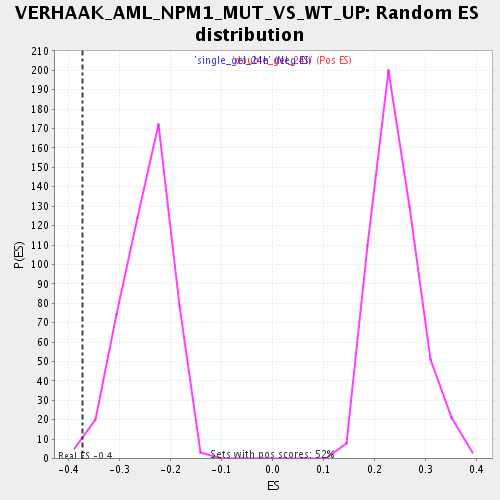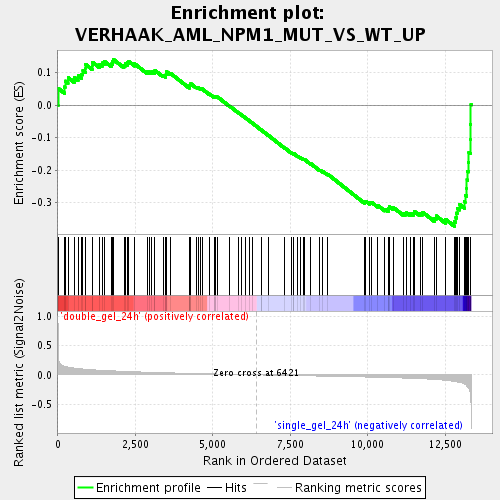
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | VERHAAK_AML_NPM1_MUT_VS_WT_UP |
| Enrichment Score (ES) | -0.37282225 |
| Normalized Enrichment Score (NES) | -1.5094647 |
| Nominal p-value | 0.008385744 |
| FDR q-value | 0.27471137 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PHLDA1 | PHLDA1 Entrez, Source | pleckstrin homology-like domain, family A, member 1 | 6 | 0.386 | 0.0516 | No |
| 2 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 215 | 0.147 | 0.0557 | No |
| 3 | AQP9 | AQP9 Entrez, Source | aquaporin 9 | 232 | 0.144 | 0.0739 | No |
| 4 | CD14 | CD14 Entrez, Source | CD14 molecule | 327 | 0.132 | 0.0847 | No |
| 5 | MMP2 | MMP2 Entrez, Source | matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase) | 527 | 0.115 | 0.0851 | No |
| 6 | PDGFD | PDGFD Entrez, Source | platelet derived growth factor D | 652 | 0.107 | 0.0902 | No |
| 7 | CX3CR1 | CX3CR1 Entrez, Source | chemokine (C-X3-C motif) receptor 1 | 768 | 0.101 | 0.0951 | No |
| 8 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 806 | 0.099 | 0.1058 | No |
| 9 | IER3 | IER3 Entrez, Source | immediate early response 3 | 890 | 0.096 | 0.1124 | No |
| 10 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 895 | 0.096 | 0.1250 | No |
| 11 | CD86 | CD86 Entrez, Source | CD86 molecule | 1107 | 0.087 | 0.1208 | No |
| 12 | STS | STS Entrez, Source | steroid sulfatase (microsomal), arylsulfatase C, isozyme S | 1112 | 0.087 | 0.1322 | No |
| 13 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 1348 | 0.079 | 0.1251 | No |
| 14 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 1427 | 0.077 | 0.1296 | No |
| 15 | PLEK | PLEK Entrez, Source | pleckstrin | 1498 | 0.075 | 0.1344 | No |
| 16 | CARD9 | CARD9 Entrez, Source | caspase recruitment domain family, member 9 | 1727 | 0.069 | 0.1265 | No |
| 17 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 1761 | 0.068 | 0.1332 | No |
| 18 | VNN1 | VNN1 Entrez, Source | vanin 1 | 1780 | 0.068 | 0.1411 | No |
| 19 | ADCY2 | ADCY2 Entrez, Source | adenylate cyclase 2 (brain) | 2134 | 0.059 | 0.1224 | No |
| 20 | HOXA4 | HOXA4 Entrez, Source | homeobox A4 | 2169 | 0.058 | 0.1277 | No |
| 21 | SECTM1 | SECTM1 Entrez, Source | secreted and transmembrane 1 | 2239 | 0.057 | 0.1301 | No |
| 22 | PTAFR | PTAFR Entrez, Source | platelet-activating factor receptor | 2286 | 0.056 | 0.1342 | No |
| 23 | C5AR1 | C5AR1 Entrez, Source | complement component 5a receptor 1 | 2477 | 0.052 | 0.1268 | No |
| 24 | SORT1 | SORT1 Entrez, Source | sortilin 1 | 2881 | 0.045 | 0.1025 | No |
| 25 | SLC15A3 | SLC15A3 Entrez, Source | solute carrier family 15, member 3 | 2965 | 0.044 | 0.1021 | No |
| 26 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 3035 | 0.043 | 0.1026 | No |
