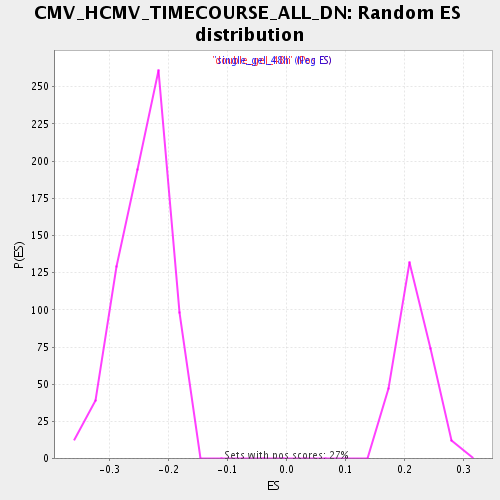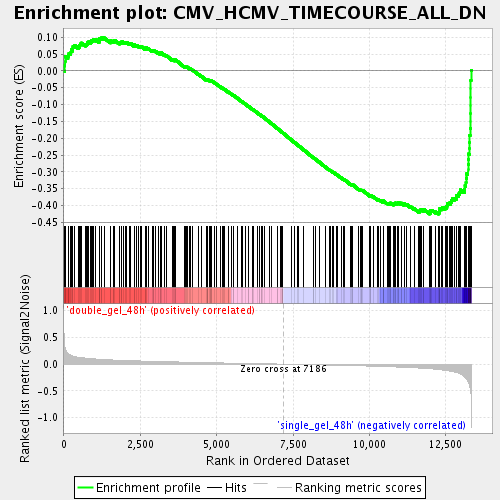
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | single_gel_48h |
| GeneSet | CMV_HCMV_TIMECOURSE_ALL_DN |
| Enrichment Score (ES) | -0.42529452 |
| Normalized Enrichment Score (NES) | -1.7562443 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.048987787 |
| FWER p-Value | 0.997 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MVD | MVD Entrez, Source | mevalonate (diphospho) decarboxylase | 21 | 0.352 | 0.0154 | No |
| 2 | GNG11 | GNG11 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 11 | 35 | 0.306 | 0.0291 | No |
| 3 | PPP1R3C | PPP1R3C Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 3C | 40 | 0.295 | 0.0431 | No |
| 4 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 134 | 0.198 | 0.0455 | No |
| 5 | ACAA2 | ACAA2 Entrez, Source | acetyl-Coenzyme A acyltransferase 2 (mitochondrial 3-oxoacyl-Coenzyme A thiolase) | 163 | 0.187 | 0.0524 | No |
| 6 | ACOX1 | ACOX1 Entrez, Source | acyl-Coenzyme A oxidase 1, palmitoyl | 227 | 0.165 | 0.0556 | No |
| 7 | IRS1 | IRS1 Entrez, Source | insulin receptor substrate 1 | 233 | 0.163 | 0.0631 | No |
| 8 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 292 | 0.150 | 0.0659 | No |
| 9 | CXCL12 | CXCL12 Entrez, Source | chemokine (C-X-C motif) ligand 12 (stromal cell-derived factor 1) | 295 | 0.150 | 0.0730 | No |
| 10 | MMP3 | MMP3 Entrez, Source | matrix metallopeptidase 3 (stromelysin 1, progelatinase) | 351 | 0.142 | 0.0756 | No |
| 11 | FAH | FAH Entrez, Source | fumarylacetoacetate hydrolase (fumarylacetoacetase) | 489 | 0.126 | 0.0712 | No |
| 12 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 506 | 0.124 | 0.0760 | No |
| 13 | IGF2BP3 | IGF2BP3 Entrez, Source | insulin-like growth factor 2 mRNA binding protein 3 | 531 | 0.123 | 0.0801 | No |
| 14 | SNUPN | SNUPN Entrez, Source | snurportin 1 | 571 | 0.120 | 0.0829 | No |
| 15 | WNT5A | WNT5A Entrez, Source | wingless-type MMTV integration site family, member 5A | 696 | 0.112 | 0.0789 | No |
| 16 | MALL | MALL Entrez, Source | mal, T-cell differentiation protein-like | 754 | 0.109 | 0.0798 | No |
| 17 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 766 | 0.108 | 0.0842 | No |
| 18 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 809 | 0.106 | 0.0861 | No |
| 19 | AKT3 | AKT3 Entrez, Source | v-akt murine thymoma viral oncogene homolog 3 (protein kinase B, gamma) | 871 | 0.104 | 0.0865 | No |
| 20 | NTF3 | NTF3 Entrez, Source | neurotrophin 3 | 895 | 0.103 | 0.0897 | No |
| 21 | PCYT2 | PCYT2 Entrez, Source | phosphate cytidylyltransferase 2, ethanolamine | 944 | 0.100 | 0.0908 | No |
| 22 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 960 | 0.099 | 0.0944 | No |
| 23 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 1041 | 0.096 | 0.0929 | No |
| 24 | FGF7 | FGF7 Entrez, Source | fibroblast growth factor 7 (keratinocyte growth factor) | 1152 | 0.091 | 0.0890 | No |
| 25 | GSTT2 | GSTT2 Entrez, Source | glutathione S-transferase theta 2 | 1160 | 0.091 | 0.0928 | No |
| 26 | ISG20L2 | ISG20L2 Entrez, Source | interferon stimulated exonuclease gene 20kDa-like 2 | 1162 | 0.091 | 0.0971 | No |
