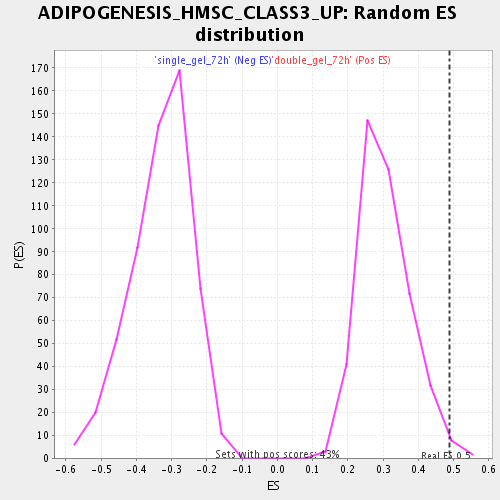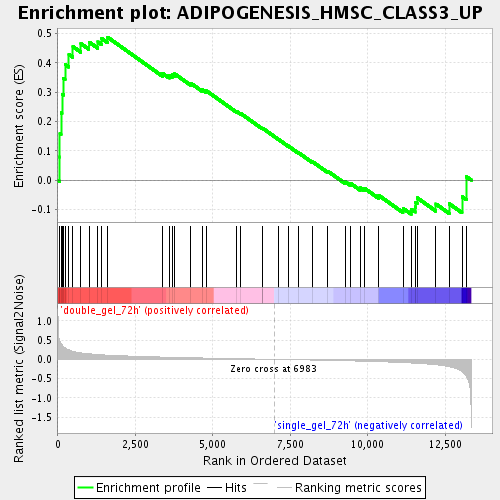
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | ADIPOGENESIS_HMSC_CLASS3_UP |
| Enrichment Score (ES) | 0.48703724 |
| Normalized Enrichment Score (NES) | 1.5921006 |
| Nominal p-value | 0.006960557 |
| FDR q-value | 0.071652435 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NFE2 | NFE2 Entrez, Source | nuclear factor (erythroid-derived 2), 45kDa | 63 | 0.480 | 0.0791 | Yes |
| 2 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 66 | 0.461 | 0.1593 | Yes |
| 3 | FASN | FASN Entrez, Source | fatty acid synthase | 101 | 0.417 | 0.2296 | Yes |
| 4 | LPIN1 | LPIN1 Entrez, Source | lipin 1 | 133 | 0.378 | 0.2932 | Yes |
| 5 | HMGCS1 | HMGCS1 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 1 (soluble) | 178 | 0.330 | 0.3475 | Yes |
| 6 | ACACA | ACACA Entrez, Source | acetyl-Coenzyme A carboxylase alpha | 231 | 0.293 | 0.3947 | Yes |
| 7 | ABCA1 | ABCA1 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 1 | 355 | 0.248 | 0.4287 | Yes |
| 8 | INSIG1 | INSIG1 Entrez, Source | insulin induced gene 1 | 485 | 0.207 | 0.4551 | Yes |
| 9 | ACACB | ACACB Entrez, Source | acetyl-Coenzyme A carboxylase beta | 742 | 0.168 | 0.4652 | Yes |
| 10 | MGP | MGP Entrez, Source | matrix Gla protein | 1003 | 0.144 | 0.4708 | Yes |
| 11 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 1291 | 0.125 | 0.4710 | Yes |
| 12 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 1413 | 0.118 | 0.4825 | Yes |
| 13 | ACSL5 | ACSL5 Entrez, Source | acyl-CoA synthetase long-chain family member 5 | 1606 | 0.109 | 0.4870 | Yes |
| 14 | EFNA1 | EFNA1 Entrez, Source | ephrin-A1 | 3378 | 0.058 | 0.3641 | No |
| 15 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 3592 | 0.054 | 0.3575 | No |
| 16 | ITGAM | ITGAM Entrez, Source | integrin, alpha M (complement component 3 receptor 3 subunit) | 3688 | 0.052 | 0.3594 | No |
| 17 | TMEM23 | TMEM23 Entrez, Source | transmembrane protein 23 | 3767 | 0.050 | 0.3624 | No |
| 18 | NAB1 | NAB1 Entrez, Source | NGFI-A binding protein 1 (EGR1 binding protein 1) | 4286 | 0.041 | 0.3307 | No |
| 19 | GAS6 | GAS6 Entrez, Source | growth arrest-specific 6 | 4656 | 0.035 | 0.3090 | No |
| 20 | LIPE | LIPE Entrez, Source | lipase, hormone-sensitive | 4794 | 0.033 | 0.3045 | No |
| 21 | PHKG1 | PHKG1 Entrez, Source | phosphorylase kinase, gamma 1 (muscle) | 5764 | 0.018 | 0.2347 | No |
