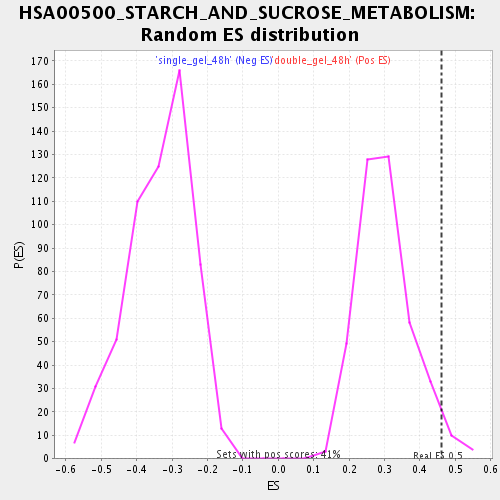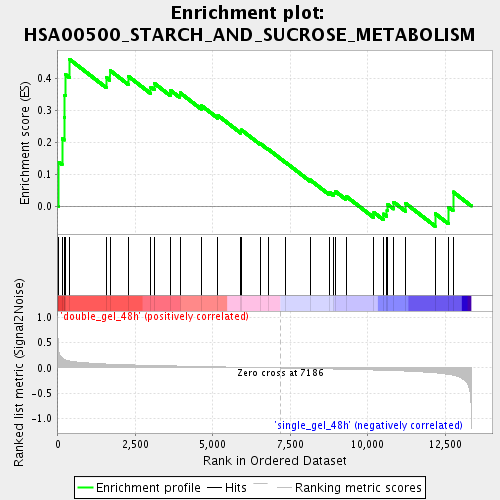
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | HSA00500_STARCH_AND_SUCROSE_METABOLISM |
| Enrichment Score (ES) | 0.45968705 |
| Normalized Enrichment Score (NES) | 1.5186617 |
| Nominal p-value | 0.033816423 |
| FDR q-value | 0.13124748 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | G6PC | G6PC Entrez, Source | glucose-6-phosphatase, catalytic subunit | 26 | 0.337 | 0.1378 | Yes |
| 2 | SI | SI Entrez, Source | sucrase-isomaltase (alpha-glucosidase) | 137 | 0.197 | 0.2113 | Yes |
| 3 | UGT2A1 | UGT2A1 Entrez, Source | UDP glucuronosyltransferase 2 family, polypeptide A1 | 201 | 0.173 | 0.2781 | Yes |
| 4 | GCK | GCK Entrez, Source | glucokinase (hexokinase 4, maturity onset diabetes of the young 2) | 218 | 0.170 | 0.3471 | Yes |
| 5 | AMY1A | AMY1A Entrez, Source | amylase, alpha 1A; salivary | 246 | 0.160 | 0.4114 | Yes |
| 6 | GYS2 | GYS2 Entrez, Source | glycogen synthase 2 (liver) | 370 | 0.139 | 0.4597 | Yes |
| 7 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 1562 | 0.078 | 0.4025 | No |
| 8 | ASCC3L1 | ASCC3L1 Entrez, Source | activating signal cointegrator 1 complex subunit 3-like 1 | 1681 | 0.075 | 0.4246 | No |
| 9 | AMY2A | AMY2A Entrez, Source | amylase, alpha 2A; pancreatic | 2276 | 0.063 | 0.4059 | No |
| 10 | GBE1 | GBE1 Entrez, Source | glucan (1,4-alpha-), branching enzyme 1 (glycogen branching enzyme, Andersen disease, glycogen storage disease type IV) | 2997 | 0.050 | 0.3725 | No |
| 11 | DDX50 | DDX50 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 50 | 3115 | 0.049 | 0.3838 | No |
| 12 | PYGM | PYGM Entrez, Source | phosphorylase, glycogen; muscle (McArdle syndrome, glycogen storage disease type V) | 3629 | 0.041 | 0.3624 | No |
| 13 | DDX18 | DDX18 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 18 | 3940 | 0.037 | 0.3546 | No |
| 14 | DDX56 | DDX56 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 56 | 4627 | 0.028 | 0.3148 | No |
| 15 | PYGL | PYGL Entrez, Source | phosphorylase, glycogen; liver (Hers disease, glycogen storage disease type VI) | 5160 | 0.022 | 0.2839 | No |
| 16 | HK1 | HK1 Entrez, Source | hexokinase 1 | 5904 | 0.014 | 0.2338 | No |
| 17 | TREH | TREH Entrez, Source | trehalase (brush-border membrane glycoprotein) | 5920 | 0.014 | 0.2383 | No |
| 18 | EP400 | EP400 Entrez, Source | E1A binding protein p400 | 6530 | 0.007 | 0.1955 | No |
| 19 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 6799 | 0.004 | 0.1771 | No |
| 20 | PGM1 | PGM1 Entrez, Source | phosphoglucomutase 1 | 7335 | -0.002 | 0.1377 | No |
| 21 | DDX47 | DDX47 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 47 | 8139 | -0.011 | 0.0819 | No |
| 22 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 8749 | -0.019 | 0.0438 | No |
| 23 | NUDT5 | NUDT5 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 5 | 8908 | -0.021 | 0.0405 | No |
| 24 | UGP2 | UGP2 Entrez, Source | UDP-glucose pyrophosphorylase 2 | 8944 | -0.021 | 0.0467 | No |
| 25 | ENPP1 | ENPP1 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 1 | 9297 | -0.026 | 0.0310 | No |
| 26 | GPI | GPI Entrez, Source | glucose phosphate isomerase | 10196 | -0.040 | -0.0200 | No |
| 27 | DDX52 | DDX52 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 52 | 10512 | -0.046 | -0.0248 | No |
| 28 | PYGB | PYGB Entrez, Source | phosphorylase, glycogen; brain | 10620 | -0.047 | -0.0132 | No |
| 29 | HK2 | HK2 Entrez, Source | hexokinase 2 | 10644 | -0.048 | 0.0049 | No |
| 30 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 10834 | -0.052 | 0.0123 | No |
| 31 | GAA | GAA Entrez, Source | glucosidase, alpha; acid (Pompe disease, glycogen storage disease type II) | 11222 | -0.061 | 0.0084 | No |
| 32 | GUSB | GUSB Entrez, Source | glucuronidase, beta | 12176 | -0.096 | -0.0236 | No |
| 33 | UXS1 | UXS1 Entrez, Source | UDP-glucuronate decarboxylase 1 | 12611 | -0.126 | -0.0040 | No |
| 34 | ENPP3 | ENPP3 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 3 | 12755 | -0.142 | 0.0441 | No |