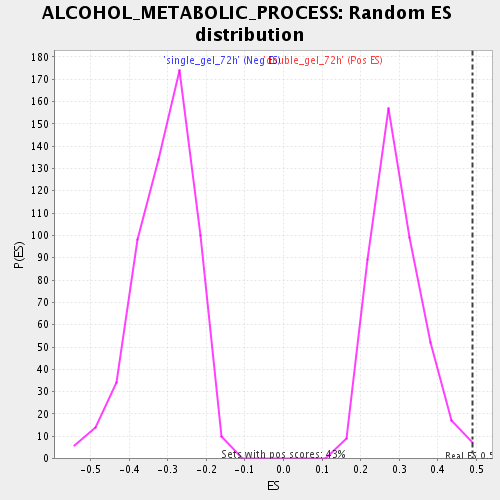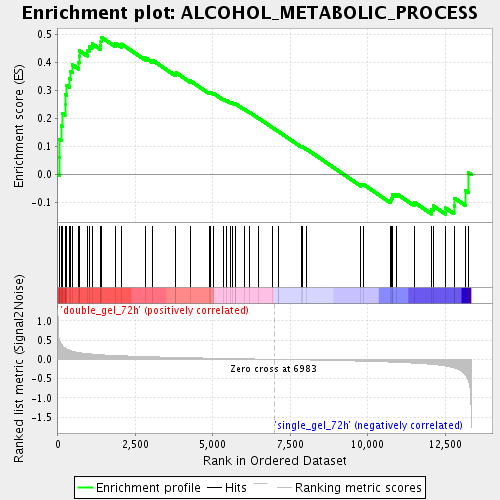
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | ALCOHOL_METABOLIC_PROCESS |
| Enrichment Score (ES) | 0.4900797 |
| Normalized Enrichment Score (NES) | 1.6764045 |
| Nominal p-value | 0.0023255814 |
| FDR q-value | 0.043369953 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FBP1 | FBP1 Entrez, Source | fructose-1,6-bisphosphatase 1 | 43 | 0.515 | 0.0627 | Yes |
| 2 | ADH7 | ADH7 Entrez, Source | alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide | 53 | 0.495 | 0.1254 | Yes |
| 3 | NR0B2 | NR0B2 Entrez, Source | nuclear receptor subfamily 0, group B, member 2 | 104 | 0.416 | 0.1749 | Yes |
| 4 | GCK | GCK Entrez, Source | glucokinase (hexokinase 4, maturity onset diabetes of the young 2) | 140 | 0.368 | 0.2193 | Yes |
| 5 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 230 | 0.293 | 0.2502 | Yes |
| 6 | MIOX | MIOX Entrez, Source | myo-inositol oxygenase | 234 | 0.291 | 0.2872 | Yes |
| 7 | ACN9 | ACN9 Entrez, Source | ACN9 homolog (S. cerevisiae) | 284 | 0.273 | 0.3185 | Yes |
| 8 | SLC25A10 | SLC25A10 Entrez, Source | solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 | 378 | 0.240 | 0.3422 | Yes |
| 9 | PDK4 | PDK4 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 4 | 412 | 0.227 | 0.3689 | Yes |
| 10 | FDPS | FDPS Entrez, Source | farnesyl diphosphate synthase (farnesyl pyrophosphate synthetase, dimethylallyltranstransferase, geranyltranstransferase) | 462 | 0.212 | 0.3924 | Yes |
| 11 | PDK1 | PDK1 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 1 | 673 | 0.176 | 0.3991 | Yes |
| 12 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 685 | 0.175 | 0.4207 | Yes |
| 13 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 693 | 0.174 | 0.4424 | Yes |
| 14 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 951 | 0.149 | 0.4421 | Yes |
| 15 | ALDOB | ALDOB Entrez, Source | aldolase B, fructose-bisphosphate | 1007 | 0.144 | 0.4564 | Yes |
| 16 | SULT1B1 | SULT1B1 Entrez, Source | sulfotransferase family, cytosolic, 1B, member 1 | 1102 | 0.137 | 0.4669 | Yes |
| 17 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 1370 | 0.121 | 0.4622 | Yes |
| 18 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 1387 | 0.119 | 0.4763 | Yes |
| 19 | COQ3 | COQ3 Entrez, Source | coenzyme Q3 homolog, methyltransferase (S. cerevisiae) | 1406 | 0.118 | 0.4901 | Yes |
| 20 | AKR1D1 | AKR1D1 Entrez, Source | aldo-keto reductase family 1, member D1 (delta 4-3-ketosteroid-5-beta-reductase) | 1859 | 0.099 | 0.4687 | No |
| 21 | ADH6 | ADH6 Entrez, Source | alcohol dehydrogenase 6 (class V) | 2053 | 0.092 | 0.4660 | No |
| 22 | PDK2 | PDK2 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 2 | 2834 | 0.071 | 0.4164 | No |
| 23 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 3058 | 0.065 | 0.4079 | No |
| 24 | ADH4 | ADH4 Entrez, Source | alcohol dehydrogenase 4 (class II), pi polypeptide | 3800 | 0.050 | 0.3585 | No |
| 25 | FPGT | FPGT Entrez, Source | fucose-1-phosphate guanylyltransferase | 3810 | 0.050 | 0.3642 | No |
| 26 | HDLBP | HDLBP Entrez, Source | high density lipoprotein binding protein (vigilin) | 4270 | 0.042 | 0.3350 | No |
| 27 | FBP2 | FBP2 Entrez, Source | fructose-1,6-bisphosphatase 2 | 4892 | 0.031 | 0.2922 | No |
