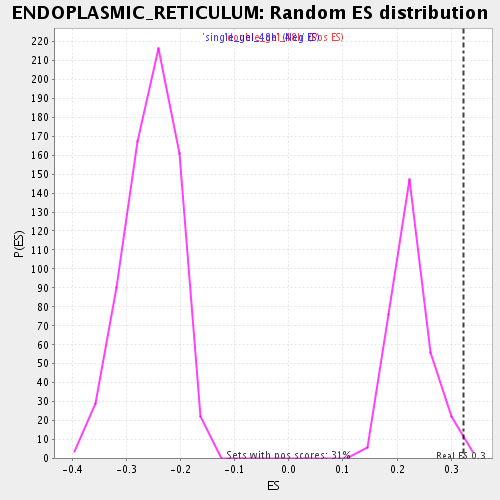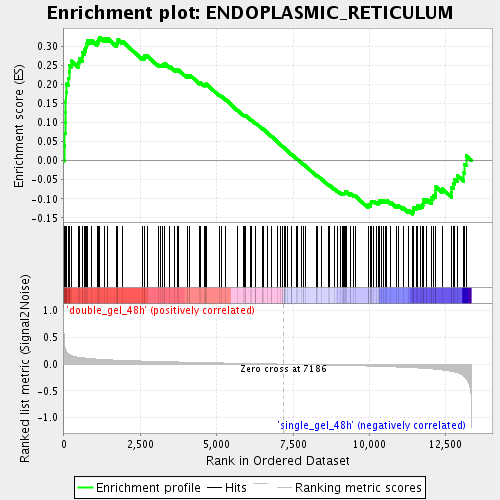
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | ENDOPLASMIC_RETICULUM |
| Enrichment Score (ES) | 0.3231985 |
| Normalized Enrichment Score (NES) | 1.4314599 |
| Nominal p-value | 0.003215434 |
| FDR q-value | 0.18969141 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 12 | 0.395 | 0.0388 | Yes |
| 2 | TM7SF2 | TM7SF2 Entrez, Source | transmembrane 7 superfamily member 2 | 24 | 0.340 | 0.0721 | Yes |
| 3 | EBP | EBP Entrez, Source | emopamil binding protein (sterol isomerase) | 45 | 0.290 | 0.0998 | Yes |
| 4 | HMGCR | HMGCR Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A reductase | 55 | 0.278 | 0.1270 | Yes |
| 5 | AADAC | AADAC Entrez, Source | arylacetamide deacetylase (esterase) | 59 | 0.269 | 0.1538 | Yes |
| 6 | HYOU1 | HYOU1 Entrez, Source | hypoxia up-regulated 1 | 68 | 0.255 | 0.1788 | Yes |
| 7 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 90 | 0.236 | 0.2010 | Yes |
| 8 | PDIA4 | PDIA4 Entrez, Source | protein disulfide isomerase family A, member 4 | 151 | 0.192 | 0.2157 | Yes |
| 9 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 167 | 0.185 | 0.2331 | Yes |
| 10 | RTN1 | RTN1 Entrez, Source | reticulon 1 | 181 | 0.178 | 0.2501 | Yes |
| 11 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 247 | 0.160 | 0.2612 | Yes |
| 12 | OPRM1 | OPRM1 Entrez, Source | opioid receptor, mu 1 | 481 | 0.127 | 0.2563 | Yes |
| 13 | PEX16 | PEX16 Entrez, Source | peroxisomal biogenesis factor 16 | 496 | 0.125 | 0.2678 | Yes |
| 14 | CLDN14 | CLDN14 Entrez, Source | claudin 14 | 599 | 0.118 | 0.2719 | Yes |
| 15 | LMAN1 | LMAN1 Entrez, Source | lectin, mannose-binding, 1 | 607 | 0.117 | 0.2832 | Yes |
| 16 | CYP19A1 | CYP19A1 Entrez, Source | cytochrome P450, family 19, subfamily A, polypeptide 1 | 670 | 0.114 | 0.2899 | Yes |
| 17 | HERPUD1 | HERPUD1 Entrez, Source | homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 | 716 | 0.111 | 0.2977 | Yes |
| 18 | THY1 | THY1 Entrez, Source | Thy-1 cell surface antigen | 748 | 0.109 | 0.3063 | Yes |
| 19 | CALR | CALR Entrez, Source | calreticulin | 768 | 0.108 | 0.3157 | Yes |
| 20 | CD320 | CD320 Entrez, Source | CD320 molecule | 901 | 0.102 | 0.3160 | Yes |
| 21 | SERP1 | SERP1 Entrez, Source | - | 1084 | 0.094 | 0.3116 | Yes |
| 22 | SSR1 | SSR1 Entrez, Source | signal sequence receptor, alpha (translocon-associated protein alpha) | 1133 | 0.092 | 0.3172 | Yes |
| 23 | ARSG | ARSG Entrez, Source | arylsulfatase G | 1175 | 0.090 | 0.3232 | Yes |
| 24 | UGCGL1 | UGCGL1 Entrez, Source | UDP-glucose ceramide glucosyltransferase-like 1 | 1335 | 0.085 | 0.3197 | No |
| 25 | PTN | PTN Entrez, Source | pleiotrophin (heparin binding growth factor 8, neurite growth-promoting factor 1) | 1439 | 0.081 | 0.3200 | No |
| 26 | CLDN8 | CLDN8 Entrez, Source | claudin 8 | 1724 | 0.074 | 0.3059 | No |
