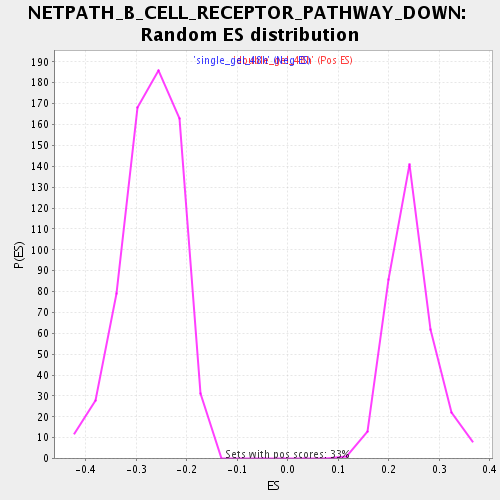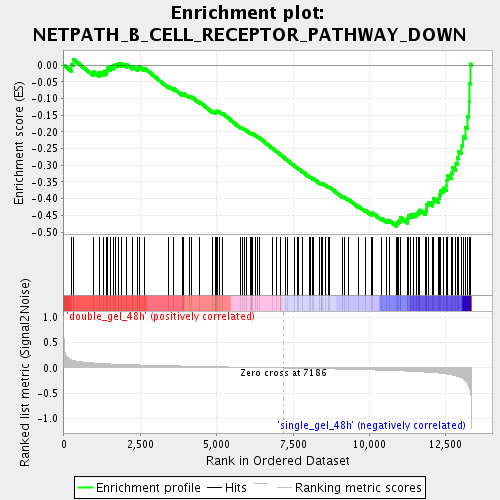
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | single_gel_48h |
| GeneSet | NETPATH_B_CELL_RECEPTOR_PATHWAY_DOWN |
| Enrichment Score (ES) | -0.4826321 |
| Normalized Enrichment Score (NES) | -1.7844107 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.04470987 |
| FWER p-Value | 0.985 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PTGER3 | PTGER3 Entrez, Source | prostaglandin E receptor 3 (subtype EP3) | 244 | 0.161 | 0.0028 | No |
| 2 | LTBP4 | LTBP4 Entrez, Source | latent transforming growth factor beta binding protein 4 | 305 | 0.149 | 0.0180 | No |
| 3 | ASGR1 | ASGR1 Entrez, Source | asialoglycoprotein receptor 1 | 970 | 0.099 | -0.0192 | No |
| 4 | PPP1R1A | PPP1R1A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 1A | 1155 | 0.091 | -0.0210 | No |
| 5 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 1285 | 0.086 | -0.0193 | No |
| 6 | CD79B | CD79B Entrez, Source | CD79b molecule, immunoglobulin-associated beta | 1390 | 0.083 | -0.0162 | No |
| 7 | ACVR2B | ACVR2B Entrez, Source | activin A receptor, type IIB | 1426 | 0.082 | -0.0080 | No |
| 8 | CSDE1 | CSDE1 Entrez, Source | cold shock domain containing E1, RNA-binding | 1522 | 0.079 | -0.0048 | No |
| 9 | PF4 | PF4 Entrez, Source | platelet factor 4 (chemokine (C-X-C motif) ligand 4) | 1615 | 0.077 | -0.0016 | No |
| 10 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 1682 | 0.075 | 0.0033 | No |
| 11 | CD40 | CD40 Entrez, Source | CD40 molecule, TNF receptor superfamily member 5 | 1790 | 0.072 | 0.0048 | No |
| 12 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 1896 | 0.070 | 0.0061 | No |
| 13 | RRAGB | RRAGB Entrez, Source | Ras-related GTP binding B | 2058 | 0.067 | 0.0028 | No |
| 14 | BMP6 | BMP6 Entrez, Source | bone morphogenetic protein 6 | 2251 | 0.063 | -0.0034 | No |
| 15 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 2418 | 0.060 | -0.0080 | No |
| 16 | IGJ | IGJ Entrez, Source | immunoglobulin J polypeptide, linker protein for immunoglobulin alpha and mu polypeptides | 2472 | 0.059 | -0.0042 | No |
| 17 | TIE1 | TIE1 Entrez, Source | tyrosine kinase with immunoglobulin-like and EGF-like domains 1 | 2639 | 0.056 | -0.0093 | No |
| 18 | TEC | TEC Entrez, Source | tec protein tyrosine kinase | 3424 | 0.044 | -0.0627 | No |
| 19 | SND1 | SND1 Entrez, Source | - | 3595 | 0.042 | -0.0700 | No |
| 20 | CUL5 | CUL5 Entrez, Source | cullin 5 | 3868 | 0.038 | -0.0855 | No |
| 21 | SRPK1 | SRPK1 Entrez, Source | SFRS protein kinase 1 | 3928 | 0.037 | -0.0850 | No |
| 22 | RIT2 | RIT2 Entrez, Source | Ras-like without CAAX 2 | 4102 | 0.035 | -0.0934 | No |
| 23 | PTPN2 | PTPN2 Entrez, Source | protein tyrosine phosphatase, non-receptor type 2 | 4170 | 0.034 | -0.0940 | No |
| 24 | P2RY6 | P2RY6 Entrez, Source | pyrimidinergic receptor P2Y, G-protein coupled, 6 | 4439 | 0.031 | -0.1102 | No |
| 25 | HINT1 | HINT1 Entrez, Source | histidine triad nucleotide binding protein 1 | 4853 | 0.026 | -0.1380 | No |
| 26 | NR3C2 | NR3C2 Entrez, Source | nuclear receptor subfamily 3, group C, member 2 | 4947 | 0.025 | -0.1418 | No |