| 28 | CLN8 | CLN8 Entrez, Source | ceroid-lipofuscinosis, neuronal 8 (epilepsy, progressive with mental retardation) | 4936 | 0.030 | 0.2928 | No |
| 29 | ARPP-19 | ARPP-19 Entrez, Source | - | 5020 | 0.029 | 0.2902 | No |
| 30 | GALK1 | GALK1 Entrez, Source | galactokinase 1 | 5339 | 0.024 | 0.2694 | No |
| 31 | CNBP | CNBP Entrez, Source | CCHC-type zinc finger, nucleic acid binding protein | 5438 | 0.023 | 0.2649 | No |
| 32 | MBTPS2 | MBTPS2 Entrez, Source | membrane-bound transcription factor peptidase, site 2 | 5566 | 0.021 | 0.2580 | No |
| 33 | UGP2 | UGP2 Entrez, Source | UDP-glucose pyrophosphorylase 2 | 5626 | 0.020 | 0.2561 | No |
| 34 | ALDOC | ALDOC Entrez, Source | aldolase C, fructose-bisphosphate | 5718 | 0.018 | 0.2516 | No |
| 35 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 5720 | 0.018 | 0.2539 | No |
| 36 | IPPK | IPPK Entrez, Source | inositol 1,3,4,5,6-pentakisphosphate 2-kinase | 6008 | 0.014 | 0.2341 | No |
| 37 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 6175 | 0.012 | 0.2231 | No |
| 38 | IL4 | IL4 Entrez, Source | interleukin 4 | 6466 | 0.007 | 0.2022 | No |
| 39 | GAPDHS | GAPDHS Entrez, Source | glyceraldehyde-3-phosphate dehydrogenase, spermatogenic | 6929 | 0.001 | 0.1675 | No |
| 40 | GALT | GALT Entrez, Source | galactose-1-phosphate uridylyltransferase | 7124 | -0.002 | 0.1531 | No |
| 41 | AKR1A1 | AKR1A1 Entrez, Source | aldo-keto reductase family 1, member A1 (aldehyde reductase) | 7127 | -0.002 | 0.1532 | No |
| 42 | GFPT2 | GFPT2 Entrez, Source | glutamine-fructose-6-phosphate transaminase 2 | 7852 | -0.013 | 0.1004 | No |
| 43 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 7883 | -0.013 | 0.0999 | No |
| 44 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 8036 | -0.015 | 0.0904 | No |
| 45 | ALDOA | ALDOA Entrez, Source | aldolase A, fructose-bisphosphate | 9779 | -0.047 | -0.0347 | No |
| 46 | GMDS | GMDS Entrez, Source | GDP-mannose 4,6-dehydratase | 9847 | -0.049 | -0.0335 | No |
| 47 | HPRT1 | HPRT1 Entrez, Source | hypoxanthine phosphoribosyltransferase 1 (Lesch-Nyhan syndrome) | 10730 | -0.071 | -0.0908 | No |
| 48 | PFKL | PFKL Entrez, Source | phosphofructokinase, liver | 10756 | -0.071 | -0.0836 | No |
| 49 | GFPT1 | GFPT1 Entrez, Source | glutamine-fructose-6-phosphate transaminase 1 | 10795 | -0.072 | -0.0772 | No |
| 50 | CEL | CEL Entrez, Source | carboxyl ester lipase (bile salt-stimulated lipase) | 10808 | -0.073 | -0.0688 | No |
| 51 | CHST1 | CHST1 Entrez, Source | carbohydrate (keratan sulfate Gal-6) sulfotransferase 1 | 10942 | -0.077 | -0.0689 | No |
| 52 | SCARB1 | SCARB1 Entrez, Source | scavenger receptor class B, member 1 | 11498 | -0.099 | -0.0980 | No |
| 53 | HK1 | HK1 Entrez, Source | hexokinase 1 | 12060 | -0.129 | -0.1238 | No |
| 54 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 12112 | -0.132 | -0.1107 | No |
| 55 | SPHK1 | SPHK1 Entrez, Source | sphingosine kinase 1 | 12507 | -0.172 | -0.1184 | No |
| 56 | ALDH3B1 | ALDH3B1 Entrez, Source | aldehyde dehydrogenase 3 family, member B1 | 12784 | -0.220 | -0.1110 | No |
| 57 | PFKM | PFKM Entrez, Source | phosphofructokinase, muscle | 12804 | -0.226 | -0.0835 | No |
| 58 | ABCG1 | ABCG1 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 1 | 13162 | -0.417 | -0.0570 | No |
| 59 | SOAT1 | SOAT1 Entrez, Source | sterol O-acyltransferase (acyl-Coenzyme A: cholesterol acyltransferase) 1 | 13248 | -0.551 | 0.0071 | No |