| 27 | DNAJB9 | DNAJB9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 9 | 1749 | 0.073 | 0.3114 | No |
| 28 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 1768 | 0.073 | 0.3174 | No |
| 29 | SLC33A1 | SLC33A1 Entrez, Source | solute carrier family 33 (acetyl-CoA transporter), member 1 | 1918 | 0.070 | 0.3131 | No |
| 30 | POR | POR Entrez, Source | P450 (cytochrome) oxidoreductase | 2561 | 0.058 | 0.2702 | No |
| 31 | DNAJB11 | DNAJB11 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 11 | 2627 | 0.056 | 0.2709 | No |
| 32 | HSD3B1 | HSD3B1 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 | 2646 | 0.056 | 0.2752 | No |
| 33 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 2730 | 0.055 | 0.2744 | No |
| 34 | RTN4R | RTN4R Entrez, Source | reticulon 4 receptor | 3109 | 0.049 | 0.2506 | No |
| 35 | GCS1 | GCS1 Entrez, Source | - | 3175 | 0.048 | 0.2505 | No |
| 36 | DNM1L | DNM1L Entrez, Source | dynamin 1-like | 3235 | 0.047 | 0.2507 | No |
| 37 | DHRS9 | DHRS9 Entrez, Source | dehydrogenase/reductase (SDR family) member 9 | 3288 | 0.046 | 0.2514 | No |
| 38 | RPN1 | RPN1 Entrez, Source | ribophorin I | 3301 | 0.046 | 0.2551 | No |
| 39 | ITPR2 | ITPR2 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 2 | 3468 | 0.043 | 0.2469 | No |
| 40 | GBA2 | GBA2 Entrez, Source | glucosidase, beta (bile acid) 2 | 3630 | 0.041 | 0.2388 | No |
| 41 | SRPR | SRPR Entrez, Source | signal recognition particle receptor ('docking protein') | 3713 | 0.040 | 0.2366 | No |
| 42 | CANX | CANX Entrez, Source | calnexin | 3753 | 0.040 | 0.2377 | No |
| 43 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 4028 | 0.036 | 0.2205 | No |
| 44 | KDELR2 | KDELR2 Entrez, Source | KDEL (Lys-Asp-Glu-Leu) endoplasmic reticulum protein retention receptor 2 | 4041 | 0.036 | 0.2232 | No |
| 45 | SREBF1 | SREBF1 Entrez, Source | sterol regulatory element binding transcription factor 1 | 4109 | 0.035 | 0.2217 | No |
| 46 | PDIA5 | PDIA5 Entrez, Source | protein disulfide isomerase family A, member 5 | 4124 | 0.035 | 0.2241 | No |
| 47 | GPC2 | GPC2 Entrez, Source | glypican 2 (cerebroglycan) | 4452 | 0.031 | 0.2024 | No |
| 48 | S100A1 | S100A1 Entrez, Source | S100 calcium binding protein A1 | 4459 | 0.030 | 0.2050 | No |
| 49 | P4HB | P4HB Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), beta polypeptide | 4586 | 0.029 | 0.1983 | No |
| 50 | PDIA6 | PDIA6 Entrez, Source | protein disulfide isomerase family A, member 6 | 4624 | 0.028 | 0.1984 | No |
| 51 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 4640 | 0.028 | 0.2001 | No |
| 52 | ACO1 | ACO1 Entrez, Source | aconitase 1, soluble | 4655 | 0.028 | 0.2018 | No |
| 53 | PDIA3 | PDIA3 Entrez, Source | protein disulfide isomerase family A, member 3 | 5084 | 0.023 | 0.1717 | No |
| 54 | BNIP1 | BNIP1 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 1 | 5150 | 0.022 | 0.1689 | No |
