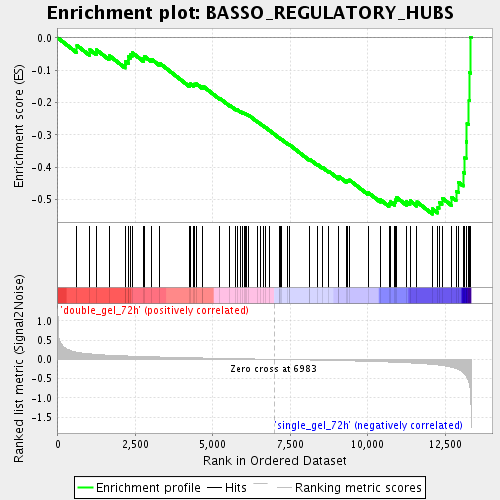
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | BASSO_REGULATORY_HUBS |
| Enrichment Score (ES) | -0.5448592 |
| Normalized Enrichment Score (NES) | -1.8266387 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.032408766 |
| FWER p-Value | 0.892 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 613 | 0.184 | -0.0231 | No |
| 2 | LGALS9 | LGALS9 Entrez, Source | lectin, galactoside-binding, soluble, 9 (galectin 9) | 1031 | 0.142 | -0.0368 | No |
| 3 | AFG3L2 | AFG3L2 Entrez, Source | AFG3 ATPase family gene 3-like 2 (yeast) | 1242 | 0.127 | -0.0367 | No |
| 4 | MRPL12 | MRPL12 Entrez, Source | mitochondrial ribosomal protein L12 | 1652 | 0.107 | -0.0542 | No |
| 5 | CASP4 | CASP4 Entrez, Source | caspase 4, apoptosis-related cysteine peptidase | 2168 | 0.089 | -0.0819 | No |
| 6 | ITPR1 | ITPR1 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 1 | 2186 | 0.089 | -0.0721 | No |
| 7 | EBNA1BP2 | EBNA1BP2 Entrez, Source | EBNA1 binding protein 2 | 2276 | 0.086 | -0.0681 | No |
| 8 | LGALS1 | LGALS1 Entrez, Source | lectin, galactoside-binding, soluble, 1 (galectin 1) | 2281 | 0.086 | -0.0576 | No |
| 9 | BYSL | BYSL Entrez, Source | bystin-like | 2347 | 0.084 | -0.0520 | No |
| 10 | PRKCB1 | PRKCB1 Entrez, Source | protein kinase C, beta 1 | 2401 | 0.082 | -0.0457 | No |
| 11 | NME1 | NME1 Entrez, Source | non-metastatic cells 1, protein (NM23A) expressed in | 2764 | 0.072 | -0.0640 | No |
| 12 | PLCB2 | PLCB2 Entrez, Source | phospholipase C, beta 2 | 2782 | 0.072 | -0.0563 | No |
| 13 | HSPE1 | HSPE1 Entrez, Source | heat shock 10kDa protein 1 (chaperonin 10) | 3020 | 0.066 | -0.0659 | No |
| 14 | TIMM17A | TIMM17A Entrez, Source | translocase of inner mitochondrial membrane 17 homolog A (yeast) | 3292 | 0.060 | -0.0788 | No |
| 15 | ASNS | ASNS Entrez, Source | asparagine synthetase | 4245 | 0.042 | -0.1453 | No |
| 16 | PRDX3 | PRDX3 Entrez, Source | peroxiredoxin 3 | 4271 | 0.042 | -0.1419 | No |
| 17 | GARS | GARS Entrez, Source | glycyl-tRNA synthetase | 4370 | 0.040 | -0.1443 | No |
| 18 | SSBP1 | SSBP1 Entrez, Source | single-stranded DNA binding protein 1 | 4411 | 0.039 | -0.1424 | No |
| 19 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 4470 | 0.038 | -0.1420 | No |
| 20 | LAPTM5 | LAPTM5 Entrez, Source | lysosomal associated multispanning membrane protein 5 | 4666 | 0.035 | -0.1524 | No |
| 21 | WDR18 | WDR18 Entrez, Source | WD repeat domain 18 | 4678 | 0.035 | -0.1489 | No |
| 22 | TOMM40 | TOMM40 Entrez, Source | translocase of outer mitochondrial membrane 40 homolog (yeast) | 5228 | 0.026 | -0.1870 | No |
| 23 | PSMD1 | PSMD1 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 1 | 5547 | 0.021 | -0.2084 | No |
| 24 | BUD31 | BUD31 Entrez, Source | BUD31 homolog (yeast) | 5741 | 0.018 | -0.2206 | No |