| 27 | RHAG | RHAG Entrez, Source | Rh-associated glycoprotein | 3123 | 0.041 | 0.1016 | No |
| 28 | CAT | CAT Entrez, Source | catalase | 3129 | 0.041 | 0.1068 | No |
| 29 | EREG | EREG Entrez, Source | epiregulin | 3391 | 0.037 | 0.0920 | No |
| 30 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 3481 | 0.035 | 0.0901 | No |
| 31 | TLR4 | TLR4 Entrez, Source | toll-like receptor 4 | 3487 | 0.035 | 0.0945 | No |
| 32 | KCNJ2 | KCNJ2 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 2 | 3503 | 0.035 | 0.0981 | No |
| 33 | PPBP | PPBP Entrez, Source | pro-platelet basic protein (chemokine (C-X-C motif) ligand 7) | 3511 | 0.035 | 0.1023 | No |
| 34 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 3636 | 0.033 | 0.0974 | No |
| 35 | SEPP1 | SEPP1 Entrez, Source | selenoprotein P, plasma, 1 | 4239 | 0.025 | 0.0554 | No |
| 36 | DEFB1 | DEFB1 Entrez, Source | defensin, beta 1 | 4246 | 0.025 | 0.0584 | No |
| 37 | FGL2 | FGL2 Entrez, Source | fibrinogen-like 2 | 4253 | 0.025 | 0.0613 | No |
| 38 | PLA2G4A | PLA2G4A Entrez, Source | phospholipase A2, group IVA (cytosolic, calcium-dependent) | 4288 | 0.025 | 0.0621 | No |
| 39 | TCIRG1 | TCIRG1 Entrez, Source | T-cell, immune regulator 1, ATPase, H+ transporting, lysosomal V0 subunit A3 | 4289 | 0.025 | 0.0655 | No |
| 40 | PELI1 | PELI1 Entrez, Source | pellino homolog 1 (Drosophila) | 4464 | 0.022 | 0.0553 | No |
| 41 | C3AR1 | C3AR1 Entrez, Source | complement component 3a receptor 1 | 4527 | 0.021 | 0.0535 | No |
| 42 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 4601 | 0.021 | 0.0508 | No |
| 43 | TBXAS1 | TBXAS1 Entrez, Source | thromboxane A synthase 1 (platelet, cytochrome P450, family 5, subfamily A) | 4650 | 0.020 | 0.0499 | No |
| 44 | C1QB | C1QB Entrez, Source | complement component 1, q subcomponent, B chain | 4891 | 0.018 | 0.0341 | No |
| 45 | HOMER3 | HOMER3 Entrez, Source | homer homolog 3 (Drosophila) | 5052 | 0.015 | 0.0241 | No |
| 46 | CSPG2 | CSPG2 Entrez, Source | chondroitin sulfate proteoglycan 2 (versican) | 5062 | 0.015 | 0.0255 | No |
| 47 | HCK | HCK Entrez, Source | hemopoietic cell kinase | 5080 | 0.015 | 0.0263 | No |
| 48 | SERPINA1 | SERPINA1 Entrez, Source | serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 1 | 5098 | 0.015 | 0.0270 | No |
| 49 | TRIB1 | TRIB1 Entrez, Source | tribbles homolog 1 (Drosophila) | 5150 | 0.014 | 0.0251 | No |
| 50 | HNMT | HNMT Entrez, Source | histamine N-methyltransferase | 5550 | 0.010 | -0.0038 | No |
| 51 | DPEP2 | DPEP2 Entrez, Source | dipeptidase 2 | 5824 | 0.006 | -0.0235 | No |
| 52 | FGR | FGR Entrez, Source | Gardner-Rasheed feline sarcoma viral (v-fgr) oncogene homolog | 5917 | 0.005 | -0.0297 | No |
| 53 | FBP1 | FBP1 Entrez, Source | fructose-1,6-bisphosphatase 1 | 6067 | 0.004 | -0.0405 | No |
