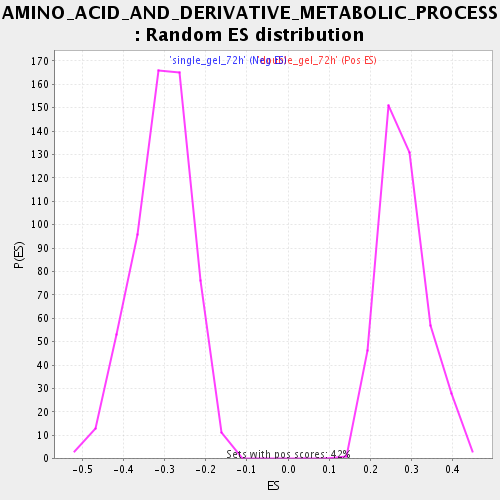
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | AMINO_ACID_AND_DERIVATIVE_METABOLIC_PROCESS |
| Enrichment Score (ES) | 0.53691113 |
| Normalized Enrichment Score (NES) | 1.9100411 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.006247458 |
| FWER p-Value | 0.413 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BBOX1 | BBOX1 Entrez, Source | butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 | 13 | 0.740 | 0.0654 | Yes |
| 2 | CDO1 | CDO1 Entrez, Source | cysteine dioxygenase, type I | 58 | 0.484 | 0.1055 | Yes |
| 3 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 110 | 0.403 | 0.1377 | Yes |
| 4 | HGD | HGD Entrez, Source | homogentisate 1,2-dioxygenase (homogentisate oxidase) | 126 | 0.385 | 0.1711 | Yes |
| 5 | ARG1 | ARG1 Entrez, Source | arginase, liver | 145 | 0.358 | 0.2019 | Yes |
| 6 | ASL | ASL Entrez, Source | argininosuccinate lyase | 183 | 0.323 | 0.2281 | Yes |
| 7 | FAH | FAH Entrez, Source | fumarylacetoacetate hydrolase (fumarylacetoacetase) | 219 | 0.299 | 0.2524 | Yes |
| 8 | GAMT | GAMT Entrez, Source | guanidinoacetate N-methyltransferase | 295 | 0.267 | 0.2707 | Yes |
| 9 | MAT1A | MAT1A Entrez, Source | methionine adenosyltransferase I, alpha | 315 | 0.262 | 0.2927 | Yes |
| 10 | SLC7A7 | SLC7A7 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 7 | 320 | 0.260 | 0.3157 | Yes |
| 11 | GLS2 | GLS2 Entrez, Source | glutaminase 2 (liver, mitochondrial) | 341 | 0.252 | 0.3368 | Yes |
| 12 | MSRA | MSRA Entrez, Source | methionine sulfoxide reductase A | 428 | 0.222 | 0.3502 | Yes |
| 13 | ALDH6A1 | ALDH6A1 Entrez, Source | aldehyde dehydrogenase 6 family, member A1 | 445 | 0.217 | 0.3685 | Yes |
| 14 | SCLY | SCLY Entrez, Source | selenocysteine lyase | 454 | 0.214 | 0.3871 | Yes |
| 15 | SLC25A15 | SLC25A15 Entrez, Source | solute carrier family 25 (mitochondrial carrier; ornithine transporter) member 15 | 595 | 0.186 | 0.3933 | Yes |
| 16 | BAAT | BAAT Entrez, Source | bile acid Coenzyme A: amino acid N-acyltransferase (glycine N-choloyltransferase) | 597 | 0.186 | 0.4099 | Yes |
| 17 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 612 | 0.184 | 0.4254 | Yes |
| 18 | PAH | PAH Entrez, Source | phenylalanine hydroxylase | 628 | 0.182 | 0.4407 | Yes |
| 19 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 651 | 0.179 | 0.4551 | Yes |
| 20 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 662 | 0.178 | 0.4703 | Yes |
| 21 | SLC7A2 | SLC7A2 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 | 776 | 0.165 | 0.4765 | Yes |
| 22 | GCSH | GCSH Entrez, Source | glycine cleavage system protein H (aminomethyl carrier) | 816 | 0.161 | 0.4880 | Yes |
| 23 | GCLC | GCLC Entrez, Source | glutamate-cysteine ligase, catalytic subunit | 823 | 0.160 | 0.5019 | Yes |
| 24 | BCKDK | BCKDK Entrez, Source | branched chain ketoacid dehydrogenase kinase | 868 | 0.156 | 0.5126 | Yes |
| 25 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 984 | 0.146 | 0.5170 | Yes |
| 26 | DHPS | DHPS Entrez, Source | deoxyhypusine synthase | 996 | 0.145 | 0.5292 | Yes |
| 27 | SULT1B1 | SULT1B1 Entrez, Source | sulfotransferase family, cytosolic, 1B, member 1 | 1102 | 0.137 | 0.5336 | Yes |
| 28 | AMT | AMT Entrez, Source | aminomethyltransferase | 1397 | 0.119 | 0.5220 | Yes |
| 29 | HDC | HDC Entrez, Source | histidine decarboxylase | 1527 | 0.112 | 0.5224 | Yes |
| 30 | QDPR | QDPR Entrez, Source | quinoid dihydropteridine reductase | 1560 | 0.110 | 0.5298 | Yes |
| 31 | ASRGL1 | ASRGL1 Entrez, Source | asparaginase like 1 | 1612 | 0.108 | 0.5357 | Yes |
| 32 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 1733 | 0.104 | 0.5360 | Yes |
| 33 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 1840 | 0.100 | 0.5369 | Yes |
| 34 | NFS1 | NFS1 Entrez, Source | NFS1 nitrogen fixation 1 (S. cerevisiae) | 2100 | 0.091 | 0.5255 | No |
| 35 | ADI1 | ADI1 Entrez, Source | acireductone dioxygenase 1 | 2159 | 0.089 | 0.5292 | No |
| 36 | PTS | PTS Entrez, Source | 6-pyruvoyltetrahydropterin synthase | 2407 | 0.082 | 0.5180 | No |
