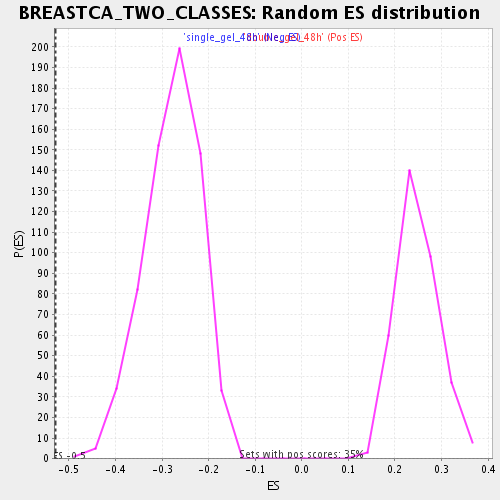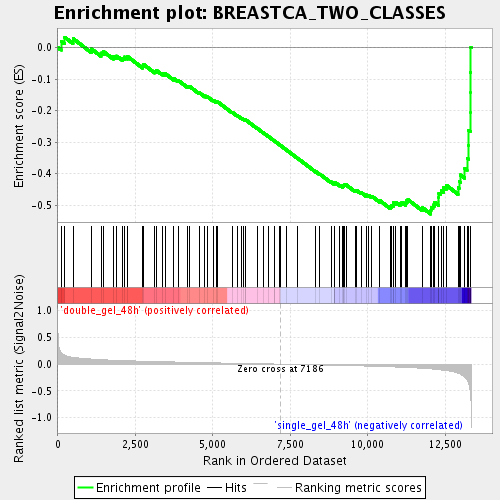
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | single_gel_48h |
| GeneSet | BREASTCA_TWO_CLASSES |
| Enrichment Score (ES) | -0.5285348 |
| Normalized Enrichment Score (NES) | -1.898565 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.024547588 |
| FWER p-Value | 0.648 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LPIN1 | LPIN1 Entrez, Source | lipin 1 | 104 | 0.220 | 0.0200 | No |
| 2 | FKBP5 | FKBP5 Entrez, Source | FK506 binding protein 5 | 219 | 0.169 | 0.0328 | No |
| 3 | FAH | FAH Entrez, Source | fumarylacetoacetate hydrolase (fumarylacetoacetase) | 489 | 0.126 | 0.0285 | No |
| 4 | GNAS | GNAS Entrez, Source | GNAS complex locus | 1079 | 0.094 | -0.0040 | No |
| 5 | MTHFD1 | MTHFD1 Entrez, Source | methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1, methenyltetrahydrofolate cyclohydrolase, formyltetrahydrofolate synthetase | 1392 | 0.083 | -0.0171 | No |
| 6 | FGFR4 | FGFR4 Entrez, Source | fibroblast growth factor receptor 4 | 1473 | 0.080 | -0.0129 | No |
| 7 | OPRS1 | OPRS1 Entrez, Source | opioid receptor, sigma 1 | 1787 | 0.072 | -0.0274 | No |
| 8 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 1896 | 0.070 | -0.0267 | No |
| 9 | GART | GART Entrez, Source | phosphoribosylglycinamide formyltransferase, phosphoribosylglycinamide synthetase, phosphoribosylaminoimidazole synthetase | 2097 | 0.066 | -0.0334 | No |
| 10 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 2158 | 0.065 | -0.0297 | No |
| 11 | SLC35B1 | SLC35B1 Entrez, Source | solute carrier family 35, member B1 | 2250 | 0.063 | -0.0286 | No |
| 12 | COG3 | COG3 Entrez, Source | component of oligomeric golgi complex 3 | 2736 | 0.055 | -0.0583 | No |
| 13 | DDT | DDT Entrez, Source | D-dopachrome tautomerase | 2762 | 0.054 | -0.0533 | No |
| 14 | GCSH | GCSH Entrez, Source | glycine cleavage system protein H (aminomethyl carrier) | 3126 | 0.048 | -0.0746 | No |
| 15 | EIF2C2 | EIF2C2 Entrez, Source | eukaryotic translation initiation factor 2C, 2 | 3179 | 0.048 | -0.0725 | No |
| 16 | DLL1 | DLL1 Entrez, Source | delta-like 1 (Drosophila) | 3375 | 0.045 | -0.0815 | No |
| 17 | WWP1 | WWP1 Entrez, Source | WW domain containing E3 ubiquitin protein ligase 1 | 3459 | 0.044 | -0.0823 | No |
| 18 | ATN1 | ATN1 Entrez, Source | atrophin 1 | 3743 | 0.040 | -0.0986 | No |
| 19 | CSDA | CSDA Entrez, Source | cold shock domain protein A | 3880 | 0.038 | -0.1040 | No |
| 20 | RAB9 | RAB9 Entrez, Source | RAB9, member RAS oncogene family | 4181 | 0.034 | -0.1223 | No |
| 21 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 4258 | 0.033 | -0.1239 | No |
| 22 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 4559 | 0.029 | -0.1428 | No |
| 23 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 4746 | 0.027 | -0.1534 | No |
| 24 | PHYH | PHYH Entrez, Source | phytanoyl-CoA 2-hydroxylase | 4827 | 0.026 | -0.1562 | No |
| 25 | MLLT3 | MLLT3 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 3 | 5005 | 0.024 | -0.1665 | No |
| 26 | PPP1CB | PPP1CB Entrez, Source | protein phosphatase 1, catalytic subunit, beta isoform | 5113 | 0.022 | -0.1718 | No |
| 27 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 5156 | 0.022 | -0.1722 | No |
| 28 | GMPS | GMPS Entrez, Source | guanine monphosphate synthetase | 5619 | 0.017 | -0.2049 | No |
| 29 | C10ORF7 | C10ORF7 Entrez, Source | chromosome 10 open reading frame 7 | 5783 | 0.015 | -0.2153 | No |
