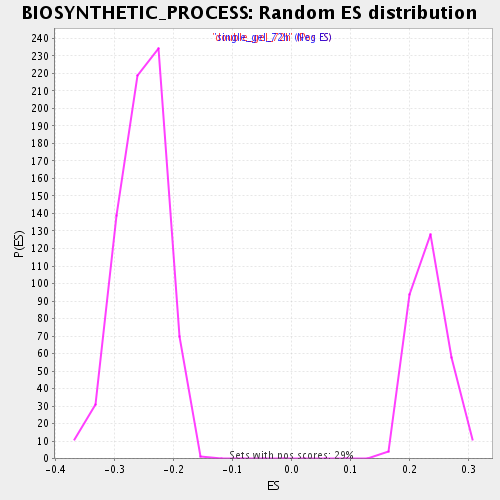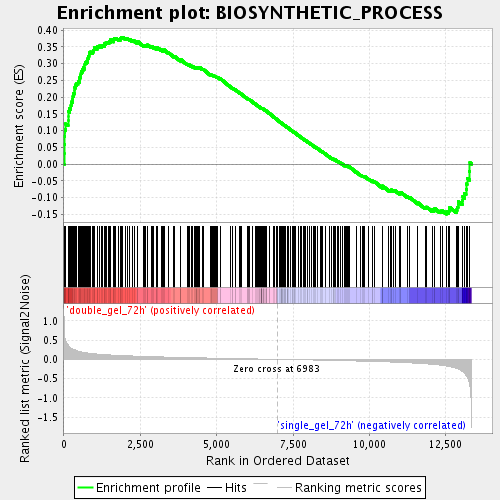
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | BIOSYNTHETIC_PROCESS |
| Enrichment Score (ES) | 0.37843755 |
| Normalized Enrichment Score (NES) | 1.6336643 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.059282463 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PEMT | PEMT Entrez, Source | phosphatidylethanolamine N-methyltransferase | 8 | 0.798 | 0.0308 | Yes |
| 2 | BBOX1 | BBOX1 Entrez, Source | butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 | 13 | 0.740 | 0.0596 | Yes |
| 3 | APOA2 | APOA2 Entrez, Source | apolipoprotein A-II | 22 | 0.601 | 0.0826 | Yes |
| 4 | LTB | LTB Entrez, Source | lymphotoxin beta (TNF superfamily, member 3) | 35 | 0.539 | 0.1029 | Yes |
| 5 | CDO1 | CDO1 Entrez, Source | cysteine dioxygenase, type I | 58 | 0.484 | 0.1202 | Yes |
| 6 | IDI1 | IDI1 Entrez, Source | isopentenyl-diphosphate delta isomerase 1 | 136 | 0.372 | 0.1290 | Yes |
| 7 | GCK | GCK Entrez, Source | glucokinase (hexokinase 4, maturity onset diabetes of the young 2) | 140 | 0.368 | 0.1432 | Yes |
| 8 | LCAT | LCAT Entrez, Source | lecithin-cholesterol acyltransferase | 142 | 0.365 | 0.1575 | Yes |
| 9 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 182 | 0.324 | 0.1672 | Yes |
| 10 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 230 | 0.293 | 0.1752 | Yes |
| 11 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 243 | 0.289 | 0.1856 | Yes |
| 12 | ACN9 | ACN9 Entrez, Source | ACN9 homolog (S. cerevisiae) | 284 | 0.273 | 0.1933 | Yes |
| 13 | GAMT | GAMT Entrez, Source | guanidinoacetate N-methyltransferase | 295 | 0.267 | 0.2030 | Yes |
| 14 | GYS2 | GYS2 Entrez, Source | glycogen synthase 2 (liver) | 314 | 0.263 | 0.2120 | Yes |
| 15 | MOCS2 | MOCS2 Entrez, Source | molybdenum cofactor synthesis 2 | 352 | 0.248 | 0.2189 | Yes |
| 16 | ABCA1 | ABCA1 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 1 | 355 | 0.248 | 0.2285 | Yes |
| 17 | SLC25A10 | SLC25A10 Entrez, Source | solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 | 378 | 0.240 | 0.2362 | Yes |
| 18 | ALAS1 | ALAS1 Entrez, Source | aminolevulinate, delta-, synthase 1 | 419 | 0.224 | 0.2420 | Yes |
| 19 | FDPS | FDPS Entrez, Source | farnesyl diphosphate synthase (farnesyl pyrophosphate synthetase, dimethylallyltranstransferase, geranyltranstransferase) | 462 | 0.212 | 0.2472 | Yes |
| 20 | DGAT2 | DGAT2 Entrez, Source | diacylglycerol O-acyltransferase homolog 2 (mouse) | 503 | 0.201 | 0.2520 | Yes |
| 21 | HRSP12 | HRSP12 Entrez, Source | heat-responsive protein 12 | 519 | 0.198 | 0.2587 | Yes |
| 22 | SIGIRR | SIGIRR Entrez, Source | single immunoglobulin and toll-interleukin 1 receptor (TIR) domain | 544 | 0.194 | 0.2644 | Yes |
| 23 | GCHFR | GCHFR Entrez, Source | GTP cyclohydrolase I feedback regulator | 558 | 0.191 | 0.2710 | Yes |
| 24 | MVK | MVK Entrez, Source | mevalonate kinase (mevalonic aciduria) | 578 | 0.188 | 0.2769 | Yes |