| 37 | BPHL | BPHL Entrez, Source | biphenyl hydrolase-like (serine hydrolase; breast epithelial mucin-associated antigen) | 2720 | 0.073 | 0.5010 | No |
| 38 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 3058 | 0.065 | 0.4814 | No |
| 39 | MCCC2 | MCCC2 Entrez, Source | methylcrotonoyl-Coenzyme A carboxylase 2 (beta) | 3517 | 0.056 | 0.4519 | No |
| 40 | P4HB | P4HB Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), beta polypeptide | 3536 | 0.055 | 0.4555 | No |
| 41 | GGTLA1 | GGTLA1 Entrez, Source | gamma-glutamyltransferase-like activity 1 | 3715 | 0.052 | 0.4467 | No |
| 42 | GSS | GSS Entrez, Source | glutathione synthetase | 4011 | 0.046 | 0.4286 | No |
| 43 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 4162 | 0.044 | 0.4212 | No |
| 44 | AARS | AARS Entrez, Source | alanyl-tRNA synthetase | 4275 | 0.042 | 0.4165 | No |
| 45 | HPD | HPD Entrez, Source | 4-hydroxyphenylpyruvate dioxygenase | 4383 | 0.040 | 0.4120 | No |
| 46 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 4470 | 0.038 | 0.4090 | No |
| 47 | GLUD1 | GLUD1 Entrez, Source | glutamate dehydrogenase 1 | 5087 | 0.028 | 0.3650 | No |
| 48 | COLQ | COLQ Entrez, Source | collagen-like tail subunit (single strand of homotrimer) of asymmetric acetylcholinesterase | 5644 | 0.020 | 0.3249 | No |
| 49 | GOT2 | GOT2 Entrez, Source | glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) | 5916 | 0.015 | 0.3059 | No |
| 50 | SCYE1 | SCYE1 Entrez, Source | small inducible cytokine subfamily E, member 1 (endothelial monocyte-activating) | 6040 | 0.014 | 0.2978 | No |
| 51 | DARS | DARS Entrez, Source | aspartyl-tRNA synthetase | 6478 | 0.007 | 0.2655 | No |
| 52 | GGT1 | GGT1 Entrez, Source | gamma-glutamyltransferase 1 | 6489 | 0.007 | 0.2653 | No |
| 53 | KARS | KARS Entrez, Source | lysyl-tRNA synthetase | 7266 | -0.004 | 0.2072 | No |
| 54 | SLC3A1 | SLC3A1 Entrez, Source | 1 | 7374 | -0.006 | 0.1997 | No |
| 55 | SLC7A8 | SLC7A8 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 8 | 8292 | -0.020 | 0.1323 | No |
| 56 | SDS | SDS Entrez, Source | serine dehydratase | 8683 | -0.026 | 0.1053 | No |
| 57 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 9012 | -0.032 | 0.0835 | No |
| 58 | OAZ1 | OAZ1 Entrez, Source | ornithine decarboxylase antizyme 1 | 9198 | -0.036 | 0.0727 | No |
| 59 | FARS2 | FARS2 Entrez, Source | phenylalanine-tRNA synthetase 2 (mitochondrial) | 9209 | -0.036 | 0.0752 | No |
| 60 | SLC7A9 | SLC7A9 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 | 9816 | -0.048 | 0.0338 | No |
| 61 | DIO2 | DIO2 Entrez, Source | deiodinase, iodothyronine, type II | 9834 | -0.049 | 0.0369 | No |
| 62 | SLC5A7 | SLC5A7 Entrez, Source | solute carrier family 5 (choline transporter), member 7 | 9983 | -0.052 | 0.0304 | No |
| 63 | HPRT1 | HPRT1 Entrez, Source | hypoxanthine phosphoribosyltransferase 1 (Lesch-Nyhan syndrome) | 10730 | -0.071 | -0.0195 | No |
| 64 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 10797 | -0.072 | -0.0180 | No |
| 65 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 10801 | -0.073 | -0.0117 | No |
| 66 | GATM | GATM Entrez, Source | glycine amidinotransferase (L-arginine:glycine amidinotransferase) | 11580 | -0.103 | -0.0611 | No |
| 67 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 12112 | -0.132 | -0.0892 | No |
| 68 | PLOD1 | PLOD1 Entrez, Source | procollagen-lysine 1, 2-oxoglutarate 5-dioxygenase 1 | 12626 | -0.189 | -0.1110 | No |
| 69 | DDAH2 | DDAH2 Entrez, Source | dimethylarginine dimethylaminohydrolase 2 | 12900 | -0.253 | -0.1089 | No |
| 70 | SLC6A6 | SLC6A6 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, taurine), member 6 | 13097 | -0.354 | -0.0919 | No |
| 71 | DDAH1 | DDAH1 Entrez, Source | dimethylarginine dimethylaminohydrolase 1 | 13118 | -0.367 | -0.0605 | No |
| 72 | SMS | SMS Entrez, Source | spermine synthase | 13163 | -0.420 | -0.0262 | No |
| 73 | BCAT1 | BCAT1 Entrez, Source | branched chain aminotransferase 1, cytosolic | 13181 | -0.442 | 0.0121 | No |