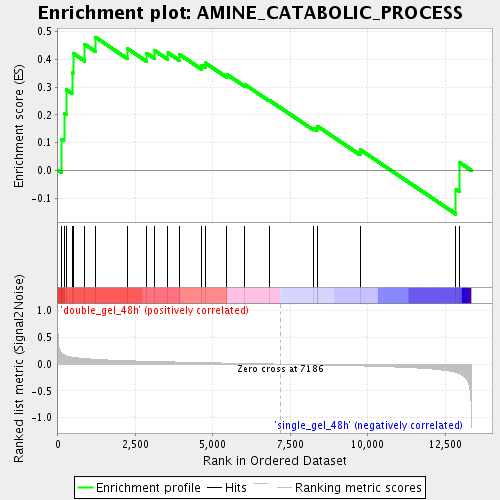
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | AMINE_CATABOLIC_PROCESS |
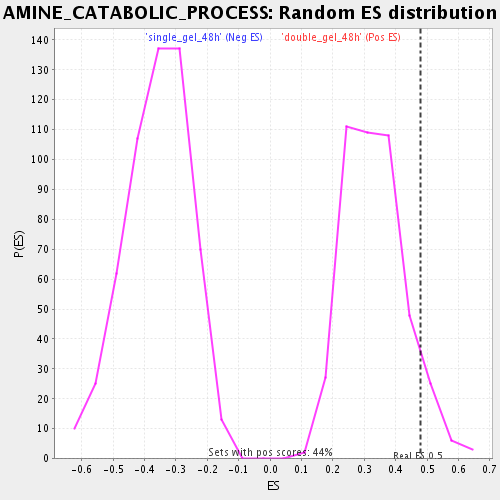
| Enrichment Score (ES) | 0.47959656 |
| Normalized Enrichment Score (NES) | 1.4395926 |
| Nominal p-value | 0.075170845 |
| FDR q-value | 0.18351328 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ARG1 | ARG1 Entrez, Source | arginase, liver | 117 | 0.209 | 0.1121 | Yes |
| 2 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 197 | 0.173 | 0.2065 | Yes |
| 3 | ASL | ASL Entrez, Source | argininosuccinate lyase | 273 | 0.154 | 0.2901 | Yes |
| 4 | HGD | HGD Entrez, Source | homogentisate 1,2-dioxygenase (homogentisate oxidase) | 457 | 0.129 | 0.3512 | Yes |
| 5 | FAH | FAH Entrez, Source | fumarylacetoacetate hydrolase (fumarylacetoacetase) | 489 | 0.126 | 0.4218 | Yes |
| 6 | BCKDK | BCKDK Entrez, Source | branched chain ketoacid dehydrogenase kinase | 866 | 0.104 | 0.4538 | Yes |
| 7 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 1212 | 0.089 | 0.4796 | Yes |
| 8 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 2252 | 0.063 | 0.4381 | No |
| 9 | ASRGL1 | ASRGL1 Entrez, Source | asparaginase like 1 | 2861 | 0.052 | 0.4227 | No |
| 10 | GCSH | GCSH Entrez, Source | glycine cleavage system protein H (aminomethyl carrier) | 3126 | 0.048 | 0.4309 | No |
| 11 | AMT | AMT Entrez, Source | aminomethyltransferase | 3552 | 0.042 | 0.4234 | No |
| 12 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 3912 | 0.038 | 0.4183 | No |
| 13 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 4645 | 0.028 | 0.3796 | No |
| 14 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 4751 | 0.027 | 0.3873 | No |
| 15 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 5455 | 0.019 | 0.3454 | No |
| 16 | COLQ | COLQ Entrez, Source | collagen-like tail subunit (single strand of homotrimer) of asymmetric acetylcholinesterase | 6022 | 0.013 | 0.3101 | No |
| 17 | GOT2 | GOT2 Entrez, Source | glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) | 6812 | 0.004 | 0.2532 | No |
| 18 | HPD | HPD Entrez, Source | 4-hydroxyphenylpyruvate dioxygenase | 8248 | -0.012 | 0.1527 | No |
| 19 | DHPS | DHPS Entrez, Source | deoxyhypusine synthase | 8363 | -0.014 | 0.1521 | No |
| 20 | GLUD1 | GLUD1 Entrez, Source | glutamate dehydrogenase 1 | 8367 | -0.014 | 0.1598 | No |
| 21 | MCCC2 | MCCC2 Entrez, Source | methylcrotonoyl-Coenzyme A carboxylase 2 (beta) | 9753 | -0.033 | 0.0748 | No |
| 22 | DDAH1 | DDAH1 Entrez, Source | dimethylarginine dimethylaminohydrolase 1 | 12847 | -0.155 | -0.0675 | No |
| 23 | DDAH2 | DDAH2 Entrez, Source | dimethylarginine dimethylaminohydrolase 2 | 12970 | -0.181 | 0.0279 | No |