| 25 | ST8SIA4 | ST8SIA4 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 4 | 611 | 0.184 | 0.2817 | Yes |
| 26 | PAH | PAH Entrez, Source | phenylalanine hydroxylase | 628 | 0.182 | 0.2877 | Yes |
| 27 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 685 | 0.175 | 0.2903 | Yes |
| 28 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 686 | 0.175 | 0.2971 | Yes |
| 29 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 693 | 0.174 | 0.3035 | Yes |
| 30 | PCYT2 | PCYT2 Entrez, Source | phosphate cytidylyltransferase 2, ethanolamine | 748 | 0.168 | 0.3060 | Yes |
| 31 | CHPT1 | CHPT1 Entrez, Source | choline phosphotransferase 1 | 778 | 0.165 | 0.3102 | Yes |
| 32 | CD37 | CD37 Entrez, Source | CD37 molecule | 784 | 0.164 | 0.3163 | Yes |
| 33 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 800 | 0.163 | 0.3215 | Yes |
| 34 | GCLC | GCLC Entrez, Source | glutamate-cysteine ligase, catalytic subunit | 823 | 0.160 | 0.3262 | Yes |
| 35 | ALAD | ALAD Entrez, Source | aminolevulinate, delta-, dehydratase | 826 | 0.160 | 0.3323 | Yes |
| 36 | EIF1 | EIF1 Entrez, Source | eukaryotic translation initiation factor 1 | 876 | 0.155 | 0.3347 | Yes |
| 37 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 951 | 0.149 | 0.3349 | Yes |
| 38 | RPL39 | RPL39 Entrez, Source | ribosomal protein L39 | 960 | 0.148 | 0.3401 | Yes |
| 39 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 984 | 0.146 | 0.3441 | Yes |
| 40 | DHPS | DHPS Entrez, Source | deoxyhypusine synthase | 996 | 0.145 | 0.3489 | Yes |
| 41 | SERP1 | SERP1 Entrez, Source | - | 1085 | 0.138 | 0.3476 | Yes |
| 42 | UCN2 | UCN2 Entrez, Source | urocortin 2 | 1097 | 0.137 | 0.3522 | Yes |
| 43 | LDLR | LDLR Entrez, Source | low density lipoprotein receptor (familial hypercholesterolemia) | 1160 | 0.133 | 0.3527 | Yes |
| 44 | ST8SIA1 | ST8SIA1 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 1 | 1223 | 0.128 | 0.3530 | Yes |
| 45 | INHA | INHA Entrez, Source | inhibin, alpha | 1270 | 0.126 | 0.3544 | Yes |
| 46 | PI4K2A | 1329 | 0.123 | 0.3548 | Yes | ||
| 47 | GALNT5 | GALNT5 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 5 (GalNAc-T5) | 1333 | 0.122 | 0.3594 | Yes |
| 48 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 1370 | 0.121 | 0.3613 | Yes |
| 49 | COQ3 | COQ3 Entrez, Source | coenzyme Q3 homolog, methyltransferase (S. cerevisiae) | 1406 | 0.118 | 0.3633 | Yes |
| 50 | GCS1 | GCS1 Entrez, Source | - | 1467 | 0.115 | 0.3633 | Yes |
| 51 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 1495 | 0.114 | 0.3657 | Yes |
| 52 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 1524 | 0.112 | 0.3679 | Yes |
| 53 | NQO1 | NQO1 Entrez, Source | NAD(P)H dehydrogenase, quinone 1 | 1532 | 0.112 | 0.3718 | Yes |
| 54 | SCP2 | SCP2 Entrez, Source | sterol carrier protein 2 | 1632 | 0.107 | 0.3684 | Yes |
| 55 | FUT1 | FUT1 Entrez, Source | fucosyltransferase 1 (galactoside 2-alpha-L-fucosyltransferase, H blood group) | 1637 | 0.107 | 0.3723 | Yes |
