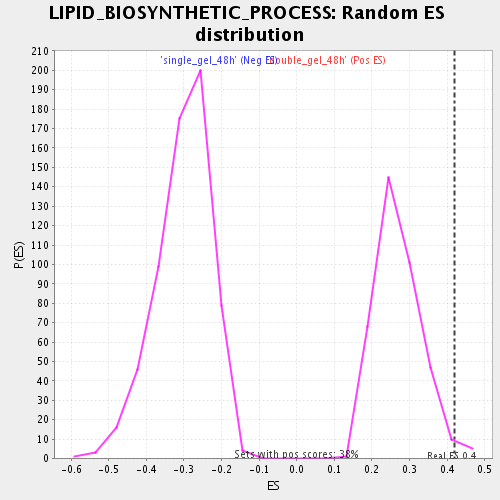
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | LIPID_BIOSYNTHETIC_PROCESS |
| Enrichment Score (ES) | 0.42045504 |
| Normalized Enrichment Score (NES) | 1.5602778 |
| Nominal p-value | 0.018567638 |
| FDR q-value | 0.1163124 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PEMT | PEMT Entrez, Source | phosphatidylethanolamine N-methyltransferase | 4 | 0.491 | 0.1032 | Yes |
| 2 | IDI1 | IDI1 Entrez, Source | isopentenyl-diphosphate delta isomerase 1 | 13 | 0.382 | 0.1831 | Yes |
| 3 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 167 | 0.185 | 0.2105 | Yes |
| 4 | FDPS | FDPS Entrez, Source | farnesyl diphosphate synthase (farnesyl pyrophosphate synthetase, dimethylallyltranstransferase, geranyltranstransferase) | 171 | 0.184 | 0.2490 | Yes |
| 5 | MVK | MVK Entrez, Source | mevalonate kinase (mevalonic aciduria) | 195 | 0.174 | 0.2839 | Yes |
| 6 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 247 | 0.160 | 0.3138 | Yes |
| 7 | LCAT | LCAT Entrez, Source | lecithin-cholesterol acyltransferase | 259 | 0.157 | 0.3461 | Yes |
| 8 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 362 | 0.140 | 0.3679 | Yes |
| 9 | CYP19A1 | CYP19A1 Entrez, Source | cytochrome P450, family 19, subfamily A, polypeptide 1 | 670 | 0.114 | 0.3688 | Yes |
| 10 | CHPT1 | CHPT1 Entrez, Source | choline phosphotransferase 1 | 789 | 0.107 | 0.3825 | Yes |
| 11 | PCYT2 | PCYT2 Entrez, Source | phosphate cytidylyltransferase 2, ethanolamine | 944 | 0.100 | 0.3919 | Yes |
| 12 | NR0B1 | NR0B1 Entrez, Source | nuclear receptor subfamily 0, group B, member 1 | 1027 | 0.096 | 0.4060 | Yes |
| 13 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 1097 | 0.093 | 0.4205 | Yes |
| 14 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 1768 | 0.073 | 0.3854 | No |
| 15 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 1932 | 0.070 | 0.3877 | No |
| 16 | AKR1D1 | AKR1D1 Entrez, Source | aldo-keto reductase family 1, member D1 (delta 4-3-ketosteroid-5-beta-reductase) | 2098 | 0.066 | 0.3892 | No |
| 17 | BMP6 | BMP6 Entrez, Source | bone morphogenetic protein 6 | 2251 | 0.063 | 0.3911 | No |
| 18 | ST6GALNAC6 | ST6GALNAC6 Entrez, Source | ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 6 | 2360 | 0.061 | 0.3957 | No |
| 19 | HSD3B1 | HSD3B1 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 | 2646 | 0.056 | 0.3861 | No |
| 20 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 3007 | 0.050 | 0.3695 | No |
| 21 | APOA1 | APOA1 Entrez, Source | apolipoprotein A-I | 3081 | 0.049 | 0.3744 | No |
| 22 | PI4K2A | 3202 | 0.047 | 0.3753 | No | ||
| 23 | SCP2 | SCP2 Entrez, Source | sterol carrier protein 2 | 3534 | 0.042 | 0.3593 | No |
| 24 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 3915 | 0.038 | 0.3386 | No |
| 25 | ST8SIA5 | ST8SIA5 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 5 | 4130 | 0.035 | 0.3298 | No |
| 26 | FDXR | FDXR Entrez, Source | ferredoxin reductase | 4149 | 0.034 | 0.3357 | No |
| 27 | DGAT2 | DGAT2 Entrez, Source | diacylglycerol O-acyltransferase homolog 2 (mouse) | 4256 | 0.033 | 0.3347 | No |