| 25 | POLR2F | POLR2F Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide F | 5780 | 0.018 | -0.2213 | No |
| 26 | PSMB5 | PSMB5 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 5 | 5908 | 0.016 | -0.2289 | No |
| 27 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 5968 | 0.015 | -0.2315 | No |
| 28 | PSMB7 | PSMB7 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 7 | 6017 | 0.014 | -0.2334 | No |
| 29 | PSMB2 | PSMB2 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 2 | 6045 | 0.014 | -0.2337 | No |
| 30 | POLD2 | POLD2 Entrez, Source | polymerase (DNA directed), delta 2, regulatory subunit 50kDa | 6095 | 0.013 | -0.2358 | No |
| 31 | EIF2S1 | EIF2S1 Entrez, Source | eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa | 6150 | 0.012 | -0.2384 | No |
| 32 | RPS26 | RPS26 Entrez, Source | ribosomal protein S26 | 6430 | 0.007 | -0.2585 | No |
| 33 | EPOR | EPOR Entrez, Source | erythropoietin receptor | 6541 | 0.006 | -0.2661 | No |
| 34 | SAR1A | SAR1A Entrez, Source | SAR1 gene homolog A (S. cerevisiae) | 6643 | 0.005 | -0.2731 | No |
| 35 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 6707 | 0.004 | -0.2774 | No |
| 36 | TGOLN2 | TGOLN2 Entrez, Source | trans-golgi network protein 2 | 6822 | 0.002 | -0.2857 | No |
| 37 | TCEB1 | TCEB1 Entrez, Source | transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) | 7156 | -0.002 | -0.3105 | No |
| 38 | ITPR2 | ITPR2 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 2 | 7164 | -0.003 | -0.3107 | No |
| 39 | EIF4G1 | EIF4G1 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 1 | 7196 | -0.003 | -0.3126 | No |
| 40 | TEP1 | TEP1 Entrez, Source | telomerase-associated protein 1 | 7217 | -0.003 | -0.3137 | No |
| 41 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 7409 | -0.006 | -0.3274 | No |
| 42 | GABARAP | GABARAP Entrez, Source | GABA(A) receptor-associated protein | 7459 | -0.007 | -0.3302 | No |
| 43 | HNRPAB | HNRPAB Entrez, Source | heterogeneous nuclear ribonucleoprotein A/B | 7472 | -0.007 | -0.3302 | No |
| 44 | SERBP1 | SERBP1 Entrez, Source | SERPINE1 mRNA binding protein 1 | 8112 | -0.016 | -0.3763 | No |
| 45 | BLMH | BLMH Entrez, Source | bleomycin hydrolase | 8135 | -0.017 | -0.3759 | No |
| 46 | PAICS | PAICS Entrez, Source | phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoimidazole succinocarboxamide synthetase | 8385 | -0.021 | -0.3920 | No |
| 47 | PSMD2 | PSMD2 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 2 | 8545 | -0.024 | -0.4010 | No |
| 48 | SSRP1 | SSRP1 Entrez, Source | structure specific recognition protein 1 | 8733 | -0.027 | -0.4117 | No |
| 49 | ANXA6 | ANXA6 Entrez, Source | annexin A6 | 9062 | -0.033 | -0.4323 | No |