| 56 | NANP | NANP Entrez, Source | N-acetylneuraminic acid phosphatase | 1648 | 0.107 | 0.3758 | Yes |
| 57 | FUT8 | FUT8 Entrez, Source | fucosyltransferase 8 (alpha (1,6) fucosyltransferase) | 1701 | 0.105 | 0.3759 | Yes |
| 58 | MTIF2 | MTIF2 Entrez, Source | mitochondrial translational initiation factor 2 | 1797 | 0.101 | 0.3726 | Yes |
| 59 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 1840 | 0.100 | 0.3733 | Yes |
| 60 | AKR1D1 | AKR1D1 Entrez, Source | aldo-keto reductase family 1, member D1 (delta 4-3-ketosteroid-5-beta-reductase) | 1859 | 0.099 | 0.3759 | Yes |
| 61 | BMP6 | BMP6 Entrez, Source | bone morphogenetic protein 6 | 1877 | 0.098 | 0.3784 | Yes |
| 62 | MRPS7 | MRPS7 Entrez, Source | mitochondrial ribosomal protein S7 | 1930 | 0.096 | 0.3782 | No |
| 63 | COQ7 | COQ7 Entrez, Source | coenzyme Q7 homolog, ubiquinone (yeast) | 2016 | 0.093 | 0.3754 | No |
| 64 | ADK | ADK Entrez, Source | adenosine kinase | 2082 | 0.091 | 0.3740 | No |
| 65 | COG3 | COG3 Entrez, Source | component of oligomeric golgi complex 3 | 2154 | 0.090 | 0.3721 | No |
| 66 | AIPL1 | AIPL1 Entrez, Source | aryl hydrocarbon receptor interacting protein-like 1 | 2256 | 0.086 | 0.3678 | No |
| 67 | EIF2B1 | EIF2B1 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 1 alpha, 26kDa | 2303 | 0.085 | 0.3676 | No |
| 68 | PTS | PTS Entrez, Source | 6-pyruvoyltetrahydropterin synthase | 2407 | 0.082 | 0.3630 | No |
| 69 | ABCF1 | ABCF1 Entrez, Source | ATP-binding cassette, sub-family F (GCN20), member 1 | 2412 | 0.082 | 0.3659 | No |
| 70 | PPAT | PPAT Entrez, Source | phosphoribosyl pyrophosphate amidotransferase | 2613 | 0.076 | 0.3536 | No |
| 71 | NDST1 | NDST1 Entrez, Source | N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 1 | 2645 | 0.075 | 0.3542 | No |
| 72 | ST6GALNAC6 | ST6GALNAC6 Entrez, Source | ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 6 | 2672 | 0.074 | 0.3551 | No |
| 73 | CHST12 | CHST12 Entrez, Source | carbohydrate (chondroitin 4) sulfotransferase 12 | 2721 | 0.073 | 0.3543 | No |
| 74 | SPN | SPN Entrez, Source | sialophorin (leukosialin, CD43) | 2741 | 0.073 | 0.3557 | No |
| 75 | COASY | COASY Entrez, Source | Coenzyme A synthase | 2861 | 0.070 | 0.3494 | No |
| 76 | SFTPD | SFTPD Entrez, Source | surfactant, pulmonary-associated protein D | 2883 | 0.069 | 0.3505 | No |
| 77 | CPOX | CPOX Entrez, Source | coproporphyrinogen oxidase | 2946 | 0.068 | 0.3484 | No |
| 78 | PEX7 | PEX7 Entrez, Source | peroxisomal biogenesis factor 7 | 3029 | 0.066 | 0.3447 | No |
| 79 | UMPS | UMPS Entrez, Source | uridine monophosphate synthetase (orotate phosphoribosyl transferase and orotidine-5'-decarboxylase) | 3035 | 0.066 | 0.3469 | No |
| 80 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 3058 | 0.065 | 0.3478 | No |
| 81 | MPDU1 | MPDU1 Entrez, Source | mannose-P-dolichol utilization defect 1 | 3177 | 0.063 | 0.3413 | No |
