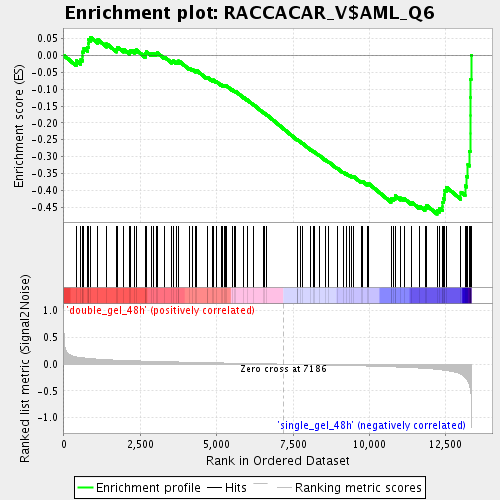
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | single_gel_48h |
| GeneSet | RACCACAR_V$AML_Q6 |
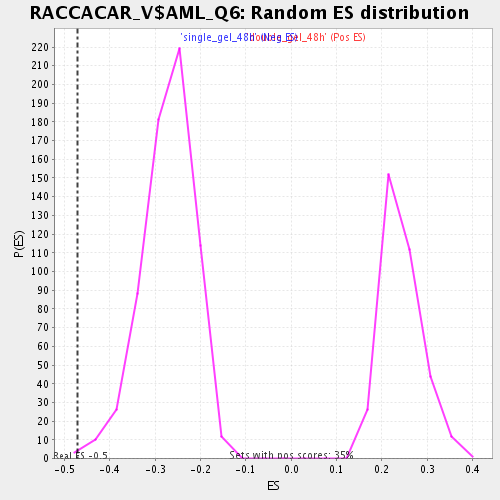
| Enrichment Score (ES) | -0.47063798 |
| Normalized Enrichment Score (NES) | -1.7296944 |
| Nominal p-value | 0.0030627872 |
| FDR q-value | 0.054521 |
| FWER p-Value | 0.999 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCR1 | CCR1 Entrez, Source | chemokine (C-C motif) receptor 1 | 410 | 0.134 | -0.0154 | No |
| 2 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 555 | 0.121 | -0.0122 | No |
| 3 | MATN1 | MATN1 Entrez, Source | matrilin 1, cartilage matrix protein | 614 | 0.117 | -0.0030 | No |
| 4 | SCNN1A | SCNN1A Entrez, Source | sodium channel, nonvoltage-gated 1 alpha | 616 | 0.117 | 0.0105 | No |
| 5 | CRY1 | CRY1 Entrez, Source | cryptochrome 1 (photolyase-like) | 652 | 0.115 | 0.0212 | No |
| 6 | CALR | CALR Entrez, Source | calreticulin | 768 | 0.108 | 0.0250 | No |
| 7 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 801 | 0.107 | 0.0350 | No |
| 8 | TRPV2 | TRPV2 Entrez, Source | transient receptor potential cation channel, subfamily V, member 2 | 808 | 0.106 | 0.0469 | No |
| 9 | FGFR1 | FGFR1 Entrez, Source | fibroblast growth factor receptor 1 (fms-related tyrosine kinase 2, Pfeiffer syndrome) | 873 | 0.104 | 0.0541 | No |
| 10 | MAFG | MAFG Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog G (avian) | 1111 | 0.093 | 0.0470 | No |
| 11 | MSRA | MSRA Entrez, Source | methionine sulfoxide reductase A | 1404 | 0.082 | 0.0345 | No |
| 12 | PCSK5 | PCSK5 Entrez, Source | proprotein convertase subtilisin/kexin type 5 | 1726 | 0.074 | 0.0188 | No |
| 13 | CDC37L1 | CDC37L1 Entrez, Source | CDC37 cell division cycle 37 homolog (S. cerevisiae)-like 1 | 1754 | 0.073 | 0.0252 | No |
| 14 | AKAP3 | AKAP3 Entrez, Source | A kinase (PRKA) anchor protein 3 | 1955 | 0.069 | 0.0181 | No |
| 15 | NAPSA | NAPSA Entrez, Source | napsin A aspartic peptidase | 2149 | 0.065 | 0.0110 | No |
| 16 | RGS14 | RGS14 Entrez, Source | regulator of G-protein signalling 14 | 2180 | 0.064 | 0.0162 | No |
| 17 | CTHRC1 | CTHRC1 Entrez, Source | collagen triple helix repeat containing 1 | 2305 | 0.062 | 0.0140 | No |
| 18 | SLC12A6 | SLC12A6 Entrez, Source | solute carrier family 12 (potassium/chloride transporters), member 6 | 2363 | 0.061 | 0.0167 | No |
| 19 | LIF | LIF Entrez, Source | leukemia inhibitory factor (cholinergic differentiation factor) | 2660 | 0.056 | 0.0009 | No |
| 20 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 2668 | 0.056 | 0.0068 | No |
| 21 | COMMD5 | COMMD5 Entrez, Source | COMM domain containing 5 | 2697 | 0.055 | 0.0111 | No |
| 22 | LIFR | LIFR Entrez, Source | leukemia inhibitory factor receptor alpha | 2859 | 0.052 | 0.0050 | No |
