Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | CHAUVIN_ANDROGEN_REGULATED_GENES |
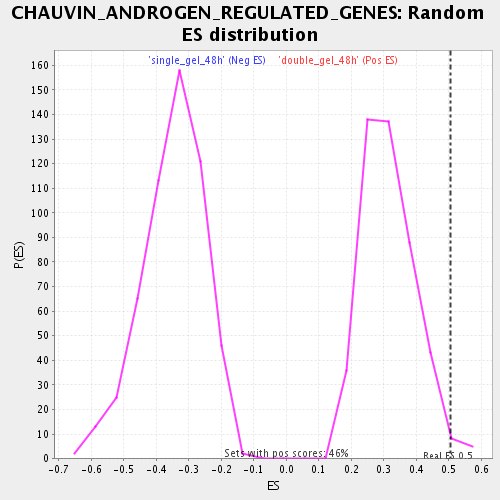
| Enrichment Score (ES) | 0.5069936 |
| Normalized Enrichment Score (NES) | 1.603581 |
| Nominal p-value | 0.017582418 |
| FDR q-value | 0.09288066 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PEMT | PEMT Entrez, Source | phosphatidylethanolamine N-methyltransferase | 4 | 0.491 | 0.1382 | Yes |
| 2 | FDFT1 | FDFT1 Entrez, Source | farnesyl-diphosphate farnesyltransferase 1 | 44 | 0.291 | 0.2173 | Yes |
| 3 | EBP | EBP Entrez, Source | emopamil binding protein (sterol isomerase) | 45 | 0.290 | 0.2990 | Yes |
| 4 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 90 | 0.236 | 0.3623 | Yes |
| 5 | FDPS | FDPS Entrez, Source | farnesyl diphosphate synthase (farnesyl pyrophosphate synthetase, dimethylallyltranstransferase, geranyltranstransferase) | 171 | 0.184 | 0.4080 | Yes |
| 6 | FKBP5 | FKBP5 Entrez, Source | FK506 binding protein 5 | 219 | 0.169 | 0.4521 | Yes |
| 7 | ACE | ACE Entrez, Source | angiotensin I converting enzyme (peptidyl-dipeptidase A) 1 | 281 | 0.152 | 0.4903 | Yes |
| 8 | RAMP3 | RAMP3 Entrez, Source | receptor (calcitonin) activity modifying protein 3 | 1115 | 0.093 | 0.4539 | Yes |
| 9 | ADAM7 | ADAM7 Entrez, Source | ADAM metallopeptidase domain 7 | 1156 | 0.091 | 0.4766 | Yes |
| 10 | GFRA1 | GFRA1 Entrez, Source | GDNF family receptor alpha 1 | 1292 | 0.086 | 0.4907 | Yes |
| 11 | CA4 | CA4 Entrez, Source | carbonic anhydrase IV | 1388 | 0.083 | 0.5070 | Yes |
| 12 | PTPRO | PTPRO Entrez, Source | protein tyrosine phosphatase, receptor type, O | 2452 | 0.059 | 0.4439 | No |
| 13 | FKBP11 | FKBP11 Entrez, Source | FK506 binding protein 11, 19 kDa | 2860 | 0.052 | 0.4281 | No |
| 14 | ALPL | ALPL Entrez, Source | alkaline phosphatase, liver/bone/kidney | 4222 | 0.034 | 0.3353 | No |
| 15 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 4751 | 0.027 | 0.3032 | No |
| 16 | CA2 | CA2 Entrez, Source | carbonic anhydrase II | 7034 | 0.002 | 0.1323 | No |
| 17 | KCNK1 | KCNK1 Entrez, Source | potassium channel, subfamily K, member 1 | 7451 | -0.003 | 0.1019 | No |
| 18 | ADRA2C | ADRA2C Entrez, Source | adrenergic, alpha-2C-, receptor | 7484 | -0.004 | 0.1005 | No |
| 19 | NUCB2 | NUCB2 Entrez, Source | nucleobindin 2 | 7742 | -0.006 | 0.0830 | No |
| 20 | GSTM1 | GSTM1 Entrez, Source | glutathione S-transferase M1 | 8992 | -0.022 | -0.0047 | No |
| 21 | ENPP1 | ENPP1 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 1 | 9297 | -0.026 | -0.0202 | No |
| 22 | GLUL | GLUL Entrez, Source | glutamate-ammonia ligase (glutamine synthetase) | 9734 | -0.033 | -0.0438 | No |
| 23 | GPC1 | GPC1 Entrez, Source | glypican 1 | 9818 | -0.034 | -0.0406 | No |
| 24 | ADCY8 | ADCY8 Entrez, Source | adenylate cyclase 8 (brain) | 10557 | -0.046 | -0.0830 | No |
| 25 | BTG3 | BTG3 Entrez, Source | BTG family, member 3 | 11098 | -0.058 | -0.1072 | No |
| 26 | TPMT | TPMT Entrez, Source | thiopurine S-methyltransferase | 12268 | -0.101 | -0.1666 | No |
| 27 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 12691 | -0.134 | -0.1605 | No |
| 28 | GJB3 | GJB3 Entrez, Source | gap junction protein, beta 3, 31kDa (connexin 31) | 13204 | -0.296 | -0.1157 | No |
| 29 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 13294 | -0.447 | 0.0036 | No |