| 27 | DUSP4 | DUSP4 Entrez, Source | dual specificity phosphatase 4 | 1225 | 0.089 | 0.0967 | No |
| 28 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 1240 | 0.088 | 0.0999 | No |
| 29 | GRIA3 | GRIA3 Entrez, Source | glutamate receptor, ionotrophic, AMPA 3 | 1320 | 0.085 | 0.0980 | No |
| 30 | ETFB | ETFB Entrez, Source | electron-transfer-flavoprotein, beta polypeptide | 1520 | 0.079 | 0.0866 | No |
| 31 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 1533 | 0.079 | 0.0895 | No |
| 32 | TWIST1 | TWIST1 Entrez, Source | twist homolog 1 (acrocephalosyndactyly 3; Saethre-Chotzen syndrome) (Drosophila) | 1621 | 0.077 | 0.0865 | No |
| 33 | CDH6 | CDH6 Entrez, Source | cadherin 6, type 2, K-cadherin (fetal kidney) | 1638 | 0.076 | 0.0890 | No |
| 34 | LRRC17 | LRRC17 Entrez, Source | leucine rich repeat containing 17 | 1657 | 0.075 | 0.0912 | No |
| 35 | GRM5 | GRM5 Entrez, Source | glutamate receptor, metabotropic 5 | 1811 | 0.072 | 0.0830 | No |
| 36 | SIX1 | SIX1 Entrez, Source | sine oculis homeobox homolog 1 (Drosophila) | 1834 | 0.071 | 0.0848 | No |
| 37 | ANPEP | ANPEP Entrez, Source | alanyl (membrane) aminopeptidase (aminopeptidase N, aminopeptidase M, microsomal aminopeptidase, CD13, p150) | 1876 | 0.070 | 0.0851 | No |
| 38 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 1896 | 0.070 | 0.0870 | No |
| 39 | NR2F2 | NR2F2 Entrez, Source | nuclear receptor subfamily 2, group F, member 2 | 1957 | 0.069 | 0.0857 | No |
| 40 | STC2 | STC2 Entrez, Source | stanniocalcin 2 | 2017 | 0.068 | 0.0845 | No |
| 41 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 2046 | 0.067 | 0.0856 | No |
| 42 | SDC2 | SDC2 Entrez, Source | syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated, fibroglycan) | 2139 | 0.065 | 0.0817 | No |
| 43 | SLC24A1 | SLC24A1 Entrez, Source | solute carrier family 24 (sodium/potassium/calcium exchanger), member 1 | 2191 | 0.064 | 0.0809 | No |
| 44 | CSTF1 | CSTF1 Entrez, Source | cleavage stimulation factor, 3' pre-RNA, subunit 1, 50kDa | 2297 | 0.062 | 0.0759 | No |
| 45 | TSPYL4 | TSPYL4 Entrez, Source | TSPY-like 4 | 2308 | 0.062 | 0.0781 | No |
| 46 | LPXN | LPXN Entrez, Source | leupaxin | 2384 | 0.061 | 0.0753 | No |
| 47 | LOXL1 | LOXL1 Entrez, Source | lysyl oxidase-like 1 | 2447 | 0.060 | 0.0735 | No |
| 48 | PTGIS | PTGIS Entrez, Source | prostaglandin I2 (prostacyclin) synthase | 2515 | 0.058 | 0.0712 | No |
| 49 | CXCR7 | CXCR7 Entrez, Source | chemokine (C-X-C motif) receptor 7 | 2536 | 0.058 | 0.0724 | No |
| 50 | FGF5 | FGF5 Entrez, Source | fibroblast growth factor 5 | 2673 | 0.056 | 0.0648 | No |
| 51 | SYNE1 | SYNE1 Entrez, Source | spectrin repeat containing, nuclear envelope 1 | 2675 | 0.056 | 0.0674 | No |
| 52 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 2688 | 0.055 | 0.0691 | No |
| 53 | DDT | DDT Entrez, Source | D-dopachrome tautomerase | 2762 | 0.054 | 0.0662 | No |
| 54 | SIP1 | SIP1 Entrez, Source | survival of motor neuron protein interacting protein 1 | 2894 | 0.052 | 0.0587 | No |
| 55 | PHLDA1 | PHLDA1 Entrez, Source | pleckstrin homology-like domain, family A, member 1 | 2920 | 0.051 | 0.0592 | No |
| 56 | MAGI2 | MAGI2 Entrez, Source | membrane associated guanylate kinase, WW and PDZ domain containing 2 | 2940 | 0.051 | 0.0603 | No |
