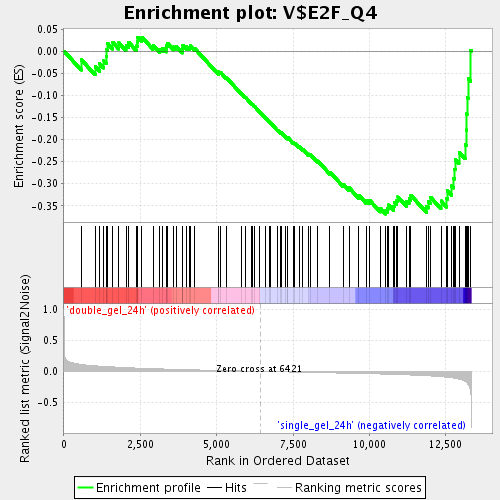
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | V$E2F_Q4 |
| Enrichment Score (ES) | -0.36862037 |
| Normalized Enrichment Score (NES) | -1.4967525 |
| Nominal p-value | 0.014227643 |
| FDR q-value | 0.27386925 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | KCND2 | KCND2 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 2 | 569 | 0.112 | -0.0196 | No |
| 2 | PAX6 | PAX6 Entrez, Source | paired box gene 6 (aniridia, keratitis) | 1024 | 0.091 | -0.0351 | No |
| 3 | MXD3 | MXD3 Entrez, Source | MAX dimerization protein 3 | 1165 | 0.085 | -0.0280 | No |
| 4 | SLCO3A1 | SLCO3A1 Entrez, Source | solute carrier organic anion transporter family, member 3A1 | 1303 | 0.080 | -0.0217 | No |
| 5 | EPHB1 | EPHB1 Entrez, Source | EPH receptor B1 | 1387 | 0.078 | -0.0118 | No |
| 6 | GRIA4 | GRIA4 Entrez, Source | glutamate receptor, ionotrophic, AMPA 4 | 1406 | 0.077 | 0.0029 | No |
| 7 | SLC38A1 | SLC38A1 Entrez, Source | solute carrier family 38, member 1 | 1420 | 0.077 | 0.0180 | No |
| 8 | DCK | DCK Entrez, Source | deoxycytidine kinase | 1598 | 0.072 | 0.0197 | No |
| 9 | KBTBD7 | KBTBD7 Entrez, Source | kelch repeat and BTB (POZ) domain containing 7 | 1799 | 0.067 | 0.0186 | No |
| 10 | HOXC10 | HOXC10 Entrez, Source | homeobox C10 | 2033 | 0.062 | 0.0138 | No |
| 11 | ILF3 | ILF3 Entrez, Source | interleukin enhancer binding factor 3, 90kDa | 2111 | 0.060 | 0.0204 | No |
| 12 | HNRPD | HNRPD Entrez, Source | heterogeneous nuclear ribonucleoprotein D (AU-rich element RNA binding protein 1, 37kDa) | 2376 | 0.054 | 0.0117 | No |
| 13 | SP3 | SP3 Entrez, Source | Sp3 transcription factor | 2400 | 0.054 | 0.0211 | No |
| 14 | FANCD2 | FANCD2 Entrez, Source | Fanconi anemia, complementation group D2 | 2414 | 0.053 | 0.0312 | No |
| 15 | HTF9C | HTF9C Entrez, Source | - | 2553 | 0.051 | 0.0313 | No |
| 16 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 2916 | 0.044 | 0.0132 | No |
| 17 | SFRS12 | SFRS12 Entrez, Source | splicing factor, arginine/serine-rich 12 | 3143 | 0.041 | 0.0047 | No |
| 18 | NRP2 | NRP2 Entrez, Source | neuropilin 2 | 3225 | 0.040 | 0.0068 | No |
| 19 | SLC6A4 | SLC6A4 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 | 3343 | 0.038 | 0.0058 | No |
| 20 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 3360 | 0.037 | 0.0123 | No |
| 21 | USP52 | USP52 Entrez, Source | ubiquitin specific peptidase 52 | 3390 | 0.037 | 0.0178 | No |
| 22 | SOAT1 | SOAT1 Entrez, Source | sterol O-acyltransferase (acyl-Coenzyme A: cholesterol acyltransferase) 1 | 3594 | 0.034 | 0.0095 | No |
| 23 | GABRB3 | GABRB3 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 3 | 3670 | 0.033 | 0.0106 | No |
| 24 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 3869 | 0.030 | 0.0019 | No |
| 25 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 3887 | 0.030 | 0.0068 | No |
| 26 | KCNA6 | KCNA6 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 6 | 3892 | 0.030 | 0.0127 | No |
| 27 | ATF5 | ATF5 Entrez, Source | activating transcription factor 5 | 4006 | 0.028 | 0.0101 | No |
| 28 | NCL | NCL Entrez, Source | nucleolin | 4105 | 0.027 | 0.0083 | No |