| 55 | VWF | VWF Entrez, Source | von Willebrand factor | 5296 | 0.020 | 0.1600 | No |
| 56 | RPN2 | RPN2 Entrez, Source | ribophorin II | 5686 | 0.016 | 0.1321 | No |
| 57 | PLA2G2A | PLA2G2A Entrez, Source | phospholipase A2, group IIA (platelets, synovial fluid) | 5884 | 0.014 | 0.1186 | No |
| 58 | SEMA4F | SEMA4F Entrez, Source | sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4F | 5918 | 0.014 | 0.1175 | No |
| 59 | PIGV | PIGV Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class V | 5939 | 0.013 | 0.1173 | No |
| 60 | ADFP | ADFP Entrez, Source | adipose differentiation-related protein | 5942 | 0.013 | 0.1185 | No |
| 61 | NCSTN | NCSTN Entrez, Source | nicastrin | 6104 | 0.012 | 0.1075 | No |
| 62 | CLN3 | CLN3 Entrez, Source | ceroid-lipofuscinosis, neuronal 3, juvenile (Batten, Spielmeyer-Vogt disease) | 6139 | 0.012 | 0.1061 | No |
| 63 | TRPC4AP | TRPC4AP Entrez, Source | transient receptor potential cation channel, subfamily C, member 4 associated protein | 6253 | 0.010 | 0.0985 | No |
| 64 | STX18 | STX18 Entrez, Source | syntaxin 18 | 6278 | 0.010 | 0.0977 | No |
| 65 | TEGT | TEGT Entrez, Source | testis enhanced gene transcript (BAX inhibitor 1) | 6284 | 0.010 | 0.0983 | No |
| 66 | CES2 | CES2 Entrez, Source | carboxylesterase 2 (intestine, liver) | 6501 | 0.008 | 0.0827 | No |
| 67 | SV2A | SV2A Entrez, Source | synaptic vesicle glycoprotein 2A | 6513 | 0.007 | 0.0826 | No |
| 68 | PPIB | PPIB Entrez, Source | peptidylprolyl isomerase B (cyclophilin B) | 6516 | 0.007 | 0.0832 | No |
| 69 | BSCL2 | BSCL2 Entrez, Source | Bernardinelli-Seip congenital lipodystrophy 2 (seipin) | 6646 | 0.006 | 0.0740 | No |
| 70 | MAL | MAL Entrez, Source | mal, T-cell differentiation protein | 6779 | 0.004 | 0.0645 | No |
| 71 | SYVN1 | SYVN1 Entrez, Source | synovial apoptosis inhibitor 1, synoviolin | 6805 | 0.004 | 0.0630 | No |
| 72 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6985 | 0.002 | 0.0496 | No |
| 73 | RETSAT | RETSAT Entrez, Source | retinol saturase (all-trans-retinol 13,14-reductase) | 7090 | 0.001 | 0.0418 | No |
| 74 | RYR1 | RYR1 Entrez, Source | ryanodine receptor 1 (skeletal) | 7158 | 0.000 | 0.0368 | No |
| 75 | PSENEN | PSENEN Entrez, Source | presenilin enhancer 2 homolog (C. elegans) | 7162 | 0.000 | 0.0366 | No |
| 76 | RPL10 | RPL10 Entrez, Source | ribosomal protein L10 | 7202 | -0.000 | 0.0337 | No |
| 77 | NPLOC4 | NPLOC4 Entrez, Source | nuclear protein localization 4 homolog (S. cerevisiae) | 7204 | -0.000 | 0.0336 | No |
| 78 | ERO1L | ERO1L Entrez, Source | ERO1-like (S. cerevisiae) | 7209 | -0.000 | 0.0333 | No |
| 79 | GNAZ | GNAZ Entrez, Source | guanine nucleotide binding protein (G protein), alpha z polypeptide | 7248 | -0.001 | 0.0305 | No |
| 80 | ATP1A3 | ATP1A3 Entrez, Source | ATPase, Na+/K+ transporting, alpha 3 polypeptide | 7330 | -0.002 | 0.0246 | No |