| 57 | GBE1 | GBE1 Entrez, Source | glucan (1,4-alpha-), branching enzyme 1 (glycogen branching enzyme, Andersen disease, glycogen storage disease type IV) | 2997 | 0.050 | 0.0584 | No |
| 58 | TMEM158 | TMEM158 Entrez, Source | transmembrane protein 158 | 3086 | 0.049 | 0.0541 | No |
| 59 | GNRH1 | GNRH1 Entrez, Source | gonadotropin-releasing hormone 1 (luteinizing-releasing hormone) | 3167 | 0.048 | 0.0502 | No |
| 60 | GCS1 | GCS1 Entrez, Source | - | 3175 | 0.048 | 0.0520 | No |
| 61 | PLEC1 | PLEC1 Entrez, Source | plectin 1, intermediate filament binding protein 500kDa | 3180 | 0.048 | 0.0540 | No |
| 62 | GJA4 | GJA4 Entrez, Source | gap junction protein, alpha 4, 37kDa (connexin 37) | 3292 | 0.046 | 0.0477 | No |
| 63 | THPO | THPO Entrez, Source | thrombopoietin (myeloproliferative leukemia virus oncogene ligand, megakaryocyte growth and development factor) | 3345 | 0.045 | 0.0460 | No |
| 64 | SMAD6 | SMAD6 Entrez, Source | SMAD, mothers against DPP homolog 6 (Drosophila) | 3551 | 0.042 | 0.0324 | No |
| 65 | PDGFRA | PDGFRA Entrez, Source | platelet-derived growth factor receptor, alpha polypeptide | 3570 | 0.042 | 0.0330 | No |
| 66 | DCAMKL1 | DCAMKL1 Entrez, Source | doublecortin and CaM kinase-like 1 | 3621 | 0.041 | 0.0312 | No |
| 67 | POLD2 | POLD2 Entrez, Source | polymerase (DNA directed), delta 2, regulatory subunit 50kDa | 3642 | 0.041 | 0.0317 | No |
| 68 | NOTCH2 | NOTCH2 Entrez, Source | Notch homolog 2 (Drosophila) | 3652 | 0.041 | 0.0329 | No |
| 69 | TCF21 | TCF21 Entrez, Source | transcription factor 21 | 3948 | 0.037 | 0.0122 | No |
| 70 | SEMA3A | SEMA3A Entrez, Source | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A | 3987 | 0.037 | 0.0111 | No |
| 71 | UMPS | UMPS Entrez, Source | uridine monophosphate synthetase (orotate phosphoribosyl transferase and orotidine-5'-decarboxylase) | 4001 | 0.036 | 0.0119 | No |
| 72 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 4028 | 0.036 | 0.0116 | No |
| 73 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 4121 | 0.035 | 0.0063 | No |
| 74 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 4134 | 0.035 | 0.0070 | No |
| 75 | CEBPD | CEBPD Entrez, Source | CCAAT/enhancer binding protein (C/EBP), delta | 4223 | 0.034 | 0.0020 | No |
| 76 | ANKRD15 | ANKRD15 Entrez, Source | ankyrin repeat domain 15 | 4401 | 0.031 | -0.0100 | No |
| 77 | LRRC15 | LRRC15 Entrez, Source | leucine rich repeat containing 15 | 4496 | 0.030 | -0.0158 | No |
| 78 | ACO1 | ACO1 Entrez, Source | aconitase 1, soluble | 4655 | 0.028 | -0.0265 | No |
| 79 | SLIT2 | SLIT2 Entrez, Source | slit homolog 2 (Drosophila) | 4685 | 0.028 | -0.0273 | No |
| 80 | PEX11B | PEX11B Entrez, Source | peroxisomal biogenesis factor 11B | 4699 | 0.028 | -0.0270 | No |
| 81 | THBS2 | THBS2 Entrez, Source | thrombospondin 2 | 4709 | 0.027 | -0.0264 | No |
| 82 | NR1D2 | NR1D2 Entrez, Source | nuclear receptor subfamily 1, group D, member 2 | 4748 | 0.027 | -0.0280 | No |
| 83 | GJA1 | GJA1 Entrez, Source | gap junction protein, alpha 1, 43kDa (connexin 43) | 4783 | 0.027 | -0.0293 | No |
| 84 | GSTO1 | GSTO1 Entrez, Source | glutathione S-transferase omega 1 | 4785 | 0.026 | -0.0281 | No |