| 50 | TCP1 | TCP1 Entrez, Source | t-complex 1 | 9066 | -0.033 | -0.4284 | No |
| 51 | MVP | MVP Entrez, Source | major vault protein | 9306 | -0.038 | -0.4416 | No |
| 52 | TUBG1 | TUBG1 Entrez, Source | tubulin, gamma 1 | 9358 | -0.039 | -0.4406 | No |
| 53 | SLC4A3 | SLC4A3 Entrez, Source | solute carrier family 4, anion exchanger, member 3 | 9406 | -0.040 | -0.4392 | No |
| 54 | PRDX4 | PRDX4 Entrez, Source | peroxiredoxin 4 | 10020 | -0.053 | -0.4788 | No |
| 55 | HHEX | HHEX Entrez, Source | homeobox, hematopoietically expressed | 10405 | -0.062 | -0.5001 | No |
| 56 | CTSH | CTSH Entrez, Source | cathepsin H | 10687 | -0.069 | -0.5126 | No |
| 57 | HPRT1 | HPRT1 Entrez, Source | hypoxanthine phosphoribosyltransferase 1 (Lesch-Nyhan syndrome) | 10730 | -0.071 | -0.5069 | No |
| 58 | LITAF | LITAF Entrez, Source | lipopolysaccharide-induced TNF factor | 10871 | -0.074 | -0.5081 | No |
| 59 | ZMYND11 | ZMYND11 Entrez, Source | zinc finger, MYND domain containing 11 | 10901 | -0.075 | -0.5009 | No |
| 60 | RBM5 | RBM5 Entrez, Source | RNA binding motif protein 5 | 10933 | -0.076 | -0.4937 | No |
| 61 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 11249 | -0.088 | -0.5065 | No |
| 62 | MGEA5 | MGEA5 Entrez, Source | meningioma expressed antigen 5 (hyaluronidase) | 11364 | -0.093 | -0.5034 | No |
| 63 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 11588 | -0.104 | -0.5072 | No |
| 64 | UCHL1 | UCHL1 Entrez, Source | ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) | 12088 | -0.130 | -0.5285 | Yes |
| 65 | TPP1 | TPP1 Entrez, Source | tripeptidyl peptidase I | 12262 | -0.145 | -0.5235 | Yes |
| 66 | NISCH | NISCH Entrez, Source | nischarin | 12301 | -0.148 | -0.5077 | Yes |
| 67 | BIN1 | BIN1 Entrez, Source | bridging integrator 1 | 12397 | -0.157 | -0.4952 | Yes |
| 68 | SLC25A4 | SLC25A4 Entrez, Source | solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 4 | 12710 | -0.207 | -0.4929 | Yes |
| 69 | TRIP13 | TRIP13 Entrez, Source | thyroid hormone receptor interactor 13 | 12851 | -0.238 | -0.4737 | Yes |
| 70 | ZYX | ZYX Entrez, Source | zyxin | 12928 | -0.262 | -0.4467 | Yes |
| 71 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 13080 | -0.340 | -0.4155 | Yes |
| 72 | CCNB1 | CCNB1 Entrez, Source | cyclin B1 | 13131 | -0.385 | -0.3711 | Yes |
| 73 | CD44 | CD44 Entrez, Source | CD44 molecule (Indian blood group) | 13174 | -0.431 | -0.3203 | Yes |
| 74 | S100A4 | S100A4 Entrez, Source | S100 calcium binding protein A4 | 13202 | -0.468 | -0.2639 | Yes |
| 75 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 13266 | -0.607 | -0.1927 | Yes |
| 76 | AHNAK | AHNAK Entrez, Source | AHNAK nucleoprotein (desmoyokin) | 13286 | -0.695 | -0.1071 | Yes |
| 77 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13314 | -0.890 | 0.0021 | Yes |