| 27 | MAP3K10 | MAP3K10 Entrez, Source | mitogen-activated protein kinase kinase kinase 10 | 4975 | 0.024 | -0.1406 | No |
| 28 | HCLS1 | HCLS1 Entrez, Source | hematopoietic cell-specific Lyn substrate 1 | 4986 | 0.024 | -0.1382 | No |
| 29 | HTR2C | HTR2C Entrez, Source | 5-hydroxytryptamine (serotonin) receptor 2C | 5015 | 0.024 | -0.1372 | No |
| 30 | CDC2L5 | CDC2L5 Entrez, Source | cell division cycle 2-like 5 (cholinesterase-related cell division controller) | 5085 | 0.023 | -0.1394 | No |
| 31 | ATP5G3 | ATP5G3 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C3 (subunit 9) | 5189 | 0.022 | -0.1443 | No |
| 32 | GZMA | GZMA Entrez, Source | granzyme A (granzyme 1, cytotoxic T-lymphocyte-associated serine esterase 3) | 5775 | 0.015 | -0.1865 | No |
| 33 | PSMB5 | PSMB5 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 5 | 5837 | 0.015 | -0.1892 | No |
| 34 | SMARCD3 | SMARCD3 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3 | 5901 | 0.014 | -0.1921 | No |
| 35 | GSPT1 | GSPT1 Entrez, Source | G1 to S phase transition 1 | 5981 | 0.013 | -0.1964 | No |
| 36 | NME2 | NME2 Entrez, Source | non-metastatic cells 2, protein (NM23B) expressed in | 6108 | 0.012 | -0.2044 | No |
| 37 | IL13RA2 | IL13RA2 Entrez, Source | interleukin 13 receptor, alpha 2 | 6144 | 0.011 | -0.2055 | No |
| 38 | RAC2 | RAC2 Entrez, Source | ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) | 6150 | 0.011 | -0.2044 | No |
| 39 | CDK10 | CDK10 Entrez, Source | cyclin-dependent kinase (CDC2-like) 10 | 6168 | 0.011 | -0.2042 | No |
| 40 | TEGT | TEGT Entrez, Source | testis enhanced gene transcript (BAX inhibitor 1) | 6284 | 0.010 | -0.2115 | No |
| 41 | CCNC | CCNC Entrez, Source | cyclin C | 6339 | 0.009 | -0.2144 | No |
| 42 | RPS3 | RPS3 Entrez, Source | ribosomal protein S3 | 6390 | 0.009 | -0.2170 | No |
| 43 | DYRK3 | DYRK3 Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 3 | 6818 | 0.004 | -0.2488 | No |
| 44 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 6964 | 0.002 | -0.2594 | No |
| 45 | CSNK2B | CSNK2B Entrez, Source | casein kinase 2, beta polypeptide | 7077 | 0.001 | -0.2677 | No |
| 46 | FRZB | FRZB Entrez, Source | frizzled-related protein | 7085 | 0.001 | -0.2681 | No |
| 47 | GNAZ | GNAZ Entrez, Source | guanine nucleotide binding protein (G protein), alpha z polypeptide | 7248 | -0.001 | -0.2803 | No |
| 48 | PRKCA | PRKCA Entrez, Source | protein kinase C, alpha | 7310 | -0.002 | -0.2847 | No |
| 49 | SRP54 | SRP54 Entrez, Source | signal recognition particle 54kDa | 7534 | -0.004 | -0.3010 | No |
| 50 | PPP2CA | PPP2CA Entrez, Source | protein phosphatase 2 (formerly 2A), catalytic subunit, alpha isoform | 7638 | -0.005 | -0.3081 | No |
| 51 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 7690 | -0.006 | -0.3112 | No |
| 52 | STK38 | STK38 Entrez, Source | serine/threonine kinase 38 | 7820 | -0.007 | -0.3200 | No |
| 53 | CD3G | CD3G Entrez, Source | CD3g molecule, gamma (CD3-TCR complex) | 8029 | -0.010 | -0.3344 | No |
| 54 | SLA | SLA Entrez, Source | Src-like-adaptor | 8073 | -0.010 | -0.3363 | No |