| 23 | IL3 | IL3 Entrez, Source | interleukin 3 (colony-stimulating factor, multiple) | 2935 | 0.051 | 0.0053 | No |
| 24 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 3016 | 0.050 | 0.0050 | No |
| 25 | ITGB7 | ITGB7 Entrez, Source | integrin, beta 7 | 3051 | 0.049 | 0.0082 | No |
| 26 | DAB2IP | DAB2IP Entrez, Source | DAB2 interacting protein | 3298 | 0.046 | -0.0051 | No |
| 27 | GTF2A1 | GTF2A1 Entrez, Source | general transcription factor IIA, 1, 19/37kDa | 3533 | 0.042 | -0.0179 | No |
| 28 | GABRA6 | GABRA6 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 6 | 3572 | 0.042 | -0.0159 | No |
| 29 | ETF1 | ETF1 Entrez, Source | eukaryotic translation termination factor 1 | 3680 | 0.041 | -0.0192 | No |
| 30 | AGTRL1 | AGTRL1 Entrez, Source | angiotensin II receptor-like 1 | 3751 | 0.040 | -0.0199 | No |
| 31 | CHAD | CHAD Entrez, Source | chondroadherin | 3762 | 0.040 | -0.0161 | No |
| 32 | RPP21 | RPP21 Entrez, Source | ribonuclease P 21kDa subunit | 4116 | 0.035 | -0.0387 | No |
| 33 | AKT2 | AKT2 Entrez, Source | v-akt murine thymoma viral oncogene homolog 2 | 4202 | 0.034 | -0.0412 | No |
| 34 | DGKZ | DGKZ Entrez, Source | diacylglycerol kinase, zeta 104kDa | 4311 | 0.032 | -0.0456 | No |
| 35 | OTP | OTP Entrez, Source | orthopedia homolog (Drosophila) | 4337 | 0.032 | -0.0438 | No |
| 36 | TBX5 | TBX5 Entrez, Source | T-box 5 | 4682 | 0.028 | -0.0665 | No |
| 37 | RUNX2 | RUNX2 Entrez, Source | runt-related transcription factor 2 | 4704 | 0.027 | -0.0649 | No |
| 38 | TRPM8 | TRPM8 Entrez, Source | transient receptor potential cation channel, subfamily M, member 8 | 4868 | 0.026 | -0.0743 | No |
| 39 | PROK2 | PROK2 Entrez, Source | prokineticin 2 | 4883 | 0.025 | -0.0724 | No |
| 40 | AGPAT4 | AGPAT4 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 4 (lysophosphatidic acid acyltransferase, delta) | 4985 | 0.024 | -0.0772 | No |
| 41 | ATP5A1 | ATP5A1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle | 5149 | 0.022 | -0.0870 | No |
| 42 | C1QTNF6 | C1QTNF6 Entrez, Source | C1q and tumor necrosis factor related protein 6 | 5205 | 0.021 | -0.0887 | No |
| 43 | CITED1 | CITED1 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 1 | 5242 | 0.021 | -0.0890 | No |
| 44 | NUTF2 | NUTF2 Entrez, Source | nuclear transport factor 2 | 5297 | 0.020 | -0.0907 | No |
| 45 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 5318 | 0.020 | -0.0899 | No |
| 46 | HEY1 | HEY1 Entrez, Source | hairy/enhancer-of-split related with YRPW motif 1 | 5501 | 0.018 | -0.1015 | No |
| 47 | GPR50 | GPR50 Entrez, Source | G protein-coupled receptor 50 | 5581 | 0.017 | -0.1054 | No |
| 48 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 5628 | 0.017 | -0.1070 | No |
| 49 | IL17RE | IL17RE Entrez, Source | interleukin 17 receptor E | 5891 | 0.014 | -0.1251 | No |