| 82 | CHST3 | CHST3 Entrez, Source | carbohydrate (chondroitin 6) sulfotransferase 3 | 3239 | 0.061 | 0.3390 | No |
| 83 | B3GALT4 | B3GALT4 Entrez, Source | UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 4 | 3273 | 0.061 | 0.3389 | No |
| 84 | EIF2B3 | EIF2B3 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 3 gamma, 58kDa | 3277 | 0.061 | 0.3410 | No |
| 85 | MGAT4A | MGAT4A Entrez, Source | mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme A | 3423 | 0.058 | 0.3322 | No |
| 86 | GMPS | GMPS Entrez, Source | guanine monphosphate synthetase | 3595 | 0.054 | 0.3212 | No |
| 87 | ST8SIA5 | ST8SIA5 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 5 | 3625 | 0.053 | 0.3211 | No |
| 88 | FUT4 | FUT4 Entrez, Source | fucosyltransferase 4 (alpha (1,3) fucosyltransferase, myeloid-specific) | 3809 | 0.050 | 0.3091 | No |
| 89 | PIGK | PIGK Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class K | 3814 | 0.050 | 0.3107 | No |
| 90 | UROS | UROS Entrez, Source | uroporphyrinogen III synthase (congenital erythropoietic porphyria) | 3820 | 0.049 | 0.3123 | No |
| 91 | AGPAT1 | AGPAT1 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) | 4028 | 0.046 | 0.2983 | No |
| 92 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 4079 | 0.045 | 0.2962 | No |
| 93 | RPN1 | RPN1 Entrez, Source | ribophorin I | 4101 | 0.045 | 0.2963 | No |
| 94 | APOA1 | APOA1 Entrez, Source | apolipoprotein A-I | 4169 | 0.043 | 0.2929 | No |
| 95 | CYP27B1 | CYP27B1 Entrez, Source | cytochrome P450, family 27, subfamily B, polypeptide 1 | 4218 | 0.043 | 0.2909 | No |
| 96 | AARS | AARS Entrez, Source | alanyl-tRNA synthetase | 4275 | 0.042 | 0.2883 | No |
| 97 | TLR4 | TLR4 Entrez, Source | toll-like receptor 4 | 4308 | 0.041 | 0.2874 | No |
| 98 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 4326 | 0.041 | 0.2877 | No |
| 99 | PIGC | PIGC Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class C | 4338 | 0.040 | 0.2885 | No |
| 100 | MRPL23 | MRPL23 Entrez, Source | mitochondrial ribosomal protein L23 | 4352 | 0.040 | 0.2891 | No |
| 101 | IL10 | IL10 Entrez, Source | interleukin 10 | 4386 | 0.040 | 0.2881 | No |
| 102 | ETF1 | ETF1 Entrez, Source | eukaryotic translation termination factor 1 | 4420 | 0.039 | 0.2871 | No |
| 103 | ADORA2A | ADORA2A Entrez, Source | adenosine A2a receptor | 4439 | 0.039 | 0.2873 | No |
| 104 | FUT9 | FUT9 Entrez, Source | fucosyltransferase 9 (alpha (1,3) fucosyltransferase) | 4443 | 0.039 | 0.2886 | No |
| 105 | HSD11B2 | HSD11B2 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 2 | 4542 | 0.037 | 0.2825 | No |
| 106 | EIF2B4 | EIF2B4 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 4 delta, 67kDa | 4558 | 0.037 | 0.2829 | No |
| 107 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 4803 | 0.032 | 0.2655 | No |
| 108 | RPL35 | RPL35 Entrez, Source | ribosomal protein L35 | 4821 | 0.032 | 0.2654 | No |