| 22 | ADFP | ADFP Entrez, Source | adipose differentiation-related protein | 5901 | 0.016 | 0.2272 | No |
| 23 | EFTUD2 | EFTUD2 Entrez, Source | elongation factor Tu GTP binding domain containing 2 | 6604 | 0.005 | 0.1754 | No |
| 24 | IGF2 | IGF2 Entrez, Source | insulin-like growth factor 2 (somatomedin A) | 6610 | 0.005 | 0.1759 | No |
| 25 | SUMO1 | SUMO1 Entrez, Source | SMT3 suppressor of mif two 3 homolog 1 (S. cerevisiae) | 6611 | 0.005 | 0.1768 | No |
| 26 | NOVA1 | NOVA1 Entrez, Source | neuro-oncological ventral antigen 1 | 7115 | -0.002 | 0.1393 | No |
| 27 | NPTXR | NPTXR Entrez, Source | neuronal pentraxin receptor | 7428 | -0.006 | 0.1169 | No |
| 28 | NID1 | NID1 Entrez, Source | nidogen 1 | 7761 | -0.011 | 0.0940 | No |
| 29 | PPP1CC | PPP1CC Entrez, Source | protein phosphatase 1, catalytic subunit, gamma isoform | 8221 | -0.018 | 0.0626 | No |
| 30 | STOM | STOM Entrez, Source | stomatin | 8714 | -0.027 | 0.0303 | No |
| 31 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 9276 | -0.037 | -0.0054 | No |
| 32 | FABP4 | FABP4 Entrez, Source | fatty acid binding protein 4, adipocyte | 9450 | -0.041 | -0.0113 | No |
| 33 | MAPK1 | MAPK1 Entrez, Source | mitogen-activated protein kinase 1 | 9755 | -0.047 | -0.0260 | No |
| 34 | CYBRD1 | CYBRD1 Entrez, Source | cytochrome b reductase 1 | 9887 | -0.050 | -0.0271 | No |
| 35 | LEPREL1 | LEPREL1 Entrez, Source | leprecan-like 1 | 10355 | -0.060 | -0.0517 | No |
| 36 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 11138 | -0.083 | -0.0960 | No |
| 37 | PPP2R1B | PPP2R1B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), beta isoform | 11399 | -0.095 | -0.0990 | No |
| 38 | IGF2R | IGF2R Entrez, Source | insulin-like growth factor 2 receptor | 11527 | -0.101 | -0.0909 | No |
| 39 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 11545 | -0.102 | -0.0745 | No |
| 40 | STX3 | STX3 Entrez, Source | syntaxin 3 | 11595 | -0.104 | -0.0600 | No |
| 41 | SLC27A4 | SLC27A4 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 4 | 12200 | -0.139 | -0.0811 | No |
| 42 | CDH13 | CDH13 Entrez, Source | cadherin 13, H-cadherin (heart) | 12630 | -0.190 | -0.0802 | No |
| 43 | PRKAR2B | PRKAR2B Entrez, Source | protein kinase, cAMP-dependent, regulatory, type II, beta | 13042 | -0.314 | -0.0563 | No |
| 44 | RAB31 | RAB31 Entrez, Source | RAB31, member RAS oncogene family | 13187 | -0.451 | 0.0117 | No |