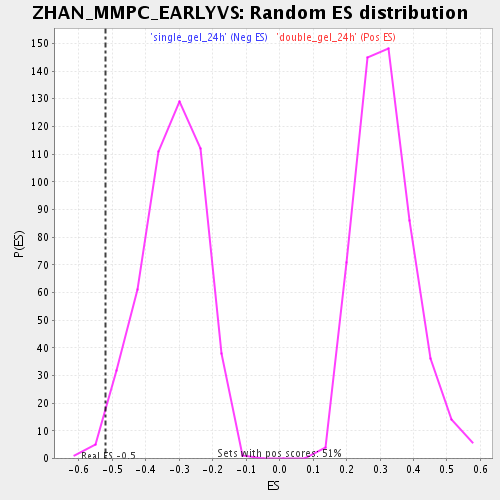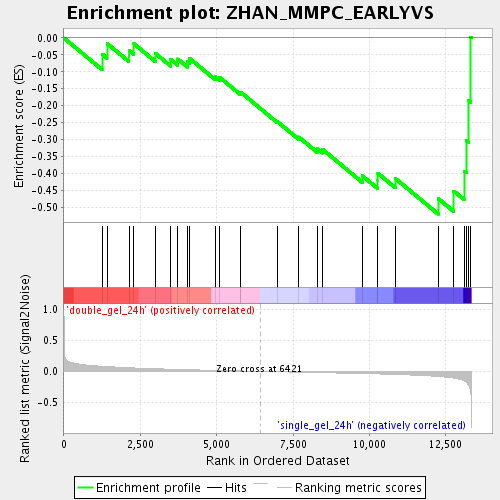
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_24h_versus_single_gel_24h.class.cls #double_gel_24h_versus_single_gel_24h_repos |
| Phenotype | class.cls#double_gel_24h_versus_single_gel_24h_repos |
| Upregulated in class | single_gel_24h |
| GeneSet | ZHAN_MMPC_EARLYVS |
| Enrichment Score (ES) | -0.5208115 |
| Normalized Enrichment Score (NES) | -1.6268631 |
| Nominal p-value | 0.010204081 |
| FDR q-value | 0.21057245 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | XBP1 | XBP1 Entrez, Source | X-box binding protein 1 | 1253 | 0.082 | -0.0479 | No |
| 2 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 1411 | 0.077 | -0.0161 | No |
| 3 | IL4R | IL4R Entrez, Source | interleukin 4 receptor | 2130 | 0.059 | -0.0366 | No |
| 4 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 2289 | 0.056 | -0.0171 | No |
| 5 | CIITA | CIITA Entrez, Source | class II, major histocompatibility complex, transactivator | 2991 | 0.043 | -0.0454 | No |
| 6 | BLK | BLK Entrez, Source | B lymphoid tyrosine kinase | 3494 | 0.035 | -0.0631 | No |
| 7 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 3724 | 0.032 | -0.0622 | No |
| 8 | BLR1 | BLR1 Entrez, Source | Burkitt lymphoma receptor 1, GTP binding protein (chemokine (C-X-C motif) receptor 5) | 4045 | 0.028 | -0.0706 | No |
| 9 | CCNF | CCNF Entrez, Source | cyclin F | 4114 | 0.027 | -0.0605 | No |
| 10 | RHOH | RHOH Entrez, Source | ras homolog gene family, member H | 4964 | 0.016 | -0.1151 | No |
| 11 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 5100 | 0.015 | -0.1168 | No |
| 12 | STCH | STCH Entrez, Source | stress 70 protein chaperone, microsome-associated, 60kDa | 5788 | 0.007 | -0.1645 | No |
| 13 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 5790 | 0.007 | -0.1607 | No |
| 14 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 6975 | -0.005 | -0.2466 | No |
| 15 | OGG1 | OGG1 Entrez, Source | 8-oxoguanine DNA glycosylase | 7683 | -0.012 | -0.2928 | No |
| 16 | ABCD3 | ABCD3 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 3 | 8286 | -0.019 | -0.3275 | No |
| 17 | TNF | TNF Entrez, Source | tumor necrosis factor (TNF superfamily, member 2) | 8457 | -0.020 | -0.3289 | No |
| 18 | RAE1 | RAE1 Entrez, Source | RAE1 RNA export 1 homolog (S. pombe) | 9764 | -0.035 | -0.4074 | No |
| 19 | RHOG | RHOG Entrez, Source | ras homolog gene family, member G (rho G) | 10269 | -0.040 | -0.4226 | No |
| 20 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 10273 | -0.040 | -0.4001 | No |
| 21 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 10848 | -0.049 | -0.4156 | No |
| 22 | ELF1 | ELF1 Entrez, Source | E74-like factor 1 (ets domain transcription factor) | 12250 | -0.081 | -0.4751 | Yes |
| 23 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 12758 | -0.107 | -0.4526 | Yes |
| 24 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 13097 | -0.150 | -0.3934 | Yes |
| 25 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 13167 | -0.169 | -0.3033 | Yes |
| 26 | LTB | LTB Entrez, Source | lymphotoxin beta (TNF superfamily, member 3) | 13250 | -0.220 | -0.1852 | Yes |
| 27 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 13314 | -0.341 | 0.0021 | Yes |