Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | FETAL_LIVER_VS_ADULT_LIVER_GNF2 |
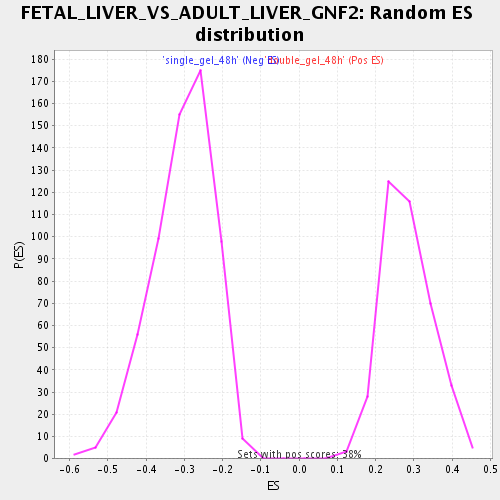
| Enrichment Score (ES) | 0.5888807 |
| Normalized Enrichment Score (NES) | 2.0843208 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0024673452 |
| FWER p-Value | 0.065 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 12 | 0.395 | 0.0936 | Yes |
| 2 | AQP9 | AQP9 Entrez, Source | aquaporin 9 | 57 | 0.272 | 0.1553 | Yes |
| 3 | PKLR | PKLR Entrez, Source | pyruvate kinase, liver and RBC | 74 | 0.247 | 0.2132 | Yes |
| 4 | CES1 | CES1 Entrez, Source | carboxylesterase 1 (monocyte/macrophage serine esterase 1) | 93 | 0.234 | 0.2676 | Yes |
| 5 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 148 | 0.193 | 0.3097 | Yes |
| 6 | UPB1 | UPB1 Entrez, Source | ureidopropionase, beta | 189 | 0.175 | 0.3486 | Yes |
| 7 | CD14 | CD14 Entrez, Source | CD14 molecule | 253 | 0.159 | 0.3818 | Yes |
| 8 | ASL | ASL Entrez, Source | argininosuccinate lyase | 273 | 0.154 | 0.4173 | Yes |
| 9 | HSD11B1 | HSD11B1 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 1 | 318 | 0.147 | 0.4491 | Yes |
| 10 | AZGP1 | AZGP1 Entrez, Source | alpha-2-glycoprotein 1, zinc | 392 | 0.137 | 0.4763 | Yes |
| 11 | ITIH4 | ITIH4 Entrez, Source | inter-alpha (globulin) inhibitor H4 (plasma Kallikrein-sensitive glycoprotein) | 429 | 0.132 | 0.5052 | Yes |
| 12 | IGFALS | IGFALS Entrez, Source | insulin-like growth factor binding protein, acid labile subunit | 500 | 0.125 | 0.5298 | Yes |
| 13 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 555 | 0.121 | 0.5547 | Yes |
| 14 | LGALS1 | LGALS1 Entrez, Source | lectin, galactoside-binding, soluble, 1 (galectin 1) | 749 | 0.109 | 0.5663 | Yes |
| 15 | SLC25A10 | SLC25A10 Entrez, Source | solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 | 790 | 0.107 | 0.5889 | Yes |
| 16 | PPP1R1A | PPP1R1A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 1A | 1155 | 0.091 | 0.5833 | No |
| 17 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 1528 | 0.079 | 0.5741 | No |
| 18 | CYP2C9 | CYP2C9 Entrez, Source | cytochrome P450, family 2, subfamily C, polypeptide 9 | 1777 | 0.073 | 0.5728 | No |
| 19 | ABCC3 | ABCC3 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 3 | 2034 | 0.067 | 0.5696 | No |
| 20 | SIGIRR | SIGIRR Entrez, Source | single immunoglobulin and toll-interleukin 1 receptor (TIR) domain | 2526 | 0.058 | 0.5465 | No |
| 21 | ACY1 | ACY1 Entrez, Source | aminoacylase 1 | 2722 | 0.055 | 0.5450 | No |
| 22 | COMT | COMT Entrez, Source | catechol-O-methyltransferase | 2832 | 0.053 | 0.5494 | No |
| 23 | ENDOG | ENDOG Entrez, Source | endonuclease G | 3299 | 0.046 | 0.5252 | No |
| 24 | SDS | SDS Entrez, Source | serine dehydratase | 3684 | 0.041 | 0.5061 | No |
