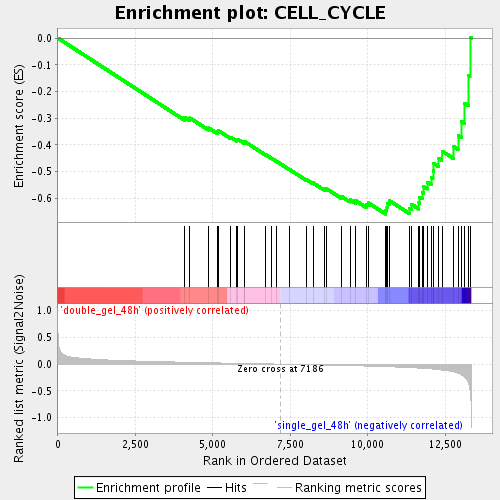
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | single_gel_48h |
| GeneSet | CELL_CYCLE |
| Enrichment Score (ES) | -0.6585951 |
| Normalized Enrichment Score (NES) | -2.0737388 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0042932653 |
| FWER p-Value | 0.069 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCNH | CCNH Entrez, Source | cyclin H | 4088 | 0.035 | -0.2965 | No |
| 2 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 4258 | 0.033 | -0.2989 | No |
| 3 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 4860 | 0.026 | -0.3362 | No |
| 4 | MAD2L2 | MAD2L2 Entrez, Source | MAD2 mitotic arrest deficient-like 2 (yeast) | 5148 | 0.022 | -0.3509 | No |
| 5 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 5178 | 0.022 | -0.3464 | No |
| 6 | CCNE1 | CCNE1 Entrez, Source | cyclin E1 | 5568 | 0.017 | -0.3702 | No |
| 7 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 5771 | 0.015 | -0.3806 | No |
| 8 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 5802 | 0.015 | -0.3783 | No |
| 9 | ORC2L | ORC2L Entrez, Source | origin recognition complex, subunit 2-like (yeast) | 6017 | 0.013 | -0.3904 | No |
| 10 | CCNA1 | CCNA1 Entrez, Source | cyclin A1 | 6019 | 0.013 | -0.3866 | No |
| 11 | BUB3 | BUB3 Entrez, Source | BUB3 budding uninhibited by benzimidazoles 3 homolog (yeast) | 6706 | 0.005 | -0.4366 | No |
| 12 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 6894 | 0.003 | -0.4497 | No |
| 13 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 7053 | 0.001 | -0.4611 | No |
| 14 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 7469 | -0.003 | -0.4913 | No |
| 15 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 8020 | -0.010 | -0.5296 | No |
| 16 | ORC4L | ORC4L Entrez, Source | origin recognition complex, subunit 4-like (yeast) | 8232 | -0.012 | -0.5417 | No |
| 17 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 8604 | -0.017 | -0.5643 | No |
| 18 | E2F5 | E2F5 Entrez, Source | E2F transcription factor 5, p130-binding | 8663 | -0.018 | -0.5632 | No |
| 19 | YWHAG | YWHAG Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide | 9159 | -0.024 | -0.5930 | No |
| 20 | ORC3L | ORC3L Entrez, Source | origin recognition complex, subunit 3-like (yeast) | 9431 | -0.028 | -0.6047 | No |
| 21 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 9613 | -0.031 | -0.6088 | No |
| 22 | HDAC3 | HDAC3 Entrez, Source | histone deacetylase 3 | 9948 | -0.036 | -0.6229 | No |
| 23 | CHEK2 | CHEK2 Entrez, Source | CHK2 checkpoint homolog (S. pombe) | 10020 | -0.037 | -0.6167 | No |
| 24 | HDAC6 | HDAC6 Entrez, Source | histone deacetylase 6 | 10578 | -0.047 | -0.6441 | Yes |
| 25 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 10619 | -0.047 | -0.6325 | Yes |
| 26 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 10625 | -0.047 | -0.6181 | Yes |
| 27 | ORC6L | ORC6L Entrez, Source | origin recognition complex, subunit 6 like (yeast) | 10702 | -0.049 | -0.6085 | Yes |
| 28 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 11337 | -0.063 | -0.6365 | Yes |
| 29 | RB1 | RB1 Entrez, Source | retinoblastoma 1 (including osteosarcoma) | 11401 | -0.065 | -0.6210 | Yes |
| 30 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 11651 | -0.074 | -0.6168 | Yes |
| 31 | GSK3B | GSK3B Entrez, Source | glycogen synthase kinase 3 beta | 11670 | -0.074 | -0.5950 | Yes |
| 32 | CCND3 | CCND3 Entrez, Source | cyclin D3 | 11760 | -0.077 | -0.5777 | Yes |
| 33 | ORC1L | ORC1L Entrez, Source | origin recognition complex, subunit 1-like (yeast) | 11803 | -0.079 | -0.5563 | Yes |
| 34 | HDAC7A | HDAC7A Entrez, Source | histone deacetylase 7A | 11917 | -0.084 | -0.5387 | Yes |
| 35 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 12064 | -0.090 | -0.5218 | Yes |
| 36 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 12106 | -0.092 | -0.4964 | Yes |
| 37 | HDAC5 | HDAC5 Entrez, Source | histone deacetylase 5 | 12129 | -0.093 | -0.4692 | Yes |
| 38 | CDK4 | CDK4 Entrez, Source | cyclin-dependent kinase 4 | 12294 | -0.103 | -0.4495 | Yes |
| 39 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 12396 | -0.111 | -0.4227 | Yes |
| 40 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 12774 | -0.145 | -0.4062 | Yes |
| 41 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 12915 | -0.168 | -0.3645 | Yes |
| 42 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 13022 | -0.195 | -0.3118 | Yes |
| 43 | CCNB1 | CCNB1 Entrez, Source | cyclin B1 | 13126 | -0.245 | -0.2435 | Yes |
| 44 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13257 | -0.367 | -0.1394 | Yes |
| 45 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13301 | -0.469 | 0.0031 | Yes |