| 50 | EIF5A | EIF5A Entrez, Source | eukaryotic translation initiation factor 5A | 5996 | 0.013 | -0.1315 | No |
| 51 | ERG | ERG Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog (avian) | 6202 | 0.011 | -0.1457 | No |
| 52 | LUC7L | LUC7L Entrez, Source | LUC7-like (S. cerevisiae) | 6518 | 0.007 | -0.1687 | No |
| 53 | KCNJ1 | KCNJ1 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 1 | 6556 | 0.007 | -0.1707 | No |
| 54 | PDE6H | PDE6H Entrez, Source | phosphodiesterase 6H, cGMP-specific, cone, gamma | 6641 | 0.006 | -0.1763 | No |
| 55 | NEK6 | NEK6 Entrez, Source | NIMA (never in mitosis gene a)-related kinase 6 | 7630 | -0.005 | -0.2503 | No |
| 56 | CCL4 | CCL4 Entrez, Source | chemokine (C-C motif) ligand 4 | 7645 | -0.005 | -0.2508 | No |
| 57 | MKLN1 | MKLN1 Entrez, Source | muskelin 1, intracellular mediator containing kelch motifs | 7647 | -0.005 | -0.2503 | No |
| 58 | NUCB2 | NUCB2 Entrez, Source | nucleobindin 2 | 7742 | -0.006 | -0.2566 | No |
| 59 | RBBP6 | RBBP6 Entrez, Source | retinoblastoma binding protein 6 | 7799 | -0.007 | -0.2600 | No |
| 60 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 8082 | -0.010 | -0.2801 | No |
| 61 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 8159 | -0.011 | -0.2846 | No |
| 62 | CD6 | CD6 Entrez, Source | CD6 molecule | 8196 | -0.012 | -0.2859 | No |
| 63 | IL17F | IL17F Entrez, Source | interleukin 17F | 8354 | -0.014 | -0.2962 | No |
| 64 | ARHGEF2 | ARHGEF2 Entrez, Source | rho/rac guanine nucleotide exchange factor (GEF) 2 | 8556 | -0.016 | -0.3095 | No |
| 65 | PTK9 | PTK9 Entrez, Source | PTK9 protein tyrosine kinase 9 | 8672 | -0.018 | -0.3162 | No |
| 66 | CXCR3 | CXCR3 Entrez, Source | chemokine (C-X-C motif) receptor 3 | 8942 | -0.021 | -0.3340 | No |
| 67 | POU2F1 | POU2F1 Entrez, Source | POU domain, class 2, transcription factor 1 | 9137 | -0.024 | -0.3459 | No |
| 68 | CSF2 | CSF2 Entrez, Source | colony stimulating factor 2 (granulocyte-macrophage) | 9237 | -0.025 | -0.3505 | No |
| 69 | RIN1 | RIN1 Entrez, Source | Ras and Rab interactor 1 | 9335 | -0.026 | -0.3548 | No |
| 70 | IFNG | IFNG Entrez, Source | interferon, gamma | 9423 | -0.028 | -0.3581 | No |
| 71 | ACIN1 | ACIN1 Entrez, Source | apoptotic chromatin condensation inducer 1 | 9476 | -0.029 | -0.3588 | No |
| 72 | DMP1 | DMP1 Entrez, Source | dentin matrix acidic phosphoprotein | 9727 | -0.032 | -0.3739 | No |
| 73 | SOST | SOST Entrez, Source | sclerosteosis | 9770 | -0.033 | -0.3732 | No |
| 74 | PGF | PGF Entrez, Source | placental growth factor, vascular endothelial growth factor-related protein | 9927 | -0.035 | -0.3809 | No |
| 75 | HEMGN | HEMGN Entrez, Source | hemogen | 9968 | -0.036 | -0.3798 | No |
| 76 | GPX1 | GPX1 Entrez, Source | glutathione peroxidase 1 | 10706 | -0.050 | -0.4297 | No |
| 77 | CD28 | CD28 Entrez, Source | CD28 molecule | 10719 | -0.050 | -0.4248 | No |
