Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_48h_versus_single_gel_48h.class.cls #double_gel_48h_versus_single_gel_48h_repos |
| Phenotype | class.cls#double_gel_48h_versus_single_gel_48h_repos |
| Upregulated in class | single_gel_48h |
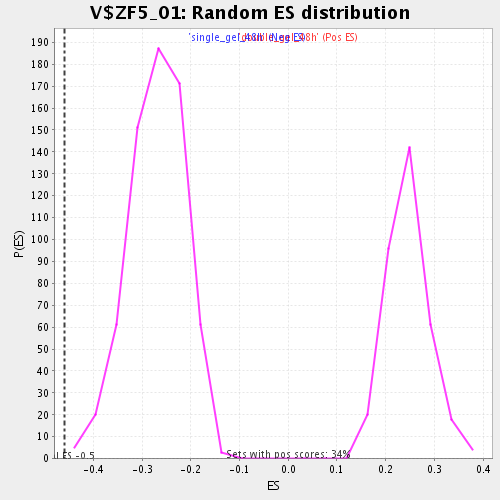
| GeneSet | V$ZF5_01 |
| Enrichment Score (ES) | -0.45941156 |
| Normalized Enrichment Score (NES) | -1.7025756 |
| Nominal p-value | 0.0015174507 |
| FDR q-value | 0.066080965 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | POU3F1 | POU3F1 Entrez, Source | POU domain, class 3, transcription factor 1 | 291 | 0.150 | -0.0009 | No |
| 2 | KCNH5 | KCNH5 Entrez, Source | potassium voltage-gated channel, subfamily H (eag-related), member 5 | 602 | 0.118 | -0.0078 | No |
| 3 | CRY1 | CRY1 Entrez, Source | cryptochrome 1 (photolyase-like) | 652 | 0.115 | 0.0047 | No |
| 4 | NXPH4 | NXPH4 Entrez, Source | neurexophilin 4 | 763 | 0.108 | 0.0116 | No |
| 5 | ACCN2 | ACCN2 Entrez, Source | amiloride-sensitive cation channel 2, neuronal | 954 | 0.099 | 0.0112 | No |
| 6 | POU4F1 | POU4F1 Entrez, Source | POU domain, class 4, transcription factor 1 | 968 | 0.099 | 0.0240 | No |
| 7 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 1228 | 0.089 | 0.0169 | No |
| 8 | GJB6 | GJB6 Entrez, Source | gap junction protein, beta 6 (connexin 30) | 1376 | 0.083 | 0.0175 | No |
| 9 | WNT11 | WNT11 Entrez, Source | wingless-type MMTV integration site family, member 11 | 1708 | 0.074 | 0.0029 | No |
| 10 | NR2F2 | NR2F2 Entrez, Source | nuclear receptor subfamily 2, group F, member 2 | 1957 | 0.069 | -0.0062 | No |
| 11 | PA2G4 | PA2G4 Entrez, Source | proliferation-associated 2G4, 38kDa | 1958 | 0.069 | 0.0035 | No |
| 12 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 2033 | 0.067 | 0.0073 | No |
| 13 | TMOD2 | TMOD2 Entrez, Source | tropomodulin 2 (neuronal) | 2592 | 0.057 | -0.0268 | No |
| 14 | LZTS2 | LZTS2 Entrez, Source | leucine zipper, putative tumor suppressor 2 | 2654 | 0.056 | -0.0236 | No |
| 15 | FOXD3 | FOXD3 Entrez, Source | forkhead box D3 | 3243 | 0.047 | -0.0614 | No |
| 16 | CACNA1A | CACNA1A Entrez, Source | calcium channel, voltage-dependent, P/Q type, alpha 1A subunit | 3318 | 0.046 | -0.0606 | No |
| 17 | GRIK3 | GRIK3 Entrez, Source | glutamate receptor, ionotropic, kainate 3 | 3319 | 0.046 | -0.0543 | No |
| 18 | SYT12 | SYT12 Entrez, Source | synaptotagmin XII | 3457 | 0.044 | -0.0585 | No |
| 19 | DCAMKL1 | DCAMKL1 Entrez, Source | doublecortin and CaM kinase-like 1 | 3621 | 0.041 | -0.0650 | No |
| 20 | RASGRF1 | RASGRF1 Entrez, Source | Ras protein-specific guanine nucleotide-releasing factor 1 | 4010 | 0.036 | -0.0892 | No |
| 21 | NSF | NSF Entrez, Source | N-ethylmaleimide-sensitive factor | 4152 | 0.034 | -0.0950 | No |
| 22 | RGS7 | RGS7 Entrez, Source | regulator of G-protein signalling 7 | 4336 | 0.032 | -0.1043 | No |
