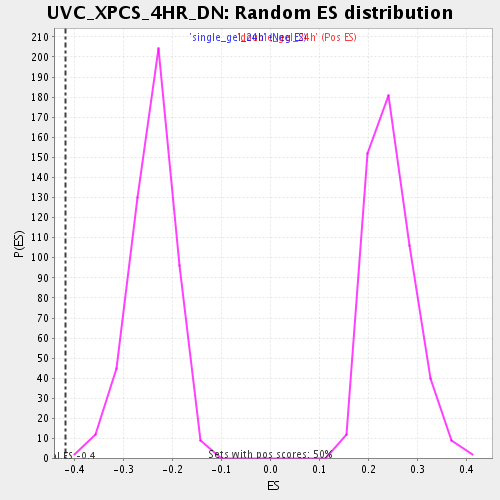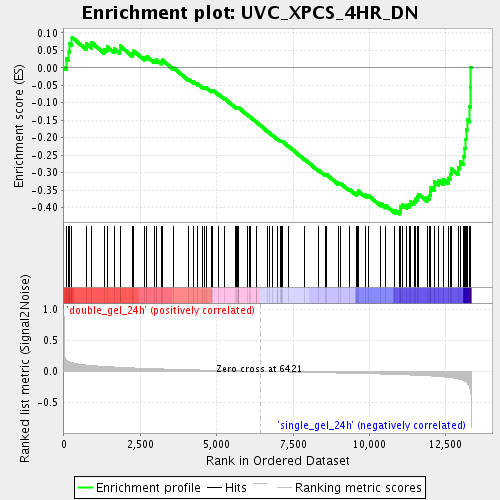
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | UVC_XPCS_4HR_DN |
| Enrichment Score (ES) | -0.4185939 |
| Normalized Enrichment Score (NES) | -1.7300035 |
| Nominal p-value | 0.002008032 |
| FDR q-value | 0.14672084 |
| FWER p-Value | 0.998 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IGF1R | IGF1R Entrez, Source | insulin-like growth factor 1 receptor | 76 | 0.189 | 0.0260 | No |
| 2 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 164 | 0.155 | 0.0456 | No |
| 3 | CREBBP | CREBBP Entrez, Source | CREB binding protein (Rubinstein-Taybi syndrome) | 179 | 0.152 | 0.0701 | No |
| 4 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 263 | 0.140 | 0.0874 | No |
| 5 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 731 | 0.103 | 0.0694 | No |
| 6 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 906 | 0.095 | 0.0723 | No |
| 7 | CENTB2 | CENTB2 Entrez, Source | centaurin, beta 2 | 1335 | 0.079 | 0.0533 | No |
| 8 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 1411 | 0.077 | 0.0606 | No |
| 9 | ZHX3 | ZHX3 Entrez, Source | zinc fingers and homeoboxes 3 | 1652 | 0.071 | 0.0545 | No |
| 10 | TGFBR3 | TGFBR3 Entrez, Source | transforming growth factor, beta receptor III (betaglycan, 300kDa) | 1835 | 0.067 | 0.0519 | No |
| 11 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 1837 | 0.067 | 0.0631 | No |
| 12 | FGF2 | FGF2 Entrez, Source | fibroblast growth factor 2 (basic) | 2253 | 0.056 | 0.0412 | No |
| 13 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 2282 | 0.056 | 0.0484 | No |
| 14 | CTBP2 | CTBP2 Entrez, Source | C-terminal binding protein 2 | 2635 | 0.049 | 0.0301 | No |
| 15 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 2716 | 0.048 | 0.0321 | No |
| 16 | KLF6 | KLF6 Entrez, Source | Kruppel-like factor 6 | 2960 | 0.044 | 0.0211 | No |
| 17 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 3035 | 0.043 | 0.0227 | No |
| 18 | WNT5A | WNT5A Entrez, Source | wingless-type MMTV integration site family, member 5A | 3195 | 0.040 | 0.0174 | No |
| 19 | MARCH7 | MARCH7 Entrez, Source | membrane-associated ring finger (C3HC4) 7 | 3217 | 0.040 | 0.0225 | No |
| 20 | RGS7 | RGS7 Entrez, Source | regulator of G-protein signalling 7 | 3598 | 0.034 | -0.0005 | No |
| 21 | PTPN12 | PTPN12 Entrez, Source | protein tyrosine phosphatase, non-receptor type 12 | 4086 | 0.027 | -0.0327 | No |
| 22 | CUTL1 | CUTL1 Entrez, Source | cut-like 1, CCAAT displacement protein (Drosophila) | 4235 | 0.025 | -0.0395 | No |
| 23 | E2F5 | E2F5 Entrez, Source | E2F transcription factor 5, p130-binding | 4371 | 0.024 | -0.0458 | No |