| 55 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 8149 | -0.011 | -0.3405 | No |
| 56 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 8159 | -0.011 | -0.3397 | No |
| 57 | LTBP2 | LTBP2 Entrez, Source | latent transforming growth factor beta binding protein 2 | 8356 | -0.014 | -0.3527 | No |
| 58 | PSMC2 | PSMC2 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 2 | 8416 | -0.014 | -0.3552 | No |
| 59 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 8419 | -0.014 | -0.3534 | No |
| 60 | ST13 | ST13 Entrez, Source | suppression of tumorigenicity 13 (colon carcinoma) (Hsp70 interacting protein) | 8448 | -0.015 | -0.3536 | No |
| 61 | ARHGEF2 | ARHGEF2 Entrez, Source | rho/rac guanine nucleotide exchange factor (GEF) 2 | 8556 | -0.016 | -0.3595 | No |
| 62 | PPP2R1A | PPP2R1A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), alpha isoform | 8667 | -0.018 | -0.3655 | No |
| 63 | PLCG2 | PLCG2 Entrez, Source | phospholipase C, gamma 2 (phosphatidylinositol-specific) | 8698 | -0.018 | -0.3654 | No |
| 64 | CDK5R1 | CDK5R1 Entrez, Source | cyclin-dependent kinase 5, regulatory subunit 1 (p35) | 9127 | -0.023 | -0.3946 | No |
| 65 | RPS6KA1 | RPS6KA1 Entrez, Source | ribosomal protein S6 kinase, 90kDa, polypeptide 1 | 9175 | -0.024 | -0.3950 | No |
| 66 | PRDX2 | PRDX2 Entrez, Source | peroxiredoxin 2 | 9329 | -0.026 | -0.4031 | No |
| 67 | HDGF | HDGF Entrez, Source | hepatoma-derived growth factor (high-mobility group protein 1-like) | 9633 | -0.031 | -0.4219 | No |
| 68 | RAD23A | RAD23A Entrez, Source | RAD23 homolog A (S. cerevisiae) | 9855 | -0.034 | -0.4341 | No |
| 69 | SELE | SELE Entrez, Source | selectin E (endothelial adhesion molecule 1) | 10062 | -0.038 | -0.4446 | No |
| 70 | EI24 | EI24 Entrez, Source | etoposide induced 2.4 mRNA | 10107 | -0.038 | -0.4429 | No |
| 71 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 10388 | -0.043 | -0.4583 | No |
| 72 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 10564 | -0.046 | -0.4654 | No |
| 73 | PPP1CA | PPP1CA Entrez, Source | protein phosphatase 1, catalytic subunit, alpha isoform | 10652 | -0.048 | -0.4656 | No |
| 74 | PSME1 | PSME1 Entrez, Source | proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) | 10878 | -0.053 | -0.4756 | Yes |
| 75 | ERBB2 | ERBB2 Entrez, Source | v-erb-b2 erythroblastic leukemia viral oncogene homolog 2, neuro/glioblastoma derived oncogene homolog (avian) | 10906 | -0.054 | -0.4705 | Yes |
| 76 | CHN1 | CHN1 Entrez, Source | chimerin (chimaerin) 1 | 10953 | -0.055 | -0.4667 | Yes |
| 77 | YWHAH | YWHAH Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide | 11003 | -0.056 | -0.4630 | Yes |
| 78 | GNB1 | GNB1 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 1 | 11004 | -0.056 | -0.4556 | Yes |
| 79 | EIF2AK2 | EIF2AK2 Entrez, Source | eukaryotic translation initiation factor 2-alpha kinase 2 | 11242 | -0.061 | -0.4655 | Yes |
| 80 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 11253 | -0.061 | -0.4581 | Yes |
| 81 | RAB11A | RAB11A Entrez, Source | RAB11A, member RAS oncogene family | 11273 | -0.062 | -0.4514 | Yes |
| 82 | ESR1 | ESR1 Entrez, Source | estrogen receptor 1 | 11349 | -0.064 | -0.4486 | Yes |