| 23 | ELAVL3 | ELAVL3 Entrez, Source | ELAV (embryonic lethal, abnormal vision, Drosophila)-like 3 (Hu antigen C) | 4360 | 0.032 | -0.1016 | No |
| 24 | DNM2 | DNM2 Entrez, Source | dynamin 2 | 4689 | 0.028 | -0.1225 | No |
| 25 | NTN1 | NTN1 Entrez, Source | netrin 1 | 4762 | 0.027 | -0.1242 | No |
| 26 | UBE2N | UBE2N Entrez, Source | ubiquitin-conjugating enzyme E2N (UBC13 homolog, yeast) | 4960 | 0.024 | -0.1357 | No |
| 27 | NUTF2 | NUTF2 Entrez, Source | nuclear transport factor 2 | 5297 | 0.020 | -0.1582 | No |
| 28 | KCND3 | KCND3 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 3 | 5549 | 0.018 | -0.1746 | No |
| 29 | CCNE1 | CCNE1 Entrez, Source | cyclin E1 | 5568 | 0.017 | -0.1735 | No |
| 30 | EIF4A1 | EIF4A1 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 1 | 5627 | 0.017 | -0.1756 | No |
| 31 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 5645 | 0.017 | -0.1745 | No |
| 32 | PPIE | PPIE Entrez, Source | peptidylprolyl isomerase E (cyclophilin E) | 5766 | 0.015 | -0.1814 | No |
| 33 | DLST | DLST Entrez, Source | dihydrolipoamide S-succinyltransferase (E2 component of 2-oxo-glutarate complex) | 5882 | 0.014 | -0.1881 | No |
| 34 | ARHGEF12 | ARHGEF12 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 12 | 5956 | 0.013 | -0.1918 | No |
| 35 | ITPKA | ITPKA Entrez, Source | inositol 1,4,5-trisphosphate 3-kinase A | 6042 | 0.012 | -0.1965 | No |
| 36 | HCN4 | HCN4 Entrez, Source | hyperpolarization activated cyclic nucleotide-gated potassium channel 4 | 6062 | 0.012 | -0.1962 | No |
| 37 | KIF1B | KIF1B Entrez, Source | kinesin family member 1B | 6130 | 0.012 | -0.1996 | No |
| 38 | GFER | GFER Entrez, Source | growth factor, augmenter of liver regeneration (ERV1 homolog, S. cerevisiae) | 6377 | 0.009 | -0.2170 | No |
| 39 | UBTF | UBTF Entrez, Source | upstream binding transcription factor, RNA polymerase I | 6405 | 0.008 | -0.2178 | No |
| 40 | HPCA | HPCA Entrez, Source | hippocalcin | 6474 | 0.008 | -0.2219 | No |
| 41 | RPL41 | RPL41 Entrez, Source | ribosomal protein L41 | 6583 | 0.007 | -0.2291 | No |
| 42 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 6992 | 0.002 | -0.2596 | No |
| 43 | STAT5B | STAT5B Entrez, Source | signal transducer and activator of transcription 5B | 7024 | 0.002 | -0.2618 | No |
| 44 | EIF3S10 | EIF3S10 Entrez, Source | eukaryotic translation initiation factor 3, subunit 10 theta, 150/170kDa | 7201 | -0.000 | -0.2750 | No |
| 45 | ABCC8 | ABCC8 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 8 | 7488 | -0.004 | -0.2961 | No |
| 46 | TLE3 | TLE3 Entrez, Source | transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) | 7718 | -0.006 | -0.3126 | No |
| 47 | FUT8 | FUT8 Entrez, Source | fucosyltransferase 8 (alpha (1,6) fucosyltransferase) | 7725 | -0.006 | -0.3122 | No |
| 48 | PHOX2A | PHOX2A Entrez, Source | paired-like (aristaless) homeobox 2a | 7751 | -0.006 | -0.3131 | No |
| 49 | APBB1 | APBB1 Entrez, Source | amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65) | 7884 | -0.008 | -0.3220 | No |