| 24 | CBLB | CBLB Entrez, Source | Cas-Br-M (murine) ecotropic retroviral transforming sequence b | 4550 | 0.021 | -0.0556 | No |
| 25 | USP15 | USP15 Entrez, Source | ubiquitin specific peptidase 15 | 4599 | 0.021 | -0.0558 | No |
| 26 | TIAM1 | TIAM1 Entrez, Source | T-cell lymphoma invasion and metastasis 1 | 4663 | 0.020 | -0.0572 | No |
| 27 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 4824 | 0.018 | -0.0662 | No |
| 28 | PRKCBP1 | PRKCBP1 Entrez, Source | protein kinase C binding protein 1 | 4831 | 0.018 | -0.0636 | No |
| 29 | EDD1 | EDD1 Entrez, Source | E3 ubiquitin protein ligase, HECT domain containing, 1 | 4878 | 0.018 | -0.0641 | No |
| 30 | PHLPP | PHLPP Entrez, Source | PH domain and leucine rich repeat protein phosphatase | 5073 | 0.015 | -0.0762 | No |
| 31 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 5269 | 0.013 | -0.0888 | No |
| 32 | PRR3 | PRR3 Entrez, Source | proline rich 3 | 5620 | 0.009 | -0.1137 | No |
| 33 | RBM16 | RBM16 Entrez, Source | RNA binding motif protein 16 | 5648 | 0.009 | -0.1143 | No |
| 34 | TERF1 | TERF1 Entrez, Source | telomeric repeat binding factor (NIMA-interacting) 1 | 5667 | 0.008 | -0.1143 | No |
| 35 | TSPAN5 | TSPAN5 Entrez, Source | tetraspanin 5 | 5710 | 0.008 | -0.1161 | No |
| 36 | MYST3 | MYST3 Entrez, Source | MYST histone acetyltransferase (monocytic leukemia) 3 | 5716 | 0.008 | -0.1152 | No |
| 37 | RSN | RSN Entrez, Source | restin (Reed-Steinberg cell-expressed intermediate filament-associated protein) | 5729 | 0.008 | -0.1148 | No |
| 38 | ATF2 | ATF2 Entrez, Source | activating transcription factor 2 | 6021 | 0.004 | -0.1361 | No |
| 39 | PUM2 | PUM2 Entrez, Source | pumilio homolog 2 (Drosophila) | 6077 | 0.004 | -0.1396 | No |
| 40 | RBM39 | RBM39 Entrez, Source | RNA binding motif protein 39 | 6107 | 0.004 | -0.1411 | No |
| 41 | FGF5 | FGF5 Entrez, Source | fibroblast growth factor 5 | 6286 | 0.001 | -0.1543 | No |
| 42 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 6305 | 0.001 | -0.1555 | No |
| 43 | TUBGCP3 | TUBGCP3 Entrez, Source | tubulin, gamma complex associated protein 3 | 6657 | -0.003 | -0.1816 | No |
| 44 | ZNF148 | ZNF148 Entrez, Source | zinc finger protein 148 | 6724 | -0.003 | -0.1860 | No |
| 45 | TRIM2 | TRIM2 Entrez, Source | tripartite motif-containing 2 | 6833 | -0.004 | -0.1935 | No |
| 46 | ZNF292 | ZNF292 Entrez, Source | zinc finger protein 292 | 6983 | -0.005 | -0.2039 | No |
| 47 | MECP2 | MECP2 Entrez, Source | methyl CpG binding protein 2 (Rett syndrome) | 6994 | -0.006 | -0.2037 | No |
| 48 | CEP170 | CEP170 Entrez, Source | centrosomal protein 170kDa | 7086 | -0.006 | -0.2095 | No |
| 49 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 7107 | -0.007 | -0.2098 | No |
