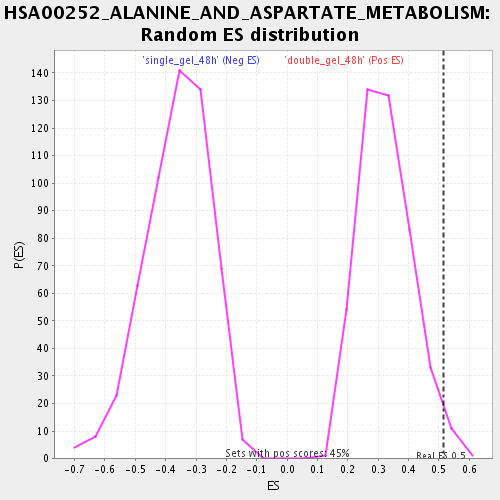
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | HSA00252_ALANINE_AND_ASPARTATE_METABOLISM |
| Enrichment Score (ES) | 0.5163962 |
| Normalized Enrichment Score (NES) | 1.5893043 |
| Nominal p-value | 0.022271715 |
| FDR q-value | 0.09755156 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 197 | 0.173 | 0.0827 | Yes |
| 2 | PC | PC Entrez, Source | pyruvate carboxylase | 270 | 0.155 | 0.1642 | Yes |
| 3 | ABAT | ABAT Entrez, Source | 4-aminobutyrate aminotransferase | 271 | 0.155 | 0.2510 | Yes |
| 4 | ASL | ASL Entrez, Source | argininosuccinate lyase | 273 | 0.154 | 0.3376 | Yes |
| 5 | PDHA2 | PDHA2 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 2 | 332 | 0.144 | 0.4144 | Yes |
| 6 | AGXT | AGXT Entrez, Source | alanine-glyoxylate aminotransferase (oxalosis I; hyperoxaluria I; glycolicaciduria; serine-pyruvate aminotransferase) | 591 | 0.118 | 0.4616 | Yes |
| 7 | PDHB | PDHB Entrez, Source | pyruvate dehydrogenase (lipoamide) beta | 1159 | 0.091 | 0.4701 | Yes |
| 8 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 1212 | 0.089 | 0.5164 | Yes |
| 9 | ASRGL1 | ASRGL1 Entrez, Source | asparaginase like 1 | 2861 | 0.052 | 0.4220 | No |
| 10 | ASS1 | ASS1 Entrez, Source | argininosuccinate synthetase 1 | 3063 | 0.049 | 0.4346 | No |
| 11 | AARS | AARS Entrez, Source | alanyl-tRNA synthetase | 3123 | 0.048 | 0.4573 | No |
| 12 | AGXT2 | AGXT2 Entrez, Source | alanine-glyoxylate aminotransferase 2 | 4019 | 0.036 | 0.4104 | No |
| 13 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 4645 | 0.028 | 0.3793 | No |
| 14 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 5455 | 0.019 | 0.3291 | No |
| 15 | NARS | NARS Entrez, Source | asparaginyl-tRNA synthetase | 5787 | 0.015 | 0.3127 | No |
| 16 | DLD | DLD Entrez, Source | dihydrolipoamide dehydrogenase | 5876 | 0.014 | 0.3140 | No |
| 17 | DARS | DARS Entrez, Source | aspartyl-tRNA synthetase | 6355 | 0.009 | 0.2832 | No |
| 18 | PDHA1 | PDHA1 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 1 | 6772 | 0.004 | 0.2545 | No |
| 19 | GOT2 | GOT2 Entrez, Source | glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) | 6812 | 0.004 | 0.2538 | No |
| 20 | DARS2 | DARS2 Entrez, Source | aspartyl-tRNA synthetase 2 (mitochondrial) | 7317 | -0.002 | 0.2169 | No |
| 21 | ASNS | ASNS Entrez, Source | asparagine synthetase | 7614 | -0.005 | 0.1975 | No |
| 22 | ACY3 | ACY3 Entrez, Source | aspartoacylase (aminocyclase) 3 | 11228 | -0.061 | -0.0397 | No |
| 23 | CRAT | CRAT Entrez, Source | carnitine acetyltransferase | 11766 | -0.078 | -0.0363 | No |
| 24 | DLAT | DLAT Entrez, Source | dihydrolipoamide S-acetyltransferase (E2 component of pyruvate dehydrogenase complex) | 12499 | -0.117 | -0.0254 | No |
| 25 | GPT | GPT Entrez, Source | glutamic-pyruvate transaminase (alanine aminotransferase) | 12858 | -0.158 | 0.0363 | No |