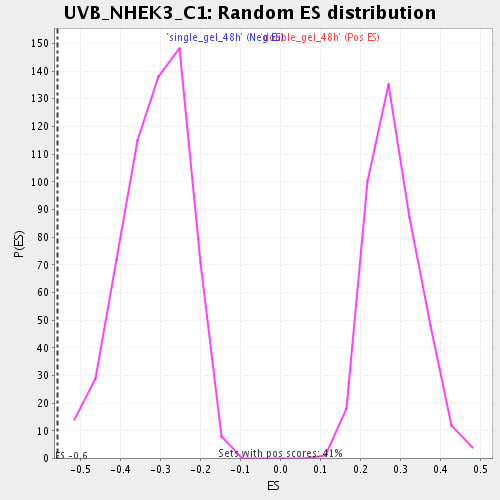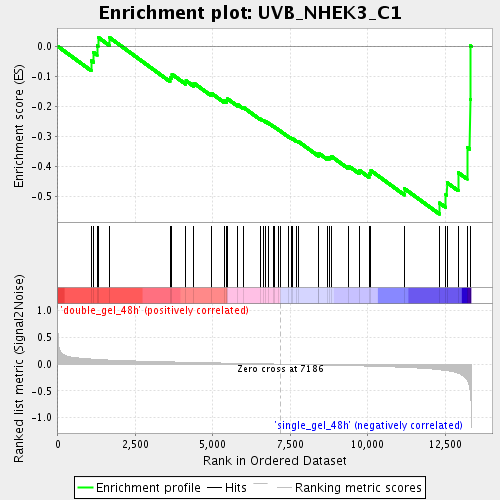
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | single_gel_48h |
| GeneSet | UVB_NHEK3_C1 |
| Enrichment Score (ES) | -0.5579799 |
| Normalized Enrichment Score (NES) | -1.7791995 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.045989603 |
| FWER p-Value | 0.988 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PBEF1 | PBEF1 Entrez, Source | pre-B-cell colony enhancing factor 1 | 1074 | 0.094 | -0.0472 | No |
| 2 | UBE2I | UBE2I Entrez, Source | ubiquitin-conjugating enzyme E2I (UBC9 homolog, yeast) | 1146 | 0.092 | -0.0199 | No |
| 3 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 1264 | 0.087 | 0.0023 | No |
| 4 | IER2 | IER2 Entrez, Source | immediate early response 2 | 1296 | 0.086 | 0.0306 | No |
| 5 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 1670 | 0.075 | 0.0292 | No |
| 6 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 3619 | 0.042 | -0.1025 | No |
| 7 | ETF1 | ETF1 Entrez, Source | eukaryotic translation termination factor 1 | 3680 | 0.041 | -0.0925 | No |
| 8 | SAFB | SAFB Entrez, Source | scaffold attachment factor B | 4133 | 0.035 | -0.1141 | No |
| 9 | DDX1 | DDX1 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 1 | 4387 | 0.031 | -0.1220 | No |
| 10 | GALNT3 | GALNT3 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 3 (GalNAc-T3) | 4949 | 0.024 | -0.1554 | No |
| 11 | COPS8 | COPS8 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 8 (Arabidopsis) | 5364 | 0.020 | -0.1796 | No |
| 12 | VDAC2 | VDAC2 Entrez, Source | voltage-dependent anion channel 2 | 5456 | 0.019 | -0.1797 | No |
| 13 | PSMA6 | PSMA6 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 6 | 5475 | 0.019 | -0.1745 | No |
| 14 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 5802 | 0.015 | -0.1936 | No |
| 15 | GSPT1 | GSPT1 Entrez, Source | G1 to S phase transition 1 | 5981 | 0.013 | -0.2024 | No |
| 16 | RNF4 | RNF4 Entrez, Source | ring finger protein 4 | 6533 | 0.007 | -0.2413 | No |
| 17 | TSG101 | TSG101 Entrez, Source | tumor susceptibility gene 101 | 6620 | 0.006 | -0.2455 | No |
| 18 | UBXD2 | UBXD2 Entrez, Source | UBX domain containing 2 | 6708 | 0.005 | -0.2502 | No |
| 19 | ATP6V1B2 | ATP6V1B2 Entrez, Source | ATPase, H+ transporting, lysosomal 56/58kDa, V1 subunit B2 | 6788 | 0.004 | -0.2546 | No |
| 20 | GPX2 | GPX2 Entrez, Source | glutathione peroxidase 2 (gastrointestinal) | 6946 | 0.003 | -0.2655 | No |
| 21 | CSNK2A1 | CSNK2A1 Entrez, Source | casein kinase 2, alpha 1 polypeptide | 6997 | 0.002 | -0.2686 | No |