| 85 | RAD17 | RAD17 Entrez, Source | RAD17 homolog (S. pombe) | 4815 | 0.026 | -0.0290 | No |
| 86 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 4836 | 0.026 | -0.0293 | No |
| 87 | LAPTM5 | LAPTM5 Entrez, Source | lysosomal associated multispanning membrane protein 5 | 4925 | 0.025 | -0.0348 | No |
| 88 | PHKA1 | PHKA1 Entrez, Source | phosphorylase kinase, alpha 1 (muscle) | 5002 | 0.024 | -0.0395 | No |
| 89 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 5117 | 0.022 | -0.0471 | No |
| 90 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 5178 | 0.022 | -0.0506 | No |
| 91 | SLC12A1 | SLC12A1 Entrez, Source | solute carrier family 12 (sodium/potassium/chloride transporters), member 1 | 5212 | 0.021 | -0.0521 | No |
| 92 | MIPEP | MIPEP Entrez, Source | mitochondrial intermediate peptidase | 5270 | 0.021 | -0.0554 | No |
| 93 | CIT | CIT Entrez, Source | citron (rho-interacting, serine/threonine kinase 21) | 5375 | 0.020 | -0.0624 | No |
| 94 | MGLL | MGLL Entrez, Source | monoglyceride lipase | 5400 | 0.019 | -0.0633 | No |
| 95 | GTF2E1 | GTF2E1 Entrez, Source | general transcription factor IIE, polypeptide 1, alpha 56kDa | 5488 | 0.018 | -0.0691 | No |
| 96 | GPSM3 | GPSM3 Entrez, Source | G-protein signalling modulator 3 (AGS3-like, C. elegans) | 5544 | 0.018 | -0.0724 | No |
| 97 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 5552 | 0.018 | -0.0721 | No |
| 98 | INPP4B | INPP4B Entrez, Source | inositol polyphosphate-4-phosphatase, type II, 105kDa | 5672 | 0.016 | -0.0804 | No |
| 99 | TSPAN5 | TSPAN5 Entrez, Source | tetraspanin 5 | 5810 | 0.015 | -0.0901 | No |
| 100 | SCN8A | SCN8A Entrez, Source | sodium channel, voltage gated, type VIII, alpha | 5856 | 0.014 | -0.0929 | No |
| 101 | ADFP | ADFP Entrez, Source | adipose differentiation-related protein | 5942 | 0.013 | -0.0987 | No |
| 102 | THOC1 | THOC1 Entrez, Source | THO complex 1 | 6041 | 0.012 | -0.1056 | No |
| 103 | ITPKA | ITPKA Entrez, Source | inositol 1,4,5-trisphosphate 3-kinase A | 6042 | 0.012 | -0.1050 | No |
| 104 | CACNA1E | CACNA1E Entrez, Source | calcium channel, voltage-dependent, alpha 1E subunit | 6182 | 0.011 | -0.1150 | No |
| 105 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 6186 | 0.011 | -0.1147 | No |
| 106 | MUC4 | MUC4 Entrez, Source | mucin 4, cell surface associated | 6207 | 0.011 | -0.1157 | No |
| 107 | OPRK1 | OPRK1 Entrez, Source | opioid receptor, kappa 1 | 6331 | 0.009 | -0.1247 | No |
| 108 | COL3A1 | COL3A1 Entrez, Source | collagen, type III, alpha 1 (Ehlers-Danlos syndrome type IV, autosomal dominant) | 6389 | 0.009 | -0.1286 | No |
| 109 | RBM16 | RBM16 Entrez, Source | RNA binding motif protein 16 | 6416 | 0.008 | -0.1302 | No |
| 110 | ADD3 | ADD3 Entrez, Source | adducin 3 (gamma) | 6472 | 0.008 | -0.1340 | No |
| 111 | MAP2K6 | MAP2K6 Entrez, Source | mitogen-activated protein kinase kinase 6 | 6481 | 0.008 | -0.1342 | No |
| 112 | ICK | ICK Entrez, Source | intestinal cell (MAK-like) kinase | 6512 | 0.007 | -0.1361 | No |
| 113 | PMP22 | PMP22 Entrez, Source | peripheral myelin protein 22 | 6574 | 0.007 | -0.1405 | No |