| 83 | CASP7 | CASP7 Entrez, Source | caspase 7, apoptosis-related cysteine peptidase | 11446 | -0.066 | -0.4471 | Yes |
| 84 | IRF7 | IRF7 Entrez, Source | interferon regulatory factor 7 | 11527 | -0.069 | -0.4440 | Yes |
| 85 | NUCB1 | NUCB1 Entrez, Source | nucleobindin 1 | 11592 | -0.072 | -0.4393 | Yes |
| 86 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 11651 | -0.074 | -0.4339 | Yes |
| 87 | MAPRE1 | MAPRE1 Entrez, Source | microtubule-associated protein, RP/EB family, member 1 | 11845 | -0.081 | -0.4378 | Yes |
| 88 | LRP2 | LRP2 Entrez, Source | low density lipoprotein-related protein 2 | 11861 | -0.081 | -0.4282 | Yes |
| 89 | CD99 | CD99 Entrez, Source | CD99 molecule | 11871 | -0.082 | -0.4180 | Yes |
| 90 | HCFC1R1 | HCFC1R1 Entrez, Source | host cell factor C1 regulator 1 (XPO1 dependent) | 11920 | -0.084 | -0.4105 | Yes |
| 91 | CAMK1 | CAMK1 Entrez, Source | calcium/calmodulin-dependent protein kinase I | 12074 | -0.090 | -0.4102 | Yes |
| 92 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 12081 | -0.091 | -0.3986 | Yes |
| 93 | PRKCI | PRKCI Entrez, Source | protein kinase C, iota | 12255 | -0.100 | -0.3984 | Yes |
| 94 | IL1R2 | IL1R2 Entrez, Source | interleukin 1 receptor, type II | 12286 | -0.103 | -0.3871 | Yes |
| 95 | PTPRC | PTPRC Entrez, Source | protein tyrosine phosphatase, receptor type, C | 12324 | -0.105 | -0.3760 | Yes |
| 96 | YWHAQ | YWHAQ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta polypeptide | 12415 | -0.112 | -0.3680 | Yes |
| 97 | IRF1 | IRF1 Entrez, Source | interferon regulatory factor 1 | 12531 | -0.120 | -0.3609 | Yes |
| 98 | PRKCD | PRKCD Entrez, Source | protein kinase C, delta | 12535 | -0.120 | -0.3452 | Yes |
| 99 | LMNA | LMNA Entrez, Source | lamin A/C | 12554 | -0.122 | -0.3304 | Yes |
| 100 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 12691 | -0.134 | -0.3230 | Yes |
| 101 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 12716 | -0.137 | -0.3067 | Yes |
| 102 | PTPRZ1 | PTPRZ1 Entrez, Source | protein tyrosine phosphatase, receptor-type, Z polypeptide 1 | 12829 | -0.152 | -0.2950 | Yes |
| 103 | GRK5 | GRK5 Entrez, Source | G protein-coupled receptor kinase 5 | 12883 | -0.162 | -0.2776 | Yes |
| 104 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 12915 | -0.168 | -0.2577 | Yes |
| 105 | VIL2 | VIL2 Entrez, Source | villin 2 (ezrin) | 13025 | -0.196 | -0.2400 | Yes |
| 106 | PRKAR2B | PRKAR2B Entrez, Source | protein kinase, cAMP-dependent, regulatory, type II, beta | 13060 | -0.212 | -0.2145 | Yes |
| 107 | PDGFA | PDGFA Entrez, Source | platelet-derived growth factor alpha polypeptide | 13145 | -0.254 | -0.1872 | Yes |
| 108 | RAB31 | RAB31 Entrez, Source | RAB31, member RAS oncogene family | 13200 | -0.292 | -0.1526 | Yes |
| 109 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13257 | -0.367 | -0.1084 | Yes |
| 110 | PEA15 | PEA15 Entrez, Source | phosphoprotein enriched in astrocytes 15 | 13282 | -0.412 | -0.0556 | Yes |
| 111 | GBP2 | GBP2 Entrez, Source | guanylate binding protein 2, interferon-inducible | 13295 | -0.454 | 0.0036 | Yes |