| 22 | LCN2 | LCN2 Entrez, Source | lipocalin 2 (oncogene 24p3) | 7114 | 0.001 | -0.2771 | No |
| 23 | PSMA5 | PSMA5 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 5 | 7199 | -0.000 | -0.2833 | No |
| 24 | KCNK1 | KCNK1 Entrez, Source | potassium channel, subfamily K, member 1 | 7451 | -0.003 | -0.3011 | No |
| 25 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 7540 | -0.004 | -0.3062 | No |
| 26 | PSMC6 | PSMC6 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 6 | 7584 | -0.005 | -0.3077 | No |
| 27 | ATP1B1 | ATP1B1 Entrez, Source | ATPase, Na+/K+ transporting, beta 1 polypeptide | 7708 | -0.006 | -0.3149 | No |
| 28 | PSMC1 | PSMC1 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 1 | 7772 | -0.007 | -0.3173 | No |
| 29 | PSMC2 | PSMC2 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 2 | 8416 | -0.014 | -0.3605 | No |
| 30 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 8419 | -0.014 | -0.3555 | No |
| 31 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 8690 | -0.018 | -0.3694 | No |
| 32 | HSP90AA1 | HSP90AA1 Entrez, Source | heat shock protein 90kDa alpha (cytosolic), class A member 1 | 8764 | -0.019 | -0.3682 | No |
| 33 | UBE2G1 | UBE2G1 Entrez, Source | ubiquitin-conjugating enzyme E2G 1 (UBC7 homolog, yeast) | 8829 | -0.020 | -0.3661 | No |
| 34 | CNN3 | CNN3 Entrez, Source | calponin 3, acidic | 9383 | -0.027 | -0.3980 | No |
| 35 | GLUL | GLUL Entrez, Source | glutamate-ammonia ligase (glutamine synthetase) | 9734 | -0.033 | -0.4127 | No |
| 36 | WBSCR1 | WBSCR1 Entrez, Source | Williams-Beuren syndrome chromosome region 1 | 10043 | -0.037 | -0.4225 | No |
| 37 | GPIAP1 | GPIAP1 Entrez, Source | GPI-anchored membrane protein 1 | 10100 | -0.038 | -0.4131 | No |
| 38 | MAP2K3 | MAP2K3 Entrez, Source | mitogen-activated protein kinase kinase 3 | 11186 | -0.060 | -0.4733 | No |
| 39 | TJP2 | TJP2 Entrez, Source | tight junction protein 2 (zona occludens 2) | 12313 | -0.104 | -0.5208 | Yes |
| 40 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 12519 | -0.119 | -0.4939 | Yes |
| 41 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 12557 | -0.122 | -0.4532 | Yes |
| 42 | CST6 | CST6 Entrez, Source | cystatin E/M | 12920 | -0.170 | -0.4199 | Yes |
| 43 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 13206 | -0.298 | -0.3354 | Yes |
| 44 | SLC31A2 | SLC31A2 Entrez, Source | solute carrier family 31 (copper transporters), member 2 | 13300 | -0.468 | -0.1760 | Yes |
| 45 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13313 | -0.503 | 0.0022 | Yes |