| 50 | AFAP | AFAP Entrez, Source | - | 7125 | -0.007 | -0.2100 | No |
| 51 | CDC2L5 | CDC2L5 Entrez, Source | cell division cycle 2-like 5 (cholinesterase-related cell division controller) | 7141 | -0.007 | -0.2099 | No |
| 52 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 7360 | -0.009 | -0.2249 | No |
| 53 | ARHGEF7 | ARHGEF7 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 7 | 7877 | -0.014 | -0.2614 | No |
| 54 | MTMR3 | MTMR3 Entrez, Source | myotubularin related protein 3 | 8338 | -0.019 | -0.2929 | No |
| 55 | MARK3 | MARK3 Entrez, Source | MAP/microtubule affinity-regulating kinase 3 | 8547 | -0.021 | -0.3051 | No |
| 56 | GLRB | GLRB Entrez, Source | glycine receptor, beta | 8597 | -0.022 | -0.3051 | No |
| 57 | ATP2B1 | ATP2B1 Entrez, Source | ATPase, Ca++ transporting, plasma membrane 1 | 8980 | -0.026 | -0.3296 | No |
| 58 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 9041 | -0.026 | -0.3297 | No |
| 59 | CEP350 | CEP350 Entrez, Source | centrosomal protein 350kDa | 9340 | -0.029 | -0.3472 | No |
| 60 | CREB1 | CREB1 Entrez, Source | cAMP responsive element binding protein 1 | 9571 | -0.032 | -0.3592 | No |
| 61 | NFIB | NFIB Entrez, Source | nuclear factor I/B | 9610 | -0.033 | -0.3565 | No |
| 62 | NIPBL | NIPBL Entrez, Source | Nipped-B homolog (Drosophila) | 9627 | -0.033 | -0.3521 | No |
| 63 | NCOA3 | NCOA3 Entrez, Source | nuclear receptor coactivator 3 | 9861 | -0.036 | -0.3637 | No |
| 64 | APPBP2 | APPBP2 Entrez, Source | amyloid beta precursor protein (cytoplasmic tail) binding protein 2 | 9965 | -0.037 | -0.3653 | No |
| 65 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 10367 | -0.041 | -0.3886 | No |
| 66 | EIF2C2 | EIF2C2 Entrez, Source | eukaryotic translation initiation factor 2C, 2 | 10536 | -0.044 | -0.3939 | No |
| 67 | GNAQ | GNAQ Entrez, Source | guanine nucleotide binding protein (G protein), q polypeptide | 10825 | -0.048 | -0.4075 | No |
| 68 | PLEKHC1 | PLEKHC1 Entrez, Source | pleckstrin homology domain containing, family C (with FERM domain) member 1 | 10973 | -0.051 | -0.4101 | Yes |
| 69 | STRN3 | STRN3 Entrez, Source | striatin, calmodulin binding protein 3 | 11023 | -0.051 | -0.4051 | Yes |
| 70 | PPP1R12A | PPP1R12A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 12A | 11026 | -0.051 | -0.3966 | Yes |
| 71 | BARD1 | BARD1 Entrez, Source | BRCA1 associated RING domain 1 | 11085 | -0.053 | -0.3922 | Yes |
| 72 | CNOT2 | CNOT2 Entrez, Source | CCR4-NOT transcription complex, subunit 2 | 11214 | -0.055 | -0.3926 | Yes |
| 73 | MTF2 | MTF2 Entrez, Source | metal response element binding transcription factor 2 | 11313 | -0.057 | -0.3904 | Yes |
| 74 | RASA1 | RASA1 Entrez, Source | RAS p21 protein activator (GTPase activating protein) 1 | 11353 | -0.058 | -0.3836 | Yes |