| 50 | PABPC4 | PABPC4 Entrez, Source | poly(A) binding protein, cytoplasmic 4 (inducible form) | 8106 | -0.011 | -0.3372 | No |
| 51 | KPNB1 | KPNB1 Entrez, Source | karyopherin (importin) beta 1 | 8243 | -0.012 | -0.3457 | No |
| 52 | TRPM7 | TRPM7 Entrez, Source | transient receptor potential cation channel, subfamily M, member 7 | 8513 | -0.016 | -0.3638 | No |
| 53 | GRIN2B | GRIN2B Entrez, Source | glutamate receptor, ionotropic, N-methyl D-aspartate 2B | 8589 | -0.017 | -0.3672 | No |
| 54 | NGFR | NGFR Entrez, Source | nerve growth factor receptor (TNFR superfamily, member 16) | 8817 | -0.019 | -0.3816 | No |
| 55 | NFATC4 | NFATC4 Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 | 8843 | -0.020 | -0.3807 | No |
| 56 | CDC42BPB | CDC42BPB Entrez, Source | CDC42 binding protein kinase beta (DMPK-like) | 8892 | -0.020 | -0.3815 | No |
| 57 | SUMO2 | SUMO2 Entrez, Source | SMT3 suppressor of mif two 3 homolog 2 (S. cerevisiae) | 8896 | -0.020 | -0.3788 | No |
| 58 | PPP1CC | PPP1CC Entrez, Source | protein phosphatase 1, catalytic subunit, gamma isoform | 8913 | -0.021 | -0.3771 | No |
| 59 | SEP15 | SEP15 Entrez, Source | - | 9382 | -0.027 | -0.4087 | No |
| 60 | RNF10 | RNF10 Entrez, Source | ring finger protein 10 | 9398 | -0.027 | -0.4059 | No |
| 61 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 9599 | -0.030 | -0.4168 | No |
| 62 | SMPD3 | SMPD3 Entrez, Source | sphingomyelin phosphodiesterase 3, neutral membrane (neutral sphingomyelinase II) | 9805 | -0.033 | -0.4276 | No |
| 63 | SENP3 | SENP3 Entrez, Source | SUMO1/sentrin/SMT3 specific peptidase 3 | 9926 | -0.035 | -0.4317 | No |
| 64 | USP48 | USP48 Entrez, Source | ubiquitin specific peptidase 48 | 9967 | -0.036 | -0.4296 | No |
| 65 | UQCRH | UQCRH Entrez, Source | ubiquinol-cytochrome c reductase hinge protein | 10061 | -0.038 | -0.4314 | No |
| 66 | XYLT2 | XYLT2 Entrez, Source | xylosyltransferase II | 10145 | -0.039 | -0.4322 | No |
| 67 | KCNC1 | KCNC1 Entrez, Source | potassium voltage-gated channel, Shaw-related subfamily, member 1 | 10154 | -0.039 | -0.4272 | No |
| 68 | TPP2 | TPP2 Entrez, Source | tripeptidyl peptidase II | 10260 | -0.041 | -0.4295 | No |
| 69 | SPRED1 | SPRED1 Entrez, Source | sprouty-related, EVH1 domain containing 1 | 10322 | -0.042 | -0.4282 | No |
| 70 | BTRC | BTRC Entrez, Source | beta-transducin repeat containing | 10328 | -0.042 | -0.4227 | No |
| 71 | SEZ6 | SEZ6 Entrez, Source | seizure related 6 homolog (mouse) | 10436 | -0.044 | -0.4246 | No |
| 72 | CALM1 | CALM1 Entrez, Source | calmodulin 1 (phosphorylase kinase, delta) | 10481 | -0.045 | -0.4216 | No |
| 73 | MAX | MAX Entrez, Source | MYC associated factor X | 10590 | -0.047 | -0.4232 | No |
| 74 | PTBP1 | PTBP1 Entrez, Source | polypyrimidine tract binding protein 1 | 10613 | -0.047 | -0.4183 | No |
| 75 | GADD45B | GADD45B Entrez, Source | growth arrest and DNA-damage-inducible, beta | 10614 | -0.047 | -0.4117 | No |