| 29 | EFNA5 | EFNA5 Entrez, Source | ephrin-A5 | 4128 | 0.027 | 0.0121 | No |
| 30 | RHD | RHD Entrez, Source | Rh blood group, D antigen | 4277 | 0.025 | 0.0062 | No |
| 31 | CSPG2 | CSPG2 Entrez, Source | chondroitin sulfate proteoglycan 2 (versican) | 5062 | 0.015 | -0.0498 | No |
| 32 | NELL2 | NELL2 Entrez, Source | NEL-like 2 (chicken) | 5064 | 0.015 | -0.0468 | No |
| 33 | CORT | CORT Entrez, Source | cortistatin | 5114 | 0.015 | -0.0474 | No |
| 34 | SFRS10 | SFRS10 Entrez, Source | splicing factor, arginine/serine-rich 10 (transformer 2 homolog, Drosophila) | 5320 | 0.012 | -0.0603 | No |
| 35 | HMGN2 | HMGN2 Entrez, Source | high-mobility group nucleosomal binding domain 2 | 5816 | 0.007 | -0.0963 | No |
| 36 | PVRL1 | PVRL1 Entrez, Source | poliovirus receptor-related 1 (herpesvirus entry mediator C; nectin) | 5943 | 0.005 | -0.1047 | No |
| 37 | UFD1L | UFD1L Entrez, Source | ubiquitin fusion degradation 1 like (yeast) | 6141 | 0.003 | -0.1189 | No |
| 38 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 6161 | 0.003 | -0.1198 | No |
| 39 | SUMO1 | SUMO1 Entrez, Source | SMT3 suppressor of mif two 3 homolog 1 (S. cerevisiae) | 6246 | 0.002 | -0.1257 | No |
| 40 | SLC9A5 | SLC9A5 Entrez, Source | solute carrier family 9 (sodium/hydrogen exchanger), member 5 | 6384 | 0.000 | -0.1360 | No |
| 41 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 6595 | -0.002 | -0.1514 | No |
| 42 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 6719 | -0.003 | -0.1601 | No |
| 43 | HCN3 | HCN3 Entrez, Source | hyperpolarization activated cyclic nucleotide-gated potassium channel 3 | 6747 | -0.003 | -0.1614 | No |
| 44 | PHF5A | PHF5A Entrez, Source | PHD finger protein 5A | 6991 | -0.006 | -0.1786 | No |
| 45 | RAB11B | RAB11B Entrez, Source | RAB11B, member RAS oncogene family | 7101 | -0.007 | -0.1855 | No |
| 46 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 7107 | -0.007 | -0.1844 | No |
| 47 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 7235 | -0.008 | -0.1924 | No |
| 48 | NUP62 | NUP62 Entrez, Source | nucleoporin 62kDa | 7323 | -0.009 | -0.1971 | No |
| 49 | DNMT1 | DNMT1 Entrez, Source | DNA (cytosine-5-)-methyltransferase 1 | 7331 | -0.009 | -0.1958 | No |
| 50 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 7506 | -0.010 | -0.2067 | No |
| 51 | SMAD6 | SMAD6 Entrez, Source | SMAD, mothers against DPP homolog 6 (Drosophila) | 7552 | -0.011 | -0.2079 | No |
| 52 | GSPT1 | GSPT1 Entrez, Source | G1 to S phase transition 1 | 7697 | -0.012 | -0.2162 | No |
| 53 | NOLC1 | NOLC1 Entrez, Source | nucleolar and coiled-body phosphoprotein 1 | 7821 | -0.014 | -0.2226 | No |
| 54 | PAQR4 | PAQR4 Entrez, Source | progestin and adipoQ receptor family member IV | 7999 | -0.016 | -0.2327 | No |
| 55 | PCSK1 | PCSK1 Entrez, Source | proprotein convertase subtilisin/kexin type 1 | 8055 | -0.016 | -0.2335 | No |
| 56 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 8303 | -0.019 | -0.2482 | No |
| 57 | KPNB1 | KPNB1 Entrez, Source | karyopherin (importin) beta 1 | 8707 | -0.023 | -0.2738 | No |
| 58 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 9143 | -0.027 | -0.3010 | No |
| 59 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 9331 | -0.029 | -0.3090 | No |