| 75 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 11458 | -0.060 | -0.3814 | Yes |
| 76 | REV3L | REV3L Entrez, Source | REV3-like, catalytic subunit of DNA polymerase zeta (yeast) | 11512 | -0.061 | -0.3752 | Yes |
| 77 | CNOT4 | CNOT4 Entrez, Source | CCR4-NOT transcription complex, subunit 4 | 11569 | -0.062 | -0.3690 | Yes |
| 78 | PTK2 | PTK2 Entrez, Source | PTK2 protein tyrosine kinase 2 | 11607 | -0.063 | -0.3613 | Yes |
| 79 | PCTK2 | PCTK2 Entrez, Source | PCTAIRE protein kinase 2 | 11883 | -0.070 | -0.3703 | Yes |
| 80 | CLASP2 | CLASP2 Entrez, Source | cytoplasmic linker associated protein 2 | 11968 | -0.072 | -0.3646 | Yes |
| 81 | PTPRK | PTPRK Entrez, Source | protein tyrosine phosphatase, receptor type, K | 11989 | -0.073 | -0.3539 | Yes |
| 82 | BTRC | BTRC Entrez, Source | beta-transducin repeat containing | 12010 | -0.073 | -0.3431 | Yes |
| 83 | IL1RAP | IL1RAP Entrez, Source | interleukin 1 receptor accessory protein | 12123 | -0.077 | -0.3387 | Yes |
| 84 | BMPR1A | BMPR1A Entrez, Source | bone morphogenetic protein receptor, type IA | 12130 | -0.077 | -0.3262 | Yes |
| 85 | PARG | PARG Entrez, Source | poly (ADP-ribose) glycohydrolase | 12259 | -0.081 | -0.3222 | Yes |
| 86 | VEGFC | VEGFC Entrez, Source | vascular endothelial growth factor C | 12416 | -0.088 | -0.3192 | Yes |
| 87 | TRIO | TRIO Entrez, Source | triple functional domain (PTPRF interacting) | 12578 | -0.095 | -0.3153 | Yes |
| 88 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 12641 | -0.099 | -0.3033 | Yes |
| 89 | STK3 | STK3 Entrez, Source | serine/threonine kinase 3 (STE20 homolog, yeast) | 12679 | -0.101 | -0.2891 | Yes |
| 90 | AKAP9 | AKAP9 Entrez, Source | A kinase (PRKA) anchor protein (yotiao) 9 | 12906 | -0.122 | -0.2856 | Yes |
| 91 | KIF2A | KIF2A Entrez, Source | kinesin heavy chain member 2A | 12962 | -0.127 | -0.2684 | Yes |
| 92 | RRAS2 | RRAS2 Entrez, Source | related RAS viral (r-ras) oncogene homolog 2 | 13088 | -0.147 | -0.2531 | Yes |
| 93 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 13119 | -0.153 | -0.2296 | Yes |
| 94 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 13153 | -0.163 | -0.2047 | Yes |
| 95 | SOS2 | SOS2 Entrez, Source | son of sevenless homolog 2 (Drosophila) | 13162 | -0.167 | -0.1772 | Yes |
| 96 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 13195 | -0.185 | -0.1485 | Yes |
| 97 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13284 | -0.263 | -0.1110 | Yes |
| 98 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 13312 | -0.331 | -0.0574 | Yes |
| 99 | GNAI1 | GNAI1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | 13318 | -0.354 | 0.0018 | Yes |