| 28 | ST8SIA1 | ST8SIA1 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 1 | 4799 | 0.026 | 0.2995 | No |
| 29 | CNBP | CNBP Entrez, Source | CCHC-type zinc finger, nucleic acid binding protein | 5336 | 0.020 | 0.2634 | No |
| 30 | CYP11A1 | CYP11A1 Entrez, Source | cytochrome P450, family 11, subfamily A, polypeptide 1 | 5484 | 0.019 | 0.2562 | No |
| 31 | PCYT1B | PCYT1B Entrez, Source | phosphate cytidylyltransferase 1, choline, beta | 5923 | 0.014 | 0.2261 | No |
| 32 | PIGV | PIGV Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class V | 5939 | 0.013 | 0.2278 | No |
| 33 | HSD11B2 | HSD11B2 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 2 | 5960 | 0.013 | 0.2291 | No |
| 34 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6985 | 0.002 | 0.1524 | No |
| 35 | PEX7 | PEX7 Entrez, Source | peroxisomal biogenesis factor 7 | 7894 | -0.008 | 0.0858 | No |
| 36 | AGPAT1 | AGPAT1 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) | 7919 | -0.009 | 0.0858 | No |
| 37 | AGPS | AGPS Entrez, Source | alkylglycerone phosphate synthase | 8182 | -0.012 | 0.0685 | No |
| 38 | PIGK | PIGK Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class K | 8287 | -0.013 | 0.0634 | No |
| 39 | HSD3B7 | HSD3B7 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 | 8545 | -0.016 | 0.0474 | No |
| 40 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 9099 | -0.023 | 0.0107 | No |
| 41 | IMPA1 | IMPA1 Entrez, Source | inositol(myo)-1(or 4)-monophosphatase 1 | 9219 | -0.025 | 0.0069 | No |
| 42 | CYP7B1 | CYP7B1 Entrez, Source | cytochrome P450, family 7, subfamily B, polypeptide 1 | 9803 | -0.033 | -0.0299 | No |
| 43 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 9958 | -0.036 | -0.0339 | No |
| 44 | PIGC | PIGC Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class C | 10066 | -0.038 | -0.0340 | No |
| 45 | DPM2 | DPM2 Entrez, Source | dolichyl-phosphate mannosyltransferase polypeptide 2, regulatory subunit | 10215 | -0.040 | -0.0367 | No |
| 46 | PIGS | PIGS Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class S | 10690 | -0.049 | -0.0620 | No |
| 47 | ST8SIA3 | ST8SIA3 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 3 | 10869 | -0.053 | -0.0643 | No |
| 48 | MIF | MIF Entrez, Source | macrophage migration inhibitory factor (glycosylation-inhibiting factor) | 11278 | -0.062 | -0.0820 | No |
| 49 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 11526 | -0.069 | -0.0860 | No |
| 50 | CD81 | CD81 Entrez, Source | CD81 molecule | 11591 | -0.072 | -0.0757 | No |
| 51 | AGPAT6 | AGPAT6 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 6 (lysophosphatidic acid acyltransferase, zeta) | 12037 | -0.089 | -0.0905 | No |
| 52 | PLCE1 | PLCE1 Entrez, Source | phospholipase C, epsilon 1 | 12550 | -0.121 | -0.1035 | No |
| 53 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 12787 | -0.146 | -0.0905 | No |
| 54 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 13105 | -0.234 | -0.0651 | No |
| 55 | ADM | ADM Entrez, Source | adrenomedullin | 13274 | -0.393 | 0.0051 | No |