| 114 | ARNT2 | ARNT2 Entrez, Source | aryl-hydrocarbon receptor nuclear translocator 2 | 6735 | 0.005 | -0.1524 | No |
| 115 | LYST | LYST Entrez, Source | lysosomal trafficking regulator | 6802 | 0.004 | -0.1573 | No |
| 116 | TGFBR3 | TGFBR3 Entrez, Source | transforming growth factor, beta receptor III (betaglycan, 300kDa) | 6986 | 0.002 | -0.1711 | No |
| 117 | TRIO | TRIO Entrez, Source | triple functional domain (PTPRF interacting) | 6995 | 0.002 | -0.1717 | No |
| 118 | ANGPTL2 | ANGPTL2 Entrez, Source | angiopoietin-like 2 | 7080 | 0.001 | -0.1780 | No |
| 119 | TGFB1I1 | TGFB1I1 Entrez, Source | transforming growth factor beta 1 induced transcript 1 | 7121 | 0.001 | -0.1810 | No |
| 120 | BDKRB2 | BDKRB2 Entrez, Source | bradykinin receptor B2 | 7148 | 0.000 | -0.1830 | No |
| 121 | YTHDC1 | YTHDC1 Entrez, Source | YTH domain containing 1 | 7152 | 0.000 | -0.1832 | No |
| 122 | SKIV2L | SKIV2L Entrez, Source | superkiller viralicidic activity 2-like (S. cerevisiae) | 7168 | 0.000 | -0.1843 | No |
| 123 | SIPA1L1 | SIPA1L1 Entrez, Source | signal-induced proliferation-associated 1 like 1 | 7445 | -0.003 | -0.2052 | No |
| 124 | KLF6 | KLF6 Entrez, Source | Kruppel-like factor 6 | 7545 | -0.004 | -0.2126 | No |
| 125 | HNRPDL | HNRPDL Entrez, Source | heterogeneous nuclear ribonucleoprotein D-like | 7633 | -0.005 | -0.2190 | No |
| 126 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 7690 | -0.006 | -0.2230 | No |
| 127 | PRKAB2 | PRKAB2 Entrez, Source | protein kinase, AMP-activated, beta 2 non-catalytic subunit | 7852 | -0.008 | -0.2349 | No |
| 128 | NT5E | NT5E Entrez, Source | 5'-nucleotidase, ecto (CD73) | 8154 | -0.011 | -0.2573 | No |
| 129 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 8159 | -0.011 | -0.2570 | No |
| 130 | ORC4L | ORC4L Entrez, Source | origin recognition complex, subunit 4-like (yeast) | 8232 | -0.012 | -0.2619 | No |
| 131 | MPG | MPG Entrez, Source | N-methylpurine-DNA glycosylase | 8351 | -0.014 | -0.2703 | No |
| 132 | ARHGEF2 | ARHGEF2 Entrez, Source | rho/rac guanine nucleotide exchange factor (GEF) 2 | 8556 | -0.016 | -0.2851 | No |
| 133 | NIT1 | NIT1 Entrez, Source | nitrilase 1 | 8696 | -0.018 | -0.2948 | No |
| 134 | RGS4 | RGS4 Entrez, Source | regulator of G-protein signalling 4 | 8726 | -0.018 | -0.2961 | No |
| 135 | PTGFR | PTGFR Entrez, Source | prostaglandin F receptor (FP) | 8805 | -0.019 | -0.3012 | No |
| 136 | CUBN | CUBN Entrez, Source | cubilin (intrinsic factor-cobalamin receptor) | 8811 | -0.019 | -0.3006 | No |
| 137 | GZMK | GZMK Entrez, Source | granzyme K (granzyme 3; tryptase II) | 8919 | -0.021 | -0.3078 | No |
| 138 | INPP5B | INPP5B Entrez, Source | inositol polyphosphate-5-phosphatase, 75kDa | 8952 | -0.021 | -0.3092 | No |
| 139 | VDR | VDR Entrez, Source | vitamin D (1,25- dihydroxyvitamin D3) receptor | 9079 | -0.023 | -0.3177 | No |
| 140 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 9147 | -0.024 | -0.3216 | No |
| 141 | REV3L | REV3L Entrez, Source | REV3-like, catalytic subunit of DNA polymerase zeta (yeast) | 9194 | -0.024 | -0.3240 | No |
| 142 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 9384 | -0.027 | -0.3371 | No |
| 143 | RBM10 | RBM10 Entrez, Source | RNA binding motif protein 10 | 9414 | -0.028 | -0.3379 | No |