| 81 | ZW10 | ZW10 Entrez, Source | ZW10, kinetochore associated, homolog (Drosophila) | 7453 | -0.003 | 0.0156 | No |
| 82 | APOC1 | APOC1 Entrez, Source | apolipoprotein C-I | 7456 | -0.003 | 0.0158 | No |
| 83 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 7461 | -0.003 | 0.0158 | No |
| 84 | TPST2 | TPST2 Entrez, Source | tyrosylprotein sulfotransferase 2 | 7610 | -0.005 | 0.0051 | No |
| 85 | SLC36A1 | SLC36A1 Entrez, Source | solute carrier family 36 (proton/amino acid symporter), member 1 | 7653 | -0.005 | 0.0025 | No |
| 86 | APOA5 | APOA5 Entrez, Source | apolipoprotein A-V | 7763 | -0.007 | -0.0051 | No |
| 87 | ICMT | ICMT Entrez, Source | isoprenylcysteine carboxyl methyltransferase | 7827 | -0.007 | -0.0092 | No |
| 88 | VCP | VCP Entrez, Source | valosin-containing protein | 7832 | -0.007 | -0.0087 | No |
| 89 | AGPAT1 | AGPAT1 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) | 7919 | -0.009 | -0.0144 | No |
| 90 | PDZD2 | PDZD2 Entrez, Source | PDZ domain containing 2 | 8263 | -0.013 | -0.0391 | No |
| 91 | CTSZ | CTSZ Entrez, Source | cathepsin Z | 8280 | -0.013 | -0.0391 | No |
| 92 | PIGK | PIGK Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class K | 8287 | -0.013 | -0.0382 | No |
| 93 | MBTPS1 | MBTPS1 Entrez, Source | membrane-bound transcription factor peptidase, site 1 | 8426 | -0.015 | -0.0472 | No |
| 94 | SERINC1 | SERINC1 Entrez, Source | serine incorporator 1 | 8655 | -0.018 | -0.0628 | No |
| 95 | ALG3 | ALG3 Entrez, Source | asparagine-linked glycosylation 3 homolog (S. cerevisiae, alpha-1,3-mannosyltransferase) | 8704 | -0.018 | -0.0646 | No |
| 96 | DAD1 | DAD1 Entrez, Source | defender against cell death 1 | 8855 | -0.020 | -0.0740 | No |
| 97 | MINPP1 | MINPP1 Entrez, Source | multiple inositol polyphosphate histidine phosphatase, 1 | 8960 | -0.021 | -0.0797 | No |
| 98 | DULLARD | DULLARD Entrez, Source | dullard homolog (Xenopus laevis) | 9054 | -0.023 | -0.0845 | No |
| 99 | TOR1A | TOR1A Entrez, Source | torsin family 1, member A (torsin A) | 9104 | -0.023 | -0.0859 | No |
| 100 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 9147 | -0.024 | -0.0867 | No |
| 101 | PLOD3 | PLOD3 Entrez, Source | procollagen-lysine, 2-oxoglutarate 5-dioxygenase 3 | 9156 | -0.024 | -0.0849 | No |
| 102 | STAU1 | STAU1 Entrez, Source | staufen, RNA binding protein, homolog 1 (Drosophila) | 9189 | -0.024 | -0.0849 | No |
| 103 | CLN8 | CLN8 Entrez, Source | ceroid-lipofuscinosis, neuronal 8 (epilepsy, progressive with mental retardation) | 9201 | -0.024 | -0.0832 | No |
| 104 | TXNDC4 | TXNDC4 Entrez, Source | thioredoxin domain containing 4 (endoplasmic reticulum) | 9203 | -0.024 | -0.0809 | No |
| 105 | TAS2R16 | TAS2R16 Entrez, Source | taste receptor, type 2, member 16 | 9240 | -0.025 | -0.0811 | No |
