.0541 | No |
| 30 | ARNT2 | ARNT2 Entrez, Source | aryl-hydrocarbon receptor nuclear translocator 2 | 4050 | 0.028 | -0.0623 | No |
| 31 | PTPN12 | PTPN12 Entrez, Source | protein tyrosine phosphatase, non-receptor type 12 | 4086 | 0.027 | -0.0619 | No |
| 32 | CUTL1 | CUTL1 Entrez, Source | cut-like 1, CCAAT displacement protein (Drosophila) | 4235 | 0.025 | -0.0703 | No |
| 33 | MDN1 | MDN1 Entrez, Source | MDN1, midasin homolog (yeast) | 4316 | 0.024 | -0.0736 | No |
| 34 | EIF4E | EIF4E Entrez, Source | eukaryotic translation initiation factor 4E | 4511 | 0.022 | -0.0859 | No |
| 35 | CBLB | CBLB Entrez, Source | Cas-Br-M (murine) ecotropic retroviral transforming sequence b | 4550 | 0.021 | -0.0864 | No |
| 36 | ITSN1 | ITSN1 Entrez, Source | intersectin 1 (SH3 domain protein) | 4584 | 0.021 | -0.0866 | No |
| 37 | ORC2L | ORC2L Entrez, Source | origin recognition complex, subunit 2-like (yeast) | 4798 | 0.019 | -0.1007 | No |
| 38 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 4824 | 0.018 | -0.1006 | No |
| 39 | PRKCBP1 | PRKCBP1 Entrez, Source | protein kinase C binding protein 1 | 4831 | 0.018 | -0.0990 | No |
| 40 | GULP1 | GULP1 Entrez, Source | GULP, engulfment adaptor PTB domain containing 1 | 4845 | 0.018 | -0.0980 | No |
| 41 | PRKCA | PRKCA Entrez, Source | protein kinase C, alpha | 4883 | 0.018 | -0.0988 | No |
| 42 | PHLPP | PHLPP Entrez, Source | PH domain and leucine rich repeat protein phosphatase | 5073 | 0.015 | -0.1115 | No |
| 43 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 5269 | 0.013 | -0.1248 | No |
| 44 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 5448 | 0.011 | -0.1371 | No |
| 45 | HOMER1 | HOMER1 Entrez, Source | homer homolog 1 (Drosophila) | 5598 | 0.009 | -0.1474 | No |
| 46 | RBM16 | RBM16 Entrez, Source | RNA binding motif protein 16 | 5648 | 0.009 | -0.1502 | No |
| 47 | TSPAN5 | TSPAN5 Entrez, Source | tetraspanin 5 | 5710 | 0.008 | -0.1539 | No |
| 48 | MYST3 | MYST3 Entrez, Source | MYST histone acetyltransferase (monocytic leukemia) 3 | 5716 | 0.008 | -0.1534 | No |
| 49 | RSN | RSN Entrez, Source | restin (Reed-Steinberg cell-expressed intermediate filament-associated protein) | 5729 | 0.008 | -0.1535 | No |
| 50 | SPOP | SPOP Entrez, Source | speckle-type POZ protein | 5734 | 0.008 | -0.1530 | No |
| 51 | PTPRM | PTPRM Entrez, Source | protein tyrosine phosphatase, receptor type, M | 5768 | 0.007 | -0.1547 | No |
| 52 | GBE1 | GBE1 Entrez, Source | glucan (1,4-alpha-), branching enzyme 1 (glycogen branching enzyme, Andersen disease, glycogen storage disease type IV) | 5875 | 0.006 | -0.1621 | No |
| 53 | ACVR2A | ACVR2A Entrez, Source | activin A receptor, type IIA | 5957 | 0.005 | -0.1677 | No |
| 54 | ATF2 | ATF2 Entrez, Source | activating transcription factor 2 | 6021 | 0.004 | -0.1719 | No |
| 55 | PUM2 | PUM2 Entrez, Source | pumilio homolog 2 (Drosophila) | 6077 | 0.004 | -0.1757 | No |
| 56 | RBM39 | RBM39 Entrez, Source | RNA binding motif protein 39 | 6107 | 0.004 | -0.1775 | No |
| 57 | FGF5 | FGF5 Entrez, Source | fibroblast growth factor 5 | 6286 | 0.001 | -0.1908 | No |
| 58 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 6305 | 0.001 | -0.1920 | No |
| 59 | GTF2F2 | GTF2F2 Entrez, Source | general transcription factor IIF, polypeptide 2, 30kDa | 6378 | 0.000 | -0.1975 | No |