| 60 | MRPL40 | MRPL40 Entrez, Source | mitochondrial ribosomal protein L40 | 9651 | -0.033 | -0.3261 | No |
| 61 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 9894 | -0.036 | -0.3369 | No |
| 62 | ACBD6 | ACBD6 Entrez, Source | acyl-Coenzyme A binding domain containing 6 | 10006 | -0.037 | -0.3376 | No |
| 63 | PPP1CC | PPP1CC Entrez, Source | protein phosphatase 1, catalytic subunit, gamma isoform | 10356 | -0.041 | -0.3553 | No |
| 64 | HNRPA1 | HNRPA1 Entrez, Source | heterogeneous nuclear ribonucleoprotein A1 | 10533 | -0.044 | -0.3595 | Yes |
| 65 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 10597 | -0.045 | -0.3550 | Yes |
| 66 | ALDH6A1 | ALDH6A1 Entrez, Source | aldehyde dehydrogenase 6 family, member A1 | 10628 | -0.045 | -0.3479 | Yes |
| 67 | FMO4 | FMO4 Entrez, Source | flavin containing monooxygenase 4 | 10796 | -0.048 | -0.3505 | Yes |
| 68 | HNRPR | HNRPR Entrez, Source | heterogeneous nuclear ribonucleoprotein R | 10811 | -0.048 | -0.3416 | Yes |
| 69 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 10896 | -0.050 | -0.3376 | Yes |
| 70 | CRK | CRK Entrez, Source | v-crk sarcoma virus CT10 oncogene homolog (avian) | 10930 | -0.050 | -0.3297 | Yes |
| 71 | ACO2 | ACO2 Entrez, Source | aconitase 2, mitochondrial | 11218 | -0.055 | -0.3399 | Yes |
| 72 | RQCD1 | RQCD1 Entrez, Source | RCD1 required for cell differentiation1 homolog (S. pombe) | 11296 | -0.057 | -0.3339 | Yes |
| 73 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 11355 | -0.058 | -0.3263 | Yes |
| 74 | XTP3TPA | XTP3TPA Entrez, Source | - | 11867 | -0.069 | -0.3506 | Yes |
| 75 | UXT | UXT Entrez, Source | ubiquitously-expressed transcript | 11925 | -0.070 | -0.3402 | Yes |
| 76 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 11994 | -0.073 | -0.3303 | Yes |
| 77 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 12342 | -0.085 | -0.3388 | Yes |
| 78 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 12529 | -0.093 | -0.3335 | Yes |
| 79 | UGCGL1 | UGCGL1 Entrez, Source | UDP-glucose ceramide glucosyltransferase-like 1 | 12555 | -0.094 | -0.3159 | Yes |
| 80 | POLD3 | POLD3 Entrez, Source | polymerase (DNA-directed), delta 3, accessory subunit | 12692 | -0.103 | -0.3048 | Yes |
| 81 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 12758 | -0.107 | -0.2874 | Yes |
| 82 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 12787 | -0.110 | -0.2668 | Yes |
| 83 | TMPO | TMPO Entrez, Source | thymopoietin | 12812 | -0.111 | -0.2455 | Yes |
| 84 | ABI2 | ABI2 Entrez, Source | abl interactor 2 | 12931 | -0.125 | -0.2285 | Yes |
| 85 | POLA2 | POLA2 Entrez, Source | polymerase (DNA directed), alpha 2 (70kD subunit) | 13133 | -0.158 | -0.2110 | Yes |
| 86 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 13174 | -0.172 | -0.1783 | Yes |
| 87 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 13185 | -0.179 | -0.1420 | Yes |
| 88 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13196 | -0.186 | -0.1041 | Yes |
| 89 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 13232 | -0.213 | -0.0625 | Yes |
| 90 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 13314 | -0.341 | 0.0021 | Yes |