| 106 | POMT1 | POMT1 Entrez, Source | protein-O-mannosyltransferase 1 | 9373 | -0.027 | -0.0884 | No |
| 107 | SEP15 | SEP15 Entrez, Source | - | 9382 | -0.027 | -0.0862 | No |
| 108 | RAB14 | RAB14 Entrez, Source | RAB14, member RAS oncogene family | 9475 | -0.029 | -0.0904 | No |
| 109 | YIPF4 | YIPF4 Entrez, Source | Yip1 domain family, member 4 | 9529 | -0.029 | -0.0914 | No |
| 110 | STS | STS Entrez, Source | steroid sulfatase (microsomal), arylsulfatase C, isozyme S | 9956 | -0.036 | -0.1201 | No |
| 111 | TMEM50B | TMEM50B Entrez, Source | transmembrane protein 50B | 9970 | -0.036 | -0.1175 | No |
| 112 | AVPR2 | AVPR2 Entrez, Source | arginine vasopressin receptor 2 (nephrogenic diabetes insipidus) | 9972 | -0.036 | -0.1139 | No |
| 113 | BDKRB1 | BDKRB1 Entrez, Source | bradykinin receptor B1 | 10044 | -0.037 | -0.1155 | No |
| 114 | YKT6 | YKT6 Entrez, Source | YKT6 v-SNARE homolog (S. cerevisiae) | 10045 | -0.038 | -0.1118 | No |
| 115 | PCMT1 | PCMT1 Entrez, Source | protein-L-isoaspartate (D-aspartate) O-methyltransferase | 10054 | -0.038 | -0.1086 | No |
| 116 | PIGC | PIGC Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class C | 10066 | -0.038 | -0.1056 | No |
| 117 | SEC22A | SEC22A Entrez, Source | SEC22 vesicle trafficking protein homolog A (S. cerevisiae) | 10135 | -0.039 | -0.1069 | No |
| 118 | DPM2 | DPM2 Entrez, Source | dolichyl-phosphate mannosyltransferase polypeptide 2, regulatory subunit | 10215 | -0.040 | -0.1088 | No |
| 119 | BAX | BAX Entrez, Source | BCL2-associated X protein | 10281 | -0.041 | -0.1096 | No |
| 120 | AKAP6 | AKAP6 Entrez, Source | A kinase (PRKA) anchor protein 6 | 10317 | -0.042 | -0.1081 | No |
| 121 | ITPR3 | ITPR3 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 3 | 10318 | -0.042 | -0.1039 | No |
| 122 | TRAM1 | TRAM1 Entrez, Source | translocation associated membrane protein 1 | 10392 | -0.043 | -0.1051 | No |
| 123 | VAPB | VAPB Entrez, Source | VAMP (vesicle-associated membrane protein)-associated protein B and C | 10452 | -0.044 | -0.1051 | No |
| 124 | ITGB1 | ITGB1 Entrez, Source | integrin, beta 1 (fibronectin receptor, beta polypeptide, antigen CD29 includes MDF2, MSK12) | 10521 | -0.046 | -0.1057 | No |
| 125 | PSEN1 | PSEN1 Entrez, Source | presenilin 1 (Alzheimer disease 3) | 10571 | -0.046 | -0.1047 | No |
| 126 | PIGS | PIGS Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class S | 10690 | -0.049 | -0.1087 | No |
| 127 | CIB1 | CIB1 Entrez, Source | calcium and integrin binding 1 (calmyrin) | 10883 | -0.053 | -0.1180 | No |
| 128 | ANP32A | ANP32A Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member A | 10934 | -0.054 | -0.1163 | No |
| 129 | TXNDC1 | TXNDC1 Entrez, Source | thioredoxin domain containing 1 | 11099 | -0.058 | -0.1229 | No |
| 130 | STX8 | STX8 Entrez, Source | syntaxin 8 | 11276 | -0.062 | -0.1300 | No |
