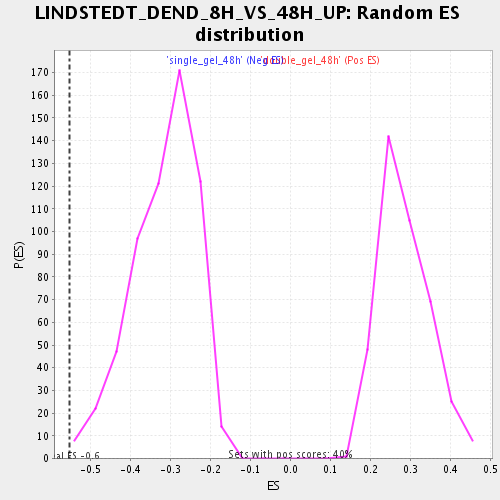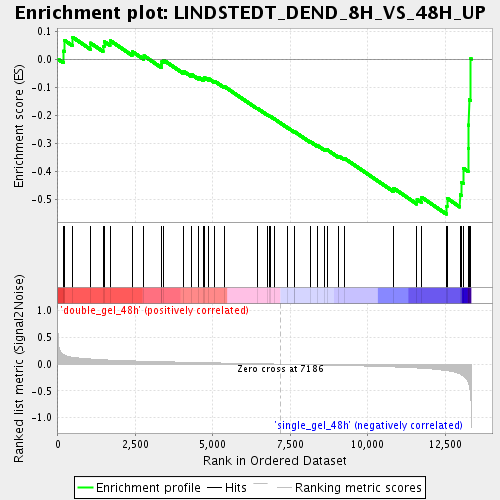
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | single_gel_48h |
| GeneSet | LINDSTEDT_DEND_8H_VS_48H_UP |
| Enrichment Score (ES) | -0.5527283 |
| Normalized Enrichment Score (NES) | -1.7568067 |
| Nominal p-value | 0.003322259 |
| FDR q-value | 0.04938339 |
| FWER p-Value | 0.997 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RTN1 | RTN1 Entrez, Source | reticulon 1 | 181 | 0.178 | 0.0291 | No |
| 2 | FKBP5 | FKBP5 Entrez, Source | FK506 binding protein 5 | 219 | 0.169 | 0.0667 | No |
| 3 | IL1RN | IL1RN Entrez, Source | interleukin 1 receptor antagonist | 475 | 0.128 | 0.0781 | No |
| 4 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 1050 | 0.095 | 0.0577 | No |
| 5 | ATF5 | ATF5 Entrez, Source | activating transcription factor 5 | 1459 | 0.081 | 0.0464 | No |
| 6 | IL1B | IL1B Entrez, Source | interleukin 1, beta | 1495 | 0.079 | 0.0628 | No |
| 7 | TNFSF9 | TNFSF9 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 9 | 1683 | 0.075 | 0.0666 | No |
| 8 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 2396 | 0.060 | 0.0275 | No |
| 9 | SLC3A2 | SLC3A2 Entrez, Source | solute carrier family 3 (activators of dibasic and neutral amino acid transport), member 2 | 2777 | 0.054 | 0.0118 | No |
| 10 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 3342 | 0.045 | -0.0198 | No |
| 11 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 3354 | 0.045 | -0.0099 | No |
| 12 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 3406 | 0.044 | -0.0031 | No |
| 13 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 4061 | 0.035 | -0.0438 | No |
| 14 | PLA2G4A | PLA2G4A Entrez, Source | phospholipase A2, group IVA (cytosolic, calcium-dependent) | 4307 | 0.033 | -0.0544 | No |
| 15 | RAB5A | RAB5A Entrez, Source | RAB5A, member RAS oncogene family | 4533 | 0.030 | -0.0642 | No |
| 16 | CCL3 | CCL3 Entrez, Source | chemokine (C-C motif) ligand 3 | 4706 | 0.027 | -0.0706 | No |
| 17 | IFIT2 | IFIT2 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 2 | 4735 | 0.027 | -0.0662 | No |
| 18 | ENTPD1 | ENTPD1 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 1 | 4858 | 0.026 | -0.0692 | No |
| 19 | STAT4 | STAT4 Entrez, Source | signal transducer and activator of transcription 4 | 5065 | 0.023 | -0.0792 | No |
| 20 | IL12B | IL12B Entrez, Source | interleukin 12B (natural killer cell stimulatory factor 2, cytotoxic lymphocyte maturation factor 2, p40) | 5367 | 0.020 | -0.0971 | No |
| 21 | PLEK | PLEK Entrez, Source | pleckstrin | 6431 | 0.008 | -0.1751 | No |
| 22 | TNF | TNF Entrez, Source | tumor necrosis factor (TNF superfamily, member 2) | 6778 | 0.004 | -0.2001 | No |
| 23 | TNFRSF4 | TNFRSF4 Entrez, Source | tumor necrosis factor receptor superfamily, member 4 | 6831 | 0.004 | -0.2031 | No |
| 24 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 6864 | 0.004 | -0.2046 | No |
| 25 | TLR2 | TLR2 Entrez, Source | toll-like receptor 2 | 6981 | 0.002 | -0.2128 | No |
| 26 | P2RX4 | P2RX4 Entrez, Source | purinergic receptor P2X, ligand-gated ion channel, 4 | 7423 | -0.003 | -0.2453 | No |
| 27 | CCL4 | CCL4 Entrez, Source | chemokine (C-C motif) ligand 4 | 7645 | -0.005 | -0.2607 | No |
| 28 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 7648 | -0.005 | -0.2596 | No |
| 29 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 8149 | -0.011 | -0.2945 | No |
| 30 | TFEC | TFEC Entrez, Source | transcription factor EC | 8378 | -0.014 | -0.3083 | No |
| 31 | ACVR2A | ACVR2A Entrez, Source | activin A receptor, type IIA | 8618 | -0.017 | -0.3222 | No |
| 32 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 8686 | -0.018 | -0.3230 | No |
| 33 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 9067 | -0.023 | -0.3461 | No |
| 34 | CSF2 | CSF2 Entrez, Source | colony stimulating factor 2 (granulocyte-macrophage) | 9237 | -0.025 | -0.3529 | No |
| 35 | CD48 | CD48 Entrez, Source | CD48 molecule | 10831 | -0.052 | -0.4602 | No |
| 36 | IL7 | IL7 Entrez, Source | interleukin 7 | 11588 | -0.071 | -0.4999 | No |
| 37 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 11740 | -0.077 | -0.4929 | No |
| 38 | PTGER4 | PTGER4 Entrez, Source | prostaglandin E receptor 4 (subtype EP4) | 12537 | -0.120 | -0.5239 | Yes |
| 39 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 12561 | -0.122 | -0.4964 | Yes |
| 40 | CD44 | CD44 Entrez, Source | CD44 molecule (Indian blood group) | 12977 | -0.184 | -0.4836 | Yes |
| 41 | EDN1 | EDN1 Entrez, Source | endothelin 1 | 13031 | -0.199 | -0.4399 | Yes |
| 42 | CXCL11 | CXCL11 Entrez, Source | chemokine (C-X-C motif) ligand 11 | 13092 | -0.227 | -0.3900 | Yes |
| 43 | PLAT | PLAT Entrez, Source | plasminogen activator, tissue | 13241 | -0.343 | -0.3189 | Yes |
| 44 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 13254 | -0.355 | -0.2348 | Yes |
| 45 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 13267 | -0.380 | -0.1446 | Yes |
| 46 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 13326 | -0.627 | 0.0012 | Yes |