| 60 | TUBGCP3 | TUBGCP3 Entrez, Source | tubulin, gamma complex associated protein 3 | 6657 | -0.003 | -0.2183 | No |
| 61 | ZNF148 | ZNF148 Entrez, Source | zinc finger protein 148 | 6724 | -0.003 | -0.2229 | No |
| 62 | MYO1B | MYO1B Entrez, Source | myosin IB | 6819 | -0.004 | -0.2296 | No |
| 63 | SEPT9 | SEPT9 Entrez, Source | septin 9 | 6891 | -0.005 | -0.2345 | No |
| 64 | ZNF292 | ZNF292 Entrez, Source | zinc finger protein 292 | 6983 | -0.005 | -0.2408 | No |
| 65 | CEP170 | CEP170 Entrez, Source | centrosomal protein 170kDa | 7086 | -0.006 | -0.2478 | No |
| 66 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 7107 | -0.007 | -0.2486 | No |
| 67 | AFAP | AFAP Entrez, Source | - | 7125 | -0.007 | -0.2491 | No |
| 68 | ABI1 | ABI1 Entrez, Source | abl-interactor 1 | 7128 | -0.007 | -0.2485 | No |
| 69 | TDRD3 | TDRD3 Entrez, Source | tudor domain containing 3 | 7165 | -0.007 | -0.2504 | No |
| 70 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 7336 | -0.009 | -0.2623 | No |
| 71 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 7360 | -0.009 | -0.2630 | No |
| 72 | MAP2K4 | MAP2K4 Entrez, Source | mitogen-activated protein kinase kinase 4 | 7416 | -0.010 | -0.2661 | No |
| 73 | CDC42BPA | CDC42BPA Entrez, Source | CDC42 binding protein kinase alpha (DMPK-like) | 7499 | -0.010 | -0.2712 | No |
| 74 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 7532 | -0.011 | -0.2724 | No |
| 75 | ARIH1 | ARIH1 Entrez, Source | ariadne homolog, ubiquitin-conjugating enzyme E2 binding protein, 1 (Drosophila) | 7627 | -0.012 | -0.2782 | No |
| 76 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 7684 | -0.012 | -0.2811 | No |
| 77 | XRCC4 | XRCC4 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 4 | 7798 | -0.014 | -0.2882 | No |
| 78 | ARHGEF7 | ARHGEF7 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 7 | 7877 | -0.014 | -0.2925 | No |
| 79 | FLNB | FLNB Entrez, Source | filamin B, beta (actin binding protein 278) | 7935 | -0.015 | -0.2951 | No |
| 80 | MSH3 | MSH3 Entrez, Source | mutS homolog 3 (E. coli) | 7967 | -0.015 | -0.2958 | No |
| 81 | STK39 | STK39 Entrez, Source | serine threonine kinase 39 (STE20/SPS1 homolog, yeast) | 8189 | -0.018 | -0.3106 | No |
| 82 | MTMR3 | MTMR3 Entrez, Source | myotubularin related protein 3 | 8338 | -0.019 | -0.3197 | No |
| 83 | NUP98 | NUP98 Entrez, Source | nucleoporin 98kDa | 8404 | -0.020 | -0.3224 | No |
| 84 | MGLL | MGLL Entrez, Source | monoglyceride lipase | 8545 | -0.021 | -0.3307 | No |
| 85 | EDG2 | EDG2 Entrez, Source | endothelial differentiation, lysophosphatidic acid G-protein-coupled receptor, 2 | 8571 | -0.022 | -0.3302 | No |
| 86 | ZMYND11 | ZMYND11 Entrez, Source | zinc finger, MYND domain containing 11 | 8725 | -0.023 | -0.3392 | No |
| 87 | ATP2B1 | ATP2B1 Entrez, Source | ATPase, Ca++ transporting, plasma membrane 1 | 8980 | -0.026 | -0.3556 | No |
| 88 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 9041 | -0.026 | -0.3572 | No |
| 89 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 9251 | -0.029 | -0.3699 | No |
| 90 | PKN2 | PKN2 Entrez, Source | protein kinase N2 | 9258 | -0.029 | -0.3671 | No |
| 91 | DPYD | DPYD Entrez, Source | dihydropyrimidine dehydrogenase | 9279 | -0.029 | -0.3654 | No |
| 92 | CEP350 | CEP350 Entrez, Source | centrosomal protein 350kDa | 9340 | -0.029 | -0.3667 | No |