| 144 | OSR2 | OSR2 Entrez, Source | odd-skipped related 2 (Drosophila) | 9457 | -0.028 | -0.3398 | No |
| 145 | MYO1B | MYO1B Entrez, Source | myosin IB | 9460 | -0.028 | -0.3386 | No |
| 146 | ATP2A2 | ATP2A2 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 | 9652 | -0.031 | -0.3516 | No |
| 147 | RAB3B | RAB3B Entrez, Source | RAB3B, member RAS oncogene family | 9706 | -0.032 | -0.3541 | No |
| 148 | AURKA | AURKA Entrez, Source | aurora kinase A | 9722 | -0.032 | -0.3537 | No |
| 149 | RND3 | RND3 Entrez, Source | Rho family GTPase 3 | 9751 | -0.033 | -0.3543 | No |
| 150 | PPP2R1B | PPP2R1B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), beta isoform | 9775 | -0.033 | -0.3544 | No |
| 151 | MGEA5 | MGEA5 Entrez, Source | meningioma expressed antigen 5 (hyaluronidase) | 10016 | -0.037 | -0.3709 | No |
| 152 | CHEK2 | CHEK2 Entrez, Source | CHK2 checkpoint homolog (S. pombe) | 10020 | -0.037 | -0.3694 | No |
| 153 | SOCS1 | SOCS1 Entrez, Source | suppressor of cytokine signaling 1 | 10039 | -0.037 | -0.3689 | No |
| 154 | GREM1 | GREM1 Entrez, Source | gremlin 1, cysteine knot superfamily, homolog (Xenopus laevis) | 10120 | -0.039 | -0.3732 | No |
| 155 | PRKAB1 | PRKAB1 Entrez, Source | protein kinase, AMP-activated, beta 1 non-catalytic subunit | 10265 | -0.041 | -0.3822 | No |
| 156 | ALDH7A1 | ALDH7A1 Entrez, Source | aldehyde dehydrogenase 7 family, member A1 | 10287 | -0.041 | -0.3818 | No |
| 157 | ODF2 | ODF2 Entrez, Source | outer dense fiber of sperm tails 2 | 10350 | -0.042 | -0.3845 | No |
| 158 | ZFP36L2 | ZFP36L2 Entrez, Source | zinc finger protein 36, C3H type-like 2 | 10449 | -0.044 | -0.3898 | No |
| 159 | VAPB | VAPB Entrez, Source | VAMP (vesicle-associated membrane protein)-associated protein B and C | 10452 | -0.044 | -0.3879 | No |
| 160 | SLC6A15 | SLC6A15 Entrez, Source | solute carrier family 6, member 15 | 10453 | -0.044 | -0.3857 | No |
| 161 | AGGF1 | AGGF1 Entrez, Source | angiogenic factor with G patch and FHA domains 1 | 10582 | -0.047 | -0.3932 | No |
| 162 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 10619 | -0.047 | -0.3937 | No |
| 163 | ATXN10 | ATXN10 Entrez, Source | ataxin 10 | 10662 | -0.048 | -0.3946 | No |
| 164 | CBR4 | CBR4 Entrez, Source | carbonyl reductase 4 | 10676 | -0.049 | -0.3932 | No |
| 165 | DNASE1L1 | DNASE1L1 Entrez, Source | deoxyribonuclease I-like 1 | 10694 | -0.049 | -0.3921 | No |
| 166 | PRIM2A | PRIM2A Entrez, Source | primase, polypeptide 2A, 58kDa | 10779 | -0.051 | -0.3961 | No |
| 167 | PKIG | PKIG Entrez, Source | protein kinase (cAMP-dependent, catalytic) inhibitor gamma | 10784 | -0.051 | -0.3939 | No |
| 168 | ACPP | ACPP Entrez, Source | acid phosphatase, prostate | 10827 | -0.052 | -0.3946 | No |
| 169 | GLMN | GLMN Entrez, Source | glomulin, FKBP associated protein | 10862 | -0.053 | -0.3947 | No |
| 170 | SERPINB2 | SERPINB2 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 2 | 10863 | -0.053 | -0.3921 | No |
| 171 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 10913 | -0.054 | -0.3932 | No |
| 172 | ANP32A | ANP32A Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member A | 10934 | -0.054 | -0.3921 | No |