| 30 | GPX4 | GPX4 Entrez, Source | glutathione peroxidase 4 (phospholipid hydroperoxidase) | 5917 | 0.014 | -0.2236 | No |
| 31 | PARD3 | PARD3 Entrez, Source | par-3 partitioning defective 3 homolog (C. elegans) | 5990 | 0.013 | -0.2274 | No |
| 32 | THOC1 | THOC1 Entrez, Source | THO complex 1 | 6041 | 0.012 | -0.2296 | No |
| 33 | TUBGCP3 | TUBGCP3 Entrez, Source | tubulin, gamma complex associated protein 3 | 6045 | 0.012 | -0.2283 | No |
| 34 | FGFR3 | FGFR3 Entrez, Source | fibroblast growth factor receptor 3 (achondroplasia, thanatophoric dwarfism) | 6436 | 0.008 | -0.2567 | No |
| 35 | MAPK1 | MAPK1 Entrez, Source | mitogen-activated protein kinase 1 | 6633 | 0.006 | -0.2707 | No |
| 36 | CREB1 | CREB1 Entrez, Source | cAMP responsive element binding protein 1 | 6781 | 0.004 | -0.2812 | No |
| 37 | NUP93 | NUP93 Entrez, Source | nucleoporin 93kDa | 6978 | 0.002 | -0.2957 | No |
| 38 | COX6C | COX6C Entrez, Source | cytochrome c oxidase subunit VIc | 6979 | 0.002 | -0.2955 | No |
| 39 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 7160 | 0.000 | -0.3090 | No |
| 40 | GRB2 | GRB2 Entrez, Source | growth factor receptor-bound protein 2 | 7179 | 0.000 | -0.3104 | No |
| 41 | SERBP1 | SERBP1 Entrez, Source | SERPINE1 mRNA binding protein 1 | 7386 | -0.002 | -0.3256 | No |
| 42 | GABRP | GABRP Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, pi | 7726 | -0.006 | -0.3504 | No |
| 43 | CEP57 | CEP57 Entrez, Source | centrosomal protein 57kDa | 8327 | -0.013 | -0.3940 | No |
| 44 | ST13 | ST13 Entrez, Source | suppression of tumorigenicity 13 (colon carcinoma) (Hsp70 interacting protein) | 8448 | -0.015 | -0.4012 | No |
| 45 | OAZ1 | OAZ1 Entrez, Source | ornithine decarboxylase antizyme 1 | 8815 | -0.019 | -0.4264 | No |
| 46 | CTNNB1 | CTNNB1 Entrez, Source | catenin (cadherin-associated protein), beta 1, 88kDa | 8921 | -0.021 | -0.4316 | No |
| 47 | SF3A3 | SF3A3 Entrez, Source | splicing factor 3a, subunit 3, 60kDa | 8924 | -0.021 | -0.4292 | No |
| 48 | GDI2 | GDI2 Entrez, Source | GDP dissociation inhibitor 2 | 8938 | -0.021 | -0.4275 | No |
| 49 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 9099 | -0.023 | -0.4366 | No |
| 50 | ATP6AP1 | ATP6AP1 Entrez, Source | ATPase, H+ transporting, lysosomal accessory protein 1 | 9190 | -0.024 | -0.4403 | No |
| 51 | RDX | RDX Entrez, Source | radixin | 9216 | -0.025 | -0.4391 | No |
| 52 | PSIP1 | PSIP1 Entrez, Source | PC4 and SFRS1 interacting protein 1 | 9227 | -0.025 | -0.4367 | No |
| 53 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 9257 | -0.025 | -0.4357 | No |
| 54 | ENPP1 | ENPP1 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 1 | 9297 | -0.026 | -0.4354 | No |
| 55 | NUP155 | NUP155 Entrez, Source | nucleoporin 155kDa | 9601 | -0.030 | -0.4544 | No |
| 56 | UBE2B | UBE2B Entrez, Source | ubiquitin-conjugating enzyme E2B (RAD6 homolog) | 9639 | -0.031 | -0.4532 | No |
| 57 | THRAP4 | THRAP4 Entrez, Source | thyroid hormone receptor associated protein 4 | 9782 | -0.033 | -0.4597 | No |
| 58 | HDAC3 | HDAC3 Entrez, Source | histone deacetylase 3 | 9948 | -0.036 | -0.4677 | No |
| 59 | ZFP36L1 | ZFP36L1 Entrez, Source | zinc finger protein 36, C3H type-like 1 | 10023 | -0.037 | -0.4686 | No |
| 60 | HMGN3 | HMGN3 Entrez, Source | high mobility group nucleosomal binding domain 3 | 10126 | -0.039 | -0.4713 | No |
| 61 | OGT | OGT Entrez, Source | O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) | 10387 | -0.043 | -0.4855 | No |