| 54 | SLC16A3 | SLC16A3 Entrez, Source | solute carrier family 16, member 3 (monocarboxylic acid transporter 4) | 6179 | 0.003 | -0.0485 | No |
| 55 | CTSL | CTSL Entrez, Source | cathepsin L | 6275 | 0.002 | -0.0555 | No |
| 56 | LMNA | LMNA Entrez, Source | lamin A/C | 6554 | -0.001 | -0.0763 | No |
| 57 | PF4 | PF4 Entrez, Source | platelet factor 4 (chemokine (C-X-C motif) ligand 4) | 6577 | -0.002 | -0.0777 | No |
| 58 | C1QA | C1QA Entrez, Source | complement component 1, q subcomponent, A chain | 6809 | -0.004 | -0.0946 | No |
| 59 | SNX10 | SNX10 Entrez, Source | sorting nexin 10 | 7325 | -0.009 | -0.1323 | No |
| 60 | CTSD | CTSD Entrez, Source | cathepsin D (lysosomal aspartyl peptidase) | 7549 | -0.011 | -0.1477 | No |
| 61 | NGFRAP1 | NGFRAP1 Entrez, Source | nerve growth factor receptor (TNFRSF16) associated protein 1 | 7599 | -0.012 | -0.1499 | No |
| 62 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 7608 | -0.012 | -0.1489 | No |
| 63 | NINJ1 | NINJ1 Entrez, Source | ninjurin 1 | 7747 | -0.013 | -0.1576 | No |
| 64 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 7827 | -0.014 | -0.1617 | No |
| 65 | SCPEP1 | SCPEP1 Entrez, Source | serine carboxypeptidase 1 | 7920 | -0.015 | -0.1666 | No |
| 66 | MAFB | MAFB Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog B (avian) | 7949 | -0.015 | -0.1667 | No |
| 67 | C2 | C2 Entrez, Source | complement component 2 | 8163 | -0.017 | -0.1805 | No |
| 68 | TNF | TNF Entrez, Source | tumor necrosis factor (TNF superfamily, member 2) | 8457 | -0.020 | -0.1999 | No |
| 69 | LAPTM4B | LAPTM4B Entrez, Source | lysosomal associated protein transmembrane 4 beta | 8548 | -0.021 | -0.2038 | No |
| 70 | HIP1 | HIP1 Entrez, Source | huntingtin interacting protein 1 | 8701 | -0.023 | -0.2122 | No |
| 71 | PRKAR2B | PRKAR2B Entrez, Source | protein kinase, cAMP-dependent, regulatory, type II, beta | 9892 | -0.036 | -0.2972 | No |
| 72 | GGT1 | GGT1 Entrez, Source | gamma-glutamyltransferase 1 | 9932 | -0.036 | -0.2953 | No |
| 73 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 10063 | -0.038 | -0.3000 | No |
| 74 | CAST | CAST Entrez, Source | calpastatin | 10118 | -0.039 | -0.2988 | No |
| 75 | CAMK1 | CAMK1 Entrez, Source | calcium/calmodulin-dependent protein kinase I | 10322 | -0.041 | -0.3087 | No |
| 76 | BCL2A1 | BCL2A1 Entrez, Source | BCL2-related protein A1 | 10553 | -0.044 | -0.3201 | No |
| 77 | CLU | CLU Entrez, Source | clusterin | 10667 | -0.046 | -0.3224 | No |
| 78 | CCR1 | CCR1 Entrez, Source | chemokine (C-C motif) receptor 1 | 10676 | -0.046 | -0.3169 | No |
| 79 | IFI30 | IFI30 Entrez, Source | interferon, gamma-inducible protein 30 | 10707 | -0.046 | -0.3129 | No |