| 173 | CHN1 | CHN1 Entrez, Source | chimerin (chimaerin) 1 | 10953 | -0.055 | -0.3908 | No |
| 174 | MLLT11 | MLLT11 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 11 | 11034 | -0.057 | -0.3942 | No |
| 175 | MNAT1 | MNAT1 Entrez, Source | menage a trois homolog 1, cyclin H assembly factor (Xenopus laevis) | 11046 | -0.057 | -0.3923 | No |
| 176 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 11131 | -0.059 | -0.3959 | No |
| 177 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 11148 | -0.059 | -0.3943 | No |
| 178 | CSPG2 | CSPG2 Entrez, Source | chondroitin sulfate proteoglycan 2 (versican) | 11206 | -0.060 | -0.3957 | No |
| 179 | AXL | AXL Entrez, Source | AXL receptor tyrosine kinase | 11348 | -0.064 | -0.4034 | No |
| 180 | AHR | AHR Entrez, Source | aryl hydrocarbon receptor | 11457 | -0.067 | -0.4084 | No |
| 181 | GABBR2 | GABBR2 Entrez, Source | gamma-aminobutyric acid (GABA) B receptor, 2 | 11619 | -0.073 | -0.4172 | No |
| 182 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 11651 | -0.074 | -0.4160 | No |
| 183 | FZD2 | FZD2 Entrez, Source | frizzled homolog 2 (Drosophila) | 11656 | -0.074 | -0.4127 | No |
| 184 | FGF2 | FGF2 Entrez, Source | fibroblast growth factor 2 (basic) | 11687 | -0.075 | -0.4114 | No |
| 185 | CIRBP | CIRBP Entrez, Source | cold inducible RNA binding protein | 11768 | -0.078 | -0.4137 | No |
| 186 | UBL3 | UBL3 Entrez, Source | ubiquitin-like 3 | 11776 | -0.078 | -0.4105 | No |
| 187 | RUFY3 | RUFY3 Entrez, Source | RUN and FYVE domain containing 3 | 11970 | -0.086 | -0.4211 | Yes |
| 188 | CDH13 | CDH13 Entrez, Source | cadherin 13, H-cadherin (heart) | 11996 | -0.087 | -0.4188 | Yes |
| 189 | TPD52L2 | TPD52L2 Entrez, Source | tumor protein D52-like 2 | 11997 | -0.087 | -0.4146 | Yes |
| 190 | GCNT1 | GCNT1 Entrez, Source | glucosaminyl (N-acetyl) transferase 1, core 2 (beta-1,6-N-acetylglucosaminyltransferase) | 12045 | -0.089 | -0.4139 | Yes |
| 191 | FABP3 | FABP3 Entrez, Source | fatty acid binding protein 3, muscle and heart (mammary-derived growth inhibitor) | 12159 | -0.094 | -0.4180 | Yes |
| 192 | KIFC1 | KIFC1 Entrez, Source | kinesin family member C1 | 12256 | -0.100 | -0.4205 | Yes |
| 193 | PDCD4 | PDCD4 Entrez, Source | programmed cell death 4 (neoplastic transformation inhibitor) | 12282 | -0.102 | -0.4175 | Yes |
| 194 | VAMP5 | VAMP5 Entrez, Source | vesicle-associated membrane protein 5 (myobrevin) | 12296 | -0.103 | -0.4135 | Yes |
| 195 | PIAS3 | PIAS3 Entrez, Source | protein inhibitor of activated STAT, 3 | 12301 | -0.103 | -0.4088 | Yes |
| 196 | MGMT | MGMT Entrez, Source | O-6-methylguanine-DNA methyltransferase | 12359 | -0.108 | -0.4079 | Yes |
| 197 | TGIF | TGIF Entrez, Source | TGFB-induced factor (TALE family homeobox) | 12388 | -0.110 | -0.4048 | Yes |
| 198 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 12473 | -0.116 | -0.4056 | Yes |
| 199 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 12519 | -0.119 | -0.4033 | Yes |
| 200 | GAS6 | GAS6 Entrez, Source | growth arrest-specific 6 | 12538 | -0.120 | -0.3989 | Yes |
| 201 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 12557 | -0.122 | -0.3943 | Yes |