| 76 | MPZL1 | MPZL1 Entrez, Source | myelin protein zero-like 1 | 10998 | -0.056 | -0.4327 | No |
| 77 | GNB1 | GNB1 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 1 | 11004 | -0.056 | -0.4253 | No |
| 78 | PAK1 | PAK1 Entrez, Source | p21/Cdc42/Rac1-activated kinase 1 (STE20 homolog, yeast) | 11055 | -0.057 | -0.4210 | No |
| 79 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 11170 | -0.060 | -0.4213 | No |
| 80 | ARHGEF7 | ARHGEF7 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 7 | 11238 | -0.061 | -0.4178 | No |
| 81 | ATP10A | ATP10A Entrez, Source | ATPase, Class V, type 10A | 11371 | -0.064 | -0.4188 | No |
| 82 | SYT7 | SYT7 Entrez, Source | synaptotagmin VII | 11610 | -0.072 | -0.4266 | No |
| 83 | LAMP1 | LAMP1 Entrez, Source | lysosomal-associated membrane protein 1 | 12040 | -0.089 | -0.4465 | Yes |
| 84 | RHOG | RHOG Entrez, Source | ras homolog gene family, member G (rho G) | 12212 | -0.097 | -0.4457 | Yes |
| 85 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 12285 | -0.102 | -0.4368 | Yes |
| 86 | HS2ST1 | HS2ST1 Entrez, Source | heparan sulfate 2-O-sulfotransferase 1 | 12339 | -0.107 | -0.4259 | Yes |
| 87 | TGIF | TGIF Entrez, Source | TGFB-induced factor (TALE family homeobox) | 12388 | -0.110 | -0.4140 | Yes |
| 88 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 12424 | -0.112 | -0.4009 | Yes |
| 89 | GPRC5C | GPRC5C Entrez, Source | G protein-coupled receptor, family C, group 5, member C | 12581 | -0.124 | -0.3953 | Yes |
| 90 | ILK | ILK Entrez, Source | integrin-linked kinase | 12613 | -0.126 | -0.3799 | Yes |
| 91 | HOMER3 | HOMER3 Entrez, Source | homer homolog 3 (Drosophila) | 12616 | -0.127 | -0.3623 | Yes |
| 92 | JUB | JUB Entrez, Source | jub, ajuba homolog (Xenopus laevis) | 12688 | -0.133 | -0.3490 | Yes |
| 93 | CAP1 | CAP1 Entrez, Source | CAP, adenylate cyclase-associated protein 1 (yeast) | 12748 | -0.141 | -0.3337 | Yes |
| 94 | PHC2 | PHC2 Entrez, Source | polyhomeotic homolog 2 (Drosophila) | 12831 | -0.153 | -0.3185 | Yes |
| 95 | DDAH1 | DDAH1 Entrez, Source | dimethylarginine dimethylaminohydrolase 1 | 12847 | -0.155 | -0.2978 | Yes |
| 96 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 12895 | -0.164 | -0.2784 | Yes |
| 97 | FOSL1 | FOSL1 Entrez, Source | FOS-like antigen 1 | 12919 | -0.170 | -0.2563 | Yes |
| 98 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 12980 | -0.185 | -0.2350 | Yes |
| 99 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 13083 | -0.223 | -0.2114 | Yes |
| 100 | RAB31 | RAB31 Entrez, Source | RAB31, member RAS oncogene family | 13200 | -0.292 | -0.1792 | Yes |
| 101 | ADM | ADM Entrez, Source | adrenomedullin | 13274 | -0.393 | -0.1295 | Yes |
| 102 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 13294 | -0.447 | -0.0682 | Yes |
| 103 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 13316 | -0.512 | 0.0020 | Yes |