| 131 | FKBP9 | FKBP9 Entrez, Source | FK506 binding protein 9, 63 kDa | 11407 | -0.065 | -0.1333 | No |
| 132 | EIF2AK3 | EIF2AK3 Entrez, Source | eukaryotic translation initiation factor 2-alpha kinase 3 | 11434 | -0.066 | -0.1287 | No |
| 133 | CASP7 | CASP7 Entrez, Source | caspase 7, apoptosis-related cysteine peptidase | 11446 | -0.066 | -0.1228 | No |
| 134 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 11526 | -0.069 | -0.1219 | No |
| 135 | TMEM98 | TMEM98 Entrez, Source | transmembrane protein 98 | 11558 | -0.070 | -0.1171 | No |
| 136 | KCNG3 | KCNG3 Entrez, Source | potassium voltage-gated channel, subfamily G, member 3 | 11675 | -0.075 | -0.1184 | No |
| 137 | PSEN2 | PSEN2 Entrez, Source | presenilin 2 (Alzheimer disease 4) | 11726 | -0.076 | -0.1146 | No |
| 138 | APH1A | APH1A Entrez, Source | anterior pharynx defective 1 homolog A (C. elegans) | 11754 | -0.077 | -0.1089 | No |
| 139 | TAP2 | TAP2 Entrez, Source | transporter 2, ATP-binding cassette, sub-family B (MDR/TAP) | 11767 | -0.078 | -0.1020 | No |
| 140 | ERP29 | ERP29 Entrez, Source | endoplasmic reticulum protein 29 | 11877 | -0.082 | -0.1020 | No |
| 141 | TLOC1 | TLOC1 Entrez, Source | translocation protein 1 | 12024 | -0.088 | -0.1041 | No |
| 142 | AGPAT6 | AGPAT6 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 6 (lysophosphatidic acid acyltransferase, zeta) | 12037 | -0.089 | -0.0961 | No |
| 143 | DHRS1 | DHRS1 Entrez, Source | dehydrogenase/reductase (SDR family) member 1 | 12095 | -0.091 | -0.0913 | No |
| 144 | CAMLG | CAMLG Entrez, Source | calcium modulating ligand | 12165 | -0.095 | -0.0870 | No |
| 145 | TAPBP | TAPBP Entrez, Source | TAP binding protein (tapasin) | 12170 | -0.095 | -0.0777 | No |
| 146 | HTRA2 | HTRA2 Entrez, Source | HtrA serine peptidase 2 | 12173 | -0.095 | -0.0683 | No |
| 147 | ABI1 | ABI1 Entrez, Source | abl-interactor 1 | 12376 | -0.109 | -0.0727 | No |
| 148 | HSD17B2 | HSD17B2 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 2 | 12678 | -0.132 | -0.0822 | No |
| 149 | SOAT1 | SOAT1 Entrez, Source | sterol O-acyltransferase (acyl-Coenzyme A: cholesterol acyltransferase) 1 | 12692 | -0.134 | -0.0697 | No |
| 150 | EHD4 | EHD4 Entrez, Source | EH-domain containing 4 | 12744 | -0.140 | -0.0595 | No |
| 151 | P4HA1 | P4HA1 Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha polypeptide I | 12791 | -0.146 | -0.0483 | No |
| 152 | BNIP3L | BNIP3L Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3-like | 12870 | -0.160 | -0.0382 | No |
| 153 | PRNP | PRNP Entrez, Source | prion protein (p27-30) (Creutzfeldt-Jakob disease, Gerstmann-Strausler-Scheinker syndrome, fatal familial insomnia) | 13090 | -0.227 | -0.0320 | No |
| 154 | RCN2 | RCN2 Entrez, Source | reticulocalbin 2, EF-hand calcium binding domain | 13116 | -0.238 | -0.0099 | No |
| 155 | RTN4 | RTN4 Entrez, Source | reticulon 4 | 13167 | -0.269 | 0.0133 | No |