| 80 | GNS | GNS Entrez, Source | glucosamine (N-acetyl)-6-sulfatase (Sanfilippo disease IIID) | 10834 | -0.049 | -0.3158 | No |
| 81 | SERPINB2 | SERPINB2 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 2 | 11162 | -0.054 | -0.3332 | No |
| 82 | IGF2R | IGF2R Entrez, Source | insulin-like growth factor 2 receptor | 11234 | -0.056 | -0.3311 | No |
| 83 | SMPDL3A | SMPDL3A Entrez, Source | sphingomyelin phosphodiesterase, acid-like 3A | 11365 | -0.058 | -0.3330 | No |
| 84 | PRKCD | PRKCD Entrez, Source | protein kinase C, delta | 11467 | -0.060 | -0.3326 | No |
| 85 | MARCKS | MARCKS Entrez, Source | myristoylated alanine-rich protein kinase C substrate | 11501 | -0.060 | -0.3269 | No |
| 86 | QPRT | QPRT Entrez, Source | quinolinate phosphoribosyltransferase (nicotinate-nucleotide pyrophosphorylase (carboxylating)) | 11690 | -0.064 | -0.3324 | No |
| 87 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 11771 | -0.066 | -0.3295 | No |
| 88 | TMEM176B | TMEM176B Entrez, Source | transmembrane protein 176B | 12155 | -0.078 | -0.3479 | No |
| 89 | IL1B | IL1B Entrez, Source | interleukin 1, beta | 12206 | -0.080 | -0.3410 | No |
| 90 | CCL4 | CCL4 Entrez, Source | chemokine (C-C motif) ligand 4 | 12509 | -0.092 | -0.3514 | No |
| 91 | NFKB2 | NFKB2 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) | 12794 | -0.110 | -0.3580 | Yes |
| 92 | LGALS3BP | LGALS3BP Entrez, Source | lectin, galactoside-binding, soluble, 3 binding protein | 12844 | -0.115 | -0.3462 | Yes |
| 93 | ABCC4 | ABCC4 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 4 | 12867 | -0.117 | -0.3320 | Yes |
| 94 | MVP | MVP Entrez, Source | major vault protein | 12888 | -0.120 | -0.3174 | Yes |
| 95 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 12964 | -0.127 | -0.3059 | Yes |
| 96 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 13119 | -0.153 | -0.2969 | Yes |
| 97 | BASP1 | BASP1 Entrez, Source | brain abundant, membrane attached signal protein 1 | 13143 | -0.159 | -0.2771 | Yes |
| 98 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 13195 | -0.185 | -0.2560 | Yes |
| 99 | LGALS2 | LGALS2 Entrez, Source | lectin, galactoside-binding, soluble, 2 (galectin 2) | 13202 | -0.192 | -0.2304 | Yes |
| 100 | EMR1 | EMR1 Entrez, Source | egf-like module containing, mucin-like, hormone receptor-like 1 | 13220 | -0.205 | -0.2040 | Yes |
| 101 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 13248 | -0.219 | -0.1765 | Yes |
| 102 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 13262 | -0.231 | -0.1463 | Yes |
| 103 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 13310 | -0.321 | -0.1064 | Yes |
| 104 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 13317 | -0.347 | -0.0601 | Yes |
| 105 | S100A6 | S100A6 Entrez, Source | S100 calcium binding protein A6 | 13330 | -0.458 | 0.0009 | Yes |