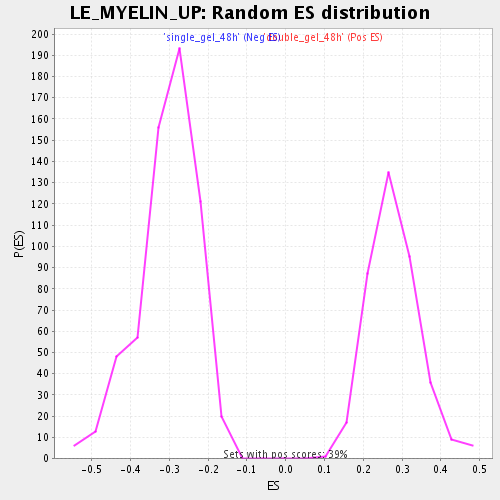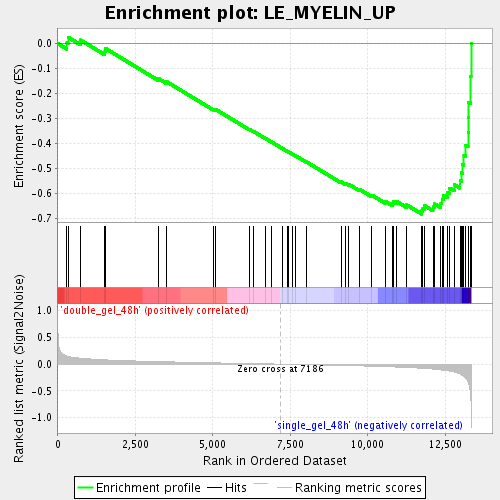
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | single_gel_48h |
| GeneSet | LE_MYELIN_UP |
| Enrichment Score (ES) | -0.68276674 |
| Normalized Enrichment Score (NES) | -2.2569125 |
| Nominal p-value | 0.0 |
| FDR q-value | 8.637701E-4 |
| FWER p-Value | 0.0030 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | POU3F1 | POU3F1 Entrez, Source | POU domain, class 3, transcription factor 1 | 291 | 0.150 | 0.0037 | No |
| 2 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 329 | 0.145 | 0.0255 | No |
| 3 | FIGNL1 | FIGNL1 Entrez, Source | fidgetin-like 1 | 723 | 0.110 | 0.0147 | No |
| 4 | EPHA5 | EPHA5 Entrez, Source | EPH receptor A5 | 1517 | 0.079 | -0.0315 | No |
| 5 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 1533 | 0.079 | -0.0193 | No |
| 6 | FOXD3 | FOXD3 Entrez, Source | forkhead box D3 | 3243 | 0.047 | -0.1399 | No |
| 7 | MCAM | MCAM Entrez, Source | melanoma cell adhesion molecule | 3499 | 0.043 | -0.1518 | No |
| 8 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 5020 | 0.024 | -0.2621 | No |
| 9 | SH3GL3 | SH3GL3 Entrez, Source | SH3-domain GRB2-like 3 | 5088 | 0.023 | -0.2633 | No |
| 10 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 6186 | 0.011 | -0.3439 | No |
| 11 | BZW2 | BZW2 Entrez, Source | basic leucine zipper and W2 domains 2 | 6314 | 0.009 | -0.3519 | No |
| 12 | PDE8A | PDE8A Entrez, Source | phosphodiesterase 8A | 6686 | 0.006 | -0.3788 | No |
| 13 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 6894 | 0.003 | -0.3939 | No |
| 14 | EDNRB | EDNRB Entrez, Source | endothelin receptor type B | 7246 | -0.001 | -0.4202 | No |
| 15 | P2RX4 | P2RX4 Entrez, Source | purinergic receptor P2X, ligand-gated ion channel, 4 | 7423 | -0.003 | -0.4329 | No |
| 16 | CXCL14 | CXCL14 Entrez, Source | chemokine (C-X-C motif) ligand 14 | 7432 | -0.003 | -0.4330 | No |
| 17 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 7563 | -0.004 | -0.4420 | No |
| 18 | RPL13A | RPL13A Entrez, Source | ribosomal protein L13a | 7679 | -0.006 | -0.4497 | No |
| 19 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 8020 | -0.010 | -0.4736 | No |
| 20 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 9136 | -0.024 | -0.5535 | No |
| 21 | NT5DC2 | NT5DC2 Entrez, Source | 5'-nucleotidase domain containing 2 | 9286 | -0.026 | -0.5604 | No |
| 22 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 9384 | -0.027 | -0.5630 | No |
| 23 | AURKA | AURKA Entrez, Source | aurora kinase A | 9722 | -0.032 | -0.5829 | No |