| 93 | PTPN2 | PTPN2 Entrez, Source | protein tyrosine phosphatase, non-receptor type 2 | 9568 | -0.032 | -0.3803 | No |
| 94 | NFIB | NFIB Entrez, Source | nuclear factor I/B | 9610 | -0.033 | -0.3798 | No |
| 95 | MNAT1 | MNAT1 Entrez, Source | menage a trois homolog 1, cyclin H assembly factor (Xenopus laevis) | 9616 | -0.033 | -0.3765 | No |
| 96 | NIPBL | NIPBL Entrez, Source | Nipped-B homolog (Drosophila) | 9627 | -0.033 | -0.3736 | No |
| 97 | OGT | OGT Entrez, Source | O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) | 9640 | -0.033 | -0.3708 | No |
| 98 | PRIM2A | PRIM2A Entrez, Source | primase, polypeptide 2A, 58kDa | 9718 | -0.034 | -0.3728 | No |
| 99 | FUBP1 | FUBP1 Entrez, Source | far upstream element (FUSE) binding protein 1 | 9728 | -0.034 | -0.3697 | No |
| 100 | NCOA3 | NCOA3 Entrez, Source | nuclear receptor coactivator 3 | 9861 | -0.036 | -0.3757 | No |
| 101 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 9911 | -0.036 | -0.3754 | No |
| 102 | WDR7 | WDR7 Entrez, Source | WD repeat domain 7 | 9968 | -0.037 | -0.3756 | No |
| 103 | GSK3B | GSK3B Entrez, Source | glycogen synthase kinase 3 beta | 10151 | -0.039 | -0.3850 | No |
| 104 | SKAP2 | SKAP2 Entrez, Source | src kinase associated phosphoprotein 2 | 10200 | -0.039 | -0.3843 | No |
| 105 | CASP8 | CASP8 Entrez, Source | caspase 8, apoptosis-related cysteine peptidase | 10368 | -0.041 | -0.3923 | No |
| 106 | ZRF1 | ZRF1 Entrez, Source | zuotin related factor 1 | 10439 | -0.042 | -0.3929 | No |
| 107 | FARS2 | FARS2 Entrez, Source | phenylalanine-tRNA synthetase 2 (mitochondrial) | 10489 | -0.043 | -0.3918 | No |
| 108 | GNAQ | GNAQ Entrez, Source | guanine nucleotide binding protein (G protein), q polypeptide | 10825 | -0.048 | -0.4118 | Yes |
| 109 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 10869 | -0.049 | -0.4096 | Yes |
| 110 | PLEKHC1 | PLEKHC1 Entrez, Source | pleckstrin homology domain containing, family C (with FERM domain) member 1 | 10973 | -0.051 | -0.4118 | Yes |
| 111 | STRN3 | STRN3 Entrez, Source | striatin, calmodulin binding protein 3 | 11023 | -0.051 | -0.4098 | Yes |
| 112 | PPP1R12A | PPP1R12A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 12A | 11026 | -0.051 | -0.4042 | Yes |
| 113 | BARD1 | BARD1 Entrez, Source | BRCA1 associated RING domain 1 | 11085 | -0.053 | -0.4028 | Yes |
| 114 | SERPINB2 | SERPINB2 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 2 | 11162 | -0.054 | -0.4025 | Yes |
| 115 | CNOT2 | CNOT2 Entrez, Source | CCR4-NOT transcription complex, subunit 2 | 11214 | -0.055 | -0.4002 | Yes |
| 116 | MTF2 | MTF2 Entrez, Source | metal response element binding transcription factor 2 | 11313 | -0.057 | -0.4013 | Yes |
| 117 | REV3L | REV3L Entrez, Source | REV3-like, catalytic subunit of DNA polymerase zeta (yeast) | 11512 | -0.061 | -0.4096 | Yes |
| 118 | CNOT4 | CNOT4 Entrez, Source | CCR4-NOT transcription complex, subunit 4 | 11569 | -0.062 | -0.4069 | Yes |
| 119 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 11598 | -0.062 | -0.4021 | Yes |
| 120 | PTK2 | PTK2 Entrez, Source | PTK2 protein tyrosine kinase 2 | 11607 | -0.063 | -0.3958 | Yes |
| 121 | FUT8 | FUT8 Entrez, Source | fucosyltransferase 8 (alpha (1,6) fucosyltransferase) | 11656 | -0.064 | -0.3923 | Yes |
| 122 | UBL3 | UBL3 Entrez, Source | ubiquitin-like 3 | 11808 | -0.068 | -0.3962 | Yes |