| 25 | SHMT2 | SHMT2 Entrez, Source | serine hydroxymethyltransferase 2 (mitochondrial) | 3728 | 0.040 | 0.5124 | No |
| 26 | CYP1A2 | CYP1A2 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 2 | 3939 | 0.037 | 0.5055 | No |
| 27 | RARRES2 | RARRES2 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 2 | 4262 | 0.033 | 0.4892 | No |
| 28 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 4345 | 0.032 | 0.4907 | No |
| 29 | HAMP | HAMP Entrez, Source | hepcidin antimicrobial peptide | 4396 | 0.031 | 0.4944 | No |
| 30 | SLC22A1 | SLC22A1 Entrez, Source | solute carrier family 22 (organic cation transporter), member 1 | 5519 | 0.018 | 0.4144 | No |
| 31 | UQCRC1 | UQCRC1 Entrez, Source | ubiquinol-cytochrome c reductase core protein I | 6394 | 0.009 | 0.3506 | No |
| 32 | CES2 | CES2 Entrez, Source | carboxylesterase 2 (intestine, liver) | 6501 | 0.008 | 0.3445 | No |
| 33 | CYP27A1 | CYP27A1 Entrez, Source | cytochrome P450, family 27, subfamily A, polypeptide 1 | 6729 | 0.005 | 0.3286 | No |
| 34 | HGFAC | HGFAC Entrez, Source | HGF activator | 6825 | 0.004 | 0.3224 | No |
| 35 | RAB26 | RAB26 Entrez, Source | RAB26, member RAS oncogene family | 6898 | 0.003 | 0.3177 | No |
| 36 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6985 | 0.002 | 0.3117 | No |
| 37 | MUCDHL | MUCDHL Entrez, Source | mucin and cadherin-like | 7100 | 0.001 | 0.3034 | No |
| 38 | APCS | APCS Entrez, Source | amyloid P component, serum | 7122 | 0.001 | 0.3020 | No |
| 39 | NKG7 | NKG7 Entrez, Source | natural killer cell group 7 sequence | 7263 | -0.001 | 0.2916 | No |
| 40 | HPD | HPD Entrez, Source | 4-hydroxyphenylpyruvate dioxygenase | 8248 | -0.012 | 0.2206 | No |
| 41 | DCPS | DCPS Entrez, Source | decapping enzyme, scavenger | 8440 | -0.015 | 0.2097 | No |
| 42 | TXNRD2 | TXNRD2 Entrez, Source | thioredoxin reductase 2 | 8441 | -0.015 | 0.2133 | No |
| 43 | GSTM1 | GSTM1 Entrez, Source | glutathione S-transferase M1 | 8992 | -0.022 | 0.1771 | No |
| 44 | NRTN | NRTN Entrez, Source | neurturin | 9082 | -0.023 | 0.1759 | No |
| 45 | STAP2 | STAP2 Entrez, Source | - | 10223 | -0.040 | 0.0998 | No |
| 46 | CYP2E1 | CYP2E1 Entrez, Source | cytochrome P450, family 2, subfamily E, polypeptide 1 | 10268 | -0.041 | 0.1062 | No |
| 47 | CD81 | CD81 Entrez, Source | CD81 molecule | 11591 | -0.072 | 0.0239 | No |
| 48 | C1R | C1R Entrez, Source | complement component 1, r subcomponent | 11719 | -0.076 | 0.0324 | No |
| 49 | GPR30 | GPR30 Entrez, Source | G protein-coupled receptor 30 | 11742 | -0.077 | 0.0492 | No |
| 50 | CD151 | CD151 Entrez, Source | CD151 molecule (Raph blood group) | 12694 | -0.134 | 0.0098 | No |
| 51 | GSTM2 | GSTM2 Entrez, Source | glutathione S-transferase M2 (muscle) | 12890 | -0.163 | 0.0340 | No |