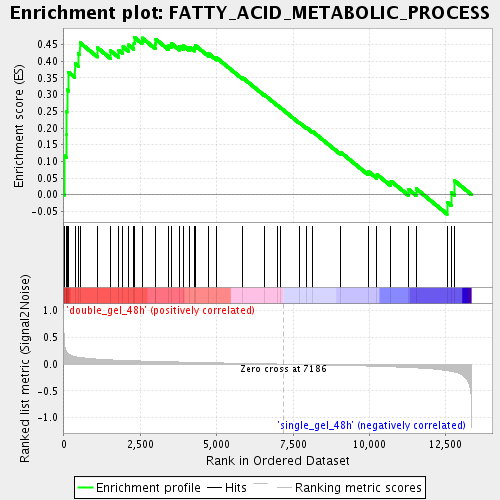
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | FATTY_ACID_METABOLIC_PROCESS |
| Enrichment Score (ES) | 0.47184515 |
| Normalized Enrichment Score (NES) | 1.6292616 |
| Nominal p-value | 0.007556675 |
| FDR q-value | 0.08511103 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 12 | 0.395 | 0.1163 | Yes |
| 2 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 90 | 0.236 | 0.1806 | Yes |
| 3 | HACL1 | HACL1 Entrez, Source | 2-hydroxyacyl-CoA lyase 1 | 94 | 0.229 | 0.2483 | Yes |
| 4 | GLYAT | GLYAT Entrez, Source | glycine-N-acyltransferase | 102 | 0.225 | 0.3144 | Yes |
| 5 | PPARA | PPARA Entrez, Source | peroxisome proliferative activated receptor, alpha | 155 | 0.190 | 0.3670 | Yes |
| 6 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 362 | 0.140 | 0.3931 | Yes |
| 7 | CPT1A | CPT1A Entrez, Source | carnitine palmitoyltransferase 1A (liver) | 469 | 0.128 | 0.4232 | Yes |
| 8 | ACADVL | ACADVL Entrez, Source | acyl-Coenzyme A dehydrogenase, very long chain | 526 | 0.123 | 0.4555 | Yes |
| 9 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 1097 | 0.093 | 0.4403 | Yes |
| 10 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 1528 | 0.079 | 0.4313 | Yes |
| 11 | PTGES | PTGES Entrez, Source | prostaglandin E synthase | 1797 | 0.072 | 0.4326 | Yes |
| 12 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 1932 | 0.070 | 0.4431 | Yes |
| 13 | FAAH | FAAH Entrez, Source | fatty acid amide hydrolase | 2116 | 0.066 | 0.4488 | Yes |
| 14 | CYP4F2 | CYP4F2 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 2 | 2291 | 0.062 | 0.4542 | Yes |
| 15 | MLYCD | MLYCD Entrez, Source | malonyl-CoA decarboxylase | 2302 | 0.062 | 0.4718 | Yes |
| 16 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 2558 | 0.058 | 0.4698 | No |
| 17 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 2994 | 0.050 | 0.4519 | No |
| 18 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 3007 | 0.050 | 0.4658 | No |
| 19 | ACOT2 | ACOT2 Entrez, Source | acyl-CoA thioesterase 2 | 3435 | 0.044 | 0.4467 | No |
| 20 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 3526 | 0.043 | 0.4526 | No |
| 21 | GNPAT | GNPAT Entrez, Source | glyceronephosphate O-acyltransferase | 3783 | 0.039 | 0.4450 | No |
| 22 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 3915 | 0.038 | 0.4463 | No |
| 23 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 4114 | 0.035 | 0.4418 | No |
| 24 | PECI | PECI Entrez, Source | peroxisomal D3,D2-enoyl-CoA isomerase | 4265 | 0.033 | 0.4403 | No |
| 25 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 4313 | 0.032 | 0.4464 | No |
| 26 | ACOX3 | ACOX3 Entrez, Source | acyl-Coenzyme A oxidase 3, pristanoyl | 4732 | 0.027 | 0.4230 | No |
| 27 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 4991 | 0.024 | 0.4107 | No |
| 28 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 5845 | 0.014 | 0.3509 | No |
| 29 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 6549 | 0.007 | 0.3001 | No |
| 30 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6985 | 0.002 | 0.2680 | No |
| 31 | ADIPOR2 | ADIPOR2 Entrez, Source | adiponectin receptor 2 | 7084 | 0.001 | 0.2610 | No |
| 32 | ACOT12 | ACOT12 Entrez, Source | acyl-CoA thioesterase 12 | 7714 | -0.006 | 0.2154 | No |
| 33 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 7949 | -0.009 | 0.2005 | No |
| 34 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 8150 | -0.011 | 0.1888 | No |
| 35 | CPT1B | CPT1B Entrez, Source | carnitine palmitoyltransferase 1B (muscle) | 9047 | -0.023 | 0.1281 | No |
| 36 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 9958 | -0.036 | 0.0703 | No |
| 37 | ABCD2 | ABCD2 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 2 | 10245 | -0.041 | 0.0608 | No |
| 38 | ADIPOR1 | ADIPOR1 Entrez, Source | adiponectin receptor 1 | 10701 | -0.049 | 0.0412 | No |
| 39 | MIF | MIF Entrez, Source | macrophage migration inhibitory factor (glycosylation-inhibiting factor) | 11278 | -0.062 | 0.0162 | No |
| 40 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 11526 | -0.069 | 0.0182 | No |
| 41 | LYPLA3 | LYPLA3 Entrez, Source | lysophospholipase 3 (lysosomal phospholipase A2) | 12539 | -0.121 | -0.0221 | No |
| 42 | CROT | CROT Entrez, Source | carnitine O-octanoyltransferase | 12676 | -0.132 | 0.0067 | No |
| 43 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 12787 | -0.146 | 0.0417 | No |