| 78 | SCN2B | SCN2B Entrez, Source | sodium channel, voltage-gated, type II, beta | 10782 | -0.051 | -0.4236 | No |
| 79 | TACC2 | TACC2 Entrez, Source | transforming, acidic coiled-coil containing protein 2 | 10838 | -0.052 | -0.4217 | No |
| 80 | TLN1 | TLN1 Entrez, Source | talin 1 | 10840 | -0.052 | -0.4157 | No |
| 81 | MAP1LC3A | MAP1LC3A Entrez, Source | microtubule-associated protein 1 light chain 3 alpha | 11020 | -0.056 | -0.4227 | No |
| 82 | ENTPD3 | ENTPD3 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 3 | 11135 | -0.059 | -0.4245 | No |
| 83 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 11375 | -0.064 | -0.4350 | No |
| 84 | WBP1 | WBP1 Entrez, Source | WW domain binding protein 1 | 11647 | -0.074 | -0.4470 | No |
| 85 | ARHGAP8 | ARHGAP8 Entrez, Source | Rho GTPase activating protein 8 | 11819 | -0.080 | -0.4506 | No |
| 86 | MTMR3 | MTMR3 Entrez, Source | myotubularin related protein 3 | 11869 | -0.082 | -0.4448 | No |
| 87 | RHOG | RHOG Entrez, Source | ras homolog gene family, member G (rho G) | 12212 | -0.097 | -0.4593 | Yes |
| 88 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 12285 | -0.102 | -0.4529 | Yes |
| 89 | HPCAL1 | HPCAL1 Entrez, Source | hippocalcin-like 1 | 12394 | -0.111 | -0.4483 | Yes |
| 90 | TNFRSF12A | TNFRSF12A Entrez, Source | tumor necrosis factor receptor superfamily, member 12A | 12397 | -0.111 | -0.4356 | Yes |
| 91 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 12424 | -0.112 | -0.4245 | Yes |
| 92 | PPP1R14C | PPP1R14C Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 14C | 12457 | -0.115 | -0.4136 | Yes |
| 93 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 12465 | -0.115 | -0.4008 | Yes |
| 94 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 12519 | -0.119 | -0.3910 | Yes |
| 95 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 12993 | -0.186 | -0.4051 | Yes |
| 96 | DAB2 | DAB2 Entrez, Source | disabled homolog 2, mitogen-responsive phosphoprotein (Drosophila) | 13130 | -0.247 | -0.3868 | Yes |
| 97 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 13180 | -0.281 | -0.3580 | Yes |
| 98 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 13220 | -0.314 | -0.3245 | Yes |
| 99 | PDGFC | PDGFC Entrez, Source | platelet derived growth factor C | 13269 | -0.383 | -0.2837 | Yes |
| 100 | CD69 | CD69 Entrez, Source | CD69 molecule | 13297 | -0.457 | -0.2328 | Yes |
| 101 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 13298 | -0.461 | -0.1793 | Yes |
| 102 | SLC31A2 | SLC31A2 Entrez, Source | solute carrier family 31 (copper transporters), member 2 | 13300 | -0.468 | -0.1252 | Yes |
| 103 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 13307 | -0.479 | -0.0701 | Yes |
| 104 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 13326 | -0.627 | 0.0012 | Yes |