| 62 | KIF5B | KIF5B Entrez, Source | kinesin family member 5B | 10742 | -0.050 | -0.5058 | No |
| 63 | PARP1 | PARP1 Entrez, Source | poly (ADP-ribose) polymerase family, member 1 | 10758 | -0.051 | -0.5005 | No |
| 64 | HP1BP3 | HP1BP3 Entrez, Source | heterochromatin protein 1, binding protein 3 | 10822 | -0.052 | -0.4987 | No |
| 65 | CD63 | CD63 Entrez, Source | CD63 molecule | 10826 | -0.052 | -0.4923 | No |
| 66 | ERBB2 | ERBB2 Entrez, Source | v-erb-b2 erythroblastic leukemia viral oncogene homolog 2, neuro/glioblastoma derived oncogene homolog (avian) | 10906 | -0.054 | -0.4915 | No |
| 67 | MNAT1 | MNAT1 Entrez, Source | menage a trois homolog 1, cyclin H assembly factor (Xenopus laevis) | 11046 | -0.057 | -0.4947 | No |
| 68 | BTG3 | BTG3 Entrez, Source | BTG family, member 3 | 11098 | -0.058 | -0.4913 | No |
| 69 | SMAD2 | SMAD2 Entrez, Source | SMAD, mothers against DPP homolog 2 (Drosophila) | 11223 | -0.061 | -0.4929 | No |
| 70 | POLR2F | POLR2F Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide F | 11240 | -0.061 | -0.4864 | No |
| 71 | FABP5 | FABP5 Entrez, Source | fatty acid binding protein 5 (psoriasis-associated) | 11296 | -0.062 | -0.4827 | No |
| 72 | CRAT | CRAT Entrez, Source | carnitine acetyltransferase | 11766 | -0.078 | -0.5082 | No |
| 73 | CALM2 | CALM2 Entrez, Source | calmodulin 2 (phosphorylase kinase, delta) | 12036 | -0.089 | -0.5173 | Yes |
| 74 | LRPAP1 | LRPAP1 Entrez, Source | low density lipoprotein receptor-related protein associated protein 1 | 12044 | -0.089 | -0.5065 | Yes |
| 75 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 12106 | -0.092 | -0.4995 | Yes |
| 76 | DEAF1 | DEAF1 Entrez, Source | deformed epidermal autoregulatory factor 1 (Drosophila) | 12157 | -0.094 | -0.4913 | Yes |
| 77 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 12287 | -0.103 | -0.4880 | Yes |
| 78 | CBX3 | CBX3 Entrez, Source | chromobox homolog 3 (HP1 gamma homolog, Drosophila) | 12288 | -0.103 | -0.4750 | Yes |
| 79 | CDK4 | CDK4 Entrez, Source | cyclin-dependent kinase 4 | 12294 | -0.103 | -0.4624 | Yes |
| 80 | TUBB | TUBB Entrez, Source | tubulin, beta | 12364 | -0.108 | -0.4539 | Yes |
| 81 | GNA12 | GNA12 Entrez, Source | guanine nucleotide binding protein (G protein) alpha 12 | 12436 | -0.113 | -0.4449 | Yes |
| 82 | IRF1 | IRF1 Entrez, Source | interferon regulatory factor 1 | 12531 | -0.120 | -0.4368 | Yes |
| 83 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 12915 | -0.168 | -0.4443 | Yes |
| 84 | PXN | PXN Entrez, Source | paxillin | 12950 | -0.176 | -0.4247 | Yes |
| 85 | ALCAM | ALCAM Entrez, Source | activated leukocyte cell adhesion molecule | 12991 | -0.186 | -0.4041 | Yes |
| 86 | VLDLR | VLDLR Entrez, Source | very low density lipoprotein receptor | 13129 | -0.246 | -0.3833 | Yes |
| 87 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 13206 | -0.298 | -0.3513 | Yes |
| 88 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 13250 | -0.352 | -0.3100 | Yes |
| 89 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13257 | -0.367 | -0.2640 | Yes |
| 90 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 13308 | -0.484 | -0.2064 | Yes |
| 91 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13313 | -0.503 | -0.1430 | Yes |
| 92 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 13318 | -0.516 | -0.0779 | Yes |
| 93 | MSN | MSN Entrez, Source | moesin | 13327 | -0.629 | 0.0011 | Yes |