| 202 | GSTA4 | GSTA4 Entrez, Source | glutathione S-transferase A4 | 12608 | -0.126 | -0.3921 | Yes |
| 203 | VEGFC | VEGFC Entrez, Source | vascular endothelial growth factor C | 12665 | -0.131 | -0.3900 | Yes |
| 204 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 12686 | -0.133 | -0.3851 | Yes |
| 205 | ZBTB1 | ZBTB1 Entrez, Source | zinc finger and BTB domain containing 1 | 12701 | -0.135 | -0.3797 | Yes |
| 206 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 12774 | -0.145 | -0.3782 | Yes |
| 207 | TRIM2 | TRIM2 Entrez, Source | tripartite motif-containing 2 | 12840 | -0.154 | -0.3757 | Yes |
| 208 | ALDH3B1 | ALDH3B1 Entrez, Source | aldehyde dehydrogenase 3 family, member B1 | 12850 | -0.156 | -0.3689 | Yes |
| 209 | NMT2 | NMT2 Entrez, Source | N-myristoyltransferase 2 | 12913 | -0.168 | -0.3655 | Yes |
| 210 | PDE4B | PDE4B Entrez, Source | phosphodiesterase 4B, cAMP-specific (phosphodiesterase E4 dunce homolog, Drosophila) | 12954 | -0.177 | -0.3601 | Yes |
| 211 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 12980 | -0.185 | -0.3531 | Yes |
| 212 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 13105 | -0.234 | -0.3512 | Yes |
| 213 | RRAS2 | RRAS2 Entrez, Source | related RAS viral (r-ras) oncogene homolog 2 | 13125 | -0.244 | -0.3409 | Yes |
| 214 | CDC42EP2 | CDC42EP2 Entrez, Source | CDC42 effector protein (Rho GTPase binding) 2 | 13158 | -0.264 | -0.3306 | Yes |
| 215 | HSPA4 | HSPA4 Entrez, Source | heat shock 70kDa protein 4 | 13162 | -0.265 | -0.3180 | Yes |
| 216 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 13180 | -0.281 | -0.3058 | Yes |
| 217 | F2R | F2R Entrez, Source | coagulation factor II (thrombin) receptor | 13224 | -0.321 | -0.2936 | Yes |
| 218 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 13226 | -0.322 | -0.2782 | Yes |
| 219 | ADORA2B | ADORA2B Entrez, Source | adenosine A2b receptor | 13236 | -0.333 | -0.2628 | Yes |
| 220 | SLC22A4 | SLC22A4 Entrez, Source | solute carrier family 22 (organic cation transporter), member 4 | 13246 | -0.348 | -0.2467 | Yes |
| 221 | GPRC5A | GPRC5A Entrez, Source | G protein-coupled receptor, family C, group 5, member A | 13268 | -0.382 | -0.2299 | Yes |
| 222 | ADM | ADM Entrez, Source | adrenomedullin | 13274 | -0.393 | -0.2113 | Yes |
| 223 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 13281 | -0.405 | -0.1922 | Yes |
| 224 | SERPINH1 | SERPINH1 Entrez, Source | serpin peptidase inhibitor, clade H (heat shock protein 47), member 1, (collagen binding protein 1) | 13296 | -0.454 | -0.1714 | Yes |
| 225 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 13307 | -0.479 | -0.1490 | Yes |
| 226 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 13308 | -0.484 | -0.1257 | Yes |
| 227 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 13312 | -0.498 | -0.1019 | Yes |
| 228 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13313 | -0.503 | -0.0776 | Yes |
| 229 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 13315 | -0.511 | -0.0530 | Yes |
| 230 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 13316 | -0.512 | -0.0283 | Yes |
| 231 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 13326 | -0.627 | 0.0012 | Yes |