| 123 | SLC4A7 | SLC4A7 Entrez, Source | solute carrier family 4, sodium bicarbonate cotransporter, member 7 | 11826 | -0.068 | -0.3899 | Yes |
| 124 | SPBC25 | SPBC25 Entrez, Source | spindle pole body component 25 homolog (S. cerevisiae) | 11828 | -0.068 | -0.3824 | Yes |
| 125 | PCTK2 | PCTK2 Entrez, Source | PCTAIRE protein kinase 2 | 11883 | -0.070 | -0.3788 | Yes |
| 126 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 11928 | -0.071 | -0.3743 | Yes |
| 127 | CLASP2 | CLASP2 Entrez, Source | cytoplasmic linker associated protein 2 | 11968 | -0.072 | -0.3692 | Yes |
| 128 | PTPRK | PTPRK Entrez, Source | protein tyrosine phosphatase, receptor type, K | 11989 | -0.073 | -0.3627 | Yes |
| 129 | TMEM23 | TMEM23 Entrez, Source | transmembrane protein 23 | 12115 | -0.076 | -0.3637 | Yes |
| 130 | IL1RAP | IL1RAP Entrez, Source | interleukin 1 receptor accessory protein | 12123 | -0.077 | -0.3556 | Yes |
| 131 | PARG | PARG Entrez, Source | poly (ADP-ribose) glycohydrolase | 12259 | -0.081 | -0.3568 | Yes |
| 132 | VEGFC | VEGFC Entrez, Source | vascular endothelial growth factor C | 12416 | -0.088 | -0.3589 | Yes |
| 133 | CASK | CASK Entrez, Source | calcium/calmodulin-dependent serine protein kinase (MAGUK family) | 12444 | -0.088 | -0.3511 | Yes |
| 134 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 12577 | -0.095 | -0.3505 | Yes |
| 135 | TRIO | TRIO Entrez, Source | triple functional domain (PTPRF interacting) | 12578 | -0.095 | -0.3399 | Yes |
| 136 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 12641 | -0.099 | -0.3336 | Yes |
| 137 | STK3 | STK3 Entrez, Source | serine/threonine kinase 3 (STE20 homolog, yeast) | 12679 | -0.101 | -0.3251 | Yes |
| 138 | BMP2K | BMP2K Entrez, Source | BMP2 inducible kinase | 12687 | -0.102 | -0.3143 | Yes |
| 139 | PDLIM5 | PDLIM5 Entrez, Source | PDZ and LIM domain 5 | 12904 | -0.122 | -0.3171 | Yes |
| 140 | SIPA1L1 | SIPA1L1 Entrez, Source | signal-induced proliferation-associated 1 like 1 | 12930 | -0.125 | -0.3051 | Yes |
| 141 | KIF2A | KIF2A Entrez, Source | kinesin heavy chain member 2A | 12962 | -0.127 | -0.2933 | Yes |
| 142 | NRG1 | NRG1 Entrez, Source | neuregulin 1 | 13047 | -0.140 | -0.2842 | Yes |
| 143 | RRAS2 | RRAS2 Entrez, Source | related RAS viral (r-ras) oncogene homolog 2 | 13088 | -0.147 | -0.2708 | Yes |
| 144 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 13119 | -0.153 | -0.2561 | Yes |
| 145 | ROCK2 | ROCK2 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 2 | 13146 | -0.160 | -0.2403 | Yes |
| 146 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 13153 | -0.163 | -0.2226 | Yes |
| 147 | SYNJ2 | SYNJ2 Entrez, Source | synaptojanin 2 | 13158 | -0.165 | -0.2045 | Yes |
| 148 | CLASP1 | CLASP1 Entrez, Source | cytoplasmic linker associated protein 1 | 13168 | -0.170 | -0.1863 | Yes |
| 149 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 13195 | -0.185 | -0.1677 | Yes |
| 150 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13284 | -0.263 | -0.1452 | Yes |
| 151 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 13302 | -0.295 | -0.1136 | Yes |
| 152 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 13312 | -0.331 | -0.0775 | Yes |
| 153 | GNAI1 | GNAI1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | 13318 | -0.354 | -0.0385 | Yes |
| 154 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 13320 | -0.362 | 0.0017 | Yes |