| 24 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 10131 | -0.039 | -0.6069 | No |
| 25 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 10564 | -0.046 | -0.6316 | No |
| 26 | PPP1R14B | PPP1R14B Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 14B | 10808 | -0.052 | -0.6410 | No |
| 27 | COL18A1 | COL18A1 Entrez, Source | collagen, type XVIII, alpha 1 | 10828 | -0.052 | -0.6336 | No |
| 28 | SPRED2 | SPRED2 Entrez, Source | sprouty-related, EVH1 domain containing 2 | 10933 | -0.054 | -0.6322 | No |
| 29 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 11253 | -0.061 | -0.6458 | No |
| 30 | TUBB6 | TUBB6 Entrez, Source | tubulin, beta 6 | 11746 | -0.077 | -0.6697 | Yes |
| 31 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 11780 | -0.078 | -0.6589 | Yes |
| 32 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 11841 | -0.081 | -0.6496 | Yes |
| 33 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 12106 | -0.092 | -0.6539 | Yes |
| 34 | PSPC1 | PSPC1 Entrez, Source | paraspeckle component 1 | 12162 | -0.095 | -0.6420 | Yes |
| 35 | KIF22 | KIF22 Entrez, Source | kinesin family member 22 | 12361 | -0.108 | -0.6385 | Yes |
| 36 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 12396 | -0.111 | -0.6223 | Yes |
| 37 | TCF19 | TCF19 Entrez, Source | transcription factor 19 (SC1) | 12445 | -0.114 | -0.6065 | Yes |
| 38 | HN1 | HN1 Entrez, Source | hematological and neurological expressed 1 | 12571 | -0.123 | -0.5951 | Yes |
| 39 | CDCA8 | CDCA8 Entrez, Source | cell division cycle associated 8 | 12643 | -0.129 | -0.5784 | Yes |
| 40 | CSRP2 | CSRP2 Entrez, Source | cysteine and glycine-rich protein 2 | 12792 | -0.147 | -0.5646 | Yes |
| 41 | CD44 | CD44 Entrez, Source | CD44 molecule (Indian blood group) | 12977 | -0.184 | -0.5472 | Yes |
| 42 | VIL2 | VIL2 Entrez, Source | villin 2 (ezrin) | 13025 | -0.196 | -0.5174 | Yes |
| 43 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 13063 | -0.214 | -0.4838 | Yes |
| 44 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 13104 | -0.234 | -0.4470 | Yes |
| 45 | BASP1 | BASP1 Entrez, Source | brain abundant, membrane attached signal protein 1 | 13161 | -0.265 | -0.4061 | Yes |
| 46 | CDC2A | 13238 | -0.335 | -0.3548 | Yes | ||
| 47 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 13250 | -0.352 | -0.2959 | Yes |
| 48 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13257 | -0.367 | -0.2340 | Yes |
| 49 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 13326 | -0.627 | -0.1325 | Yes |
| 50 | MARCKSL1 | MARCKSL1 Entrez, Source | MARCKS-like 1 | 13335 | -0.786 | 0.0005 | Yes |