| 109 | EIF2B5 | EIF2B5 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 5 epsilon, 82kDa | 4827 | 0.032 | 0.2663 | No |
| 110 | DHRS9 | DHRS9 Entrez, Source | dehydrogenase/reductase (SDR family) member 9 | 4865 | 0.031 | 0.2647 | No |
| 111 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 4887 | 0.031 | 0.2643 | No |
| 112 | MRPL41 | MRPL41 Entrez, Source | mitochondrial ribosomal protein L41 | 4921 | 0.030 | 0.2630 | No |
| 113 | TSC1 | TSC1 Entrez, Source | tuberous sclerosis 1 | 4962 | 0.030 | 0.2611 | No |
| 114 | HSD3B1 | HSD3B1 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 | 4997 | 0.029 | 0.2597 | No |
| 115 | ARPP-19 | ARPP-19 Entrez, Source | - | 5020 | 0.029 | 0.2591 | No |
| 116 | COPS5 | COPS5 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 5 (Arabidopsis) | 5028 | 0.029 | 0.2597 | No |
| 117 | POMT1 | POMT1 Entrez, Source | protein-O-mannosyltransferase 1 | 5110 | 0.028 | 0.2546 | No |
| 118 | RPL21 | RPL21 Entrez, Source | ribosomal protein L21 | 5125 | 0.027 | 0.2546 | No |
| 119 | CNBP | CNBP Entrez, Source | CCHC-type zinc finger, nucleic acid binding protein | 5438 | 0.023 | 0.2316 | No |
| 120 | PGGT1B | PGGT1B Entrez, Source | protein geranylgeranyltransferase type I, beta subunit | 5519 | 0.021 | 0.2263 | No |
| 121 | GRM8 | GRM8 Entrez, Source | glutamate receptor, metabotropic 8 | 5525 | 0.021 | 0.2268 | No |
| 122 | ALAS2 | ALAS2 Entrez, Source | aminolevulinate, delta-, synthase 2 (sideroblastic/hypochromic anemia) | 5606 | 0.020 | 0.2215 | No |
| 123 | PTPRC | PTPRC Entrez, Source | protein tyrosine phosphatase, receptor type, C | 5608 | 0.020 | 0.2222 | No |
| 124 | HBS1L | HBS1L Entrez, Source | HBS1-like (S. cerevisiae) | 5747 | 0.018 | 0.2124 | No |
| 125 | RPL4 | RPL4 Entrez, Source | ribosomal protein L4 | 5769 | 0.018 | 0.2115 | No |
| 126 | RPL13 | RPL13 Entrez, Source | ribosomal protein L13 | 5821 | 0.017 | 0.2082 | No |
| 127 | APBB1 | APBB1 Entrez, Source | amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65) | 6000 | 0.014 | 0.1952 | No |
| 128 | IL12B | IL12B Entrez, Source | interleukin 12B (natural killer cell stimulatory factor 2, cytotoxic lymphocyte maturation factor 2, p40) | 6003 | 0.014 | 0.1956 | No |
| 129 | GALNT1 | GALNT1 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 1 (GalNAc-T1) | 6034 | 0.014 | 0.1938 | No |
| 130 | SCYE1 | SCYE1 Entrez, Source | small inducible cytokine subfamily E, member 1 (endothelial monocyte-activating) | 6040 | 0.014 | 0.1940 | No |
| 131 | RPL41 | RPL41 Entrez, Source | ribosomal protein L41 | 6066 | 0.013 | 0.1926 | No |
| 132 | AGPS | AGPS Entrez, Source | alkylglycerone phosphate synthase | 6079 | 0.013 | 0.1922 | No |
| 133 | PCYT1B | PCYT1B Entrez, Source | phosphate cytidylyltransferase 1, choline, beta | 6186 | 0.012 | 0.1845 | No |
| 134 | RPL14 | RPL14 Entrez, Source | ribosomal protein L14 | 6253 | 0.010 | 0.1799 | No |
| 135 | EIF5 | EIF5 Entrez, Source | eukaryotic translation initiation factor 5 | 6311 | 0.009 | 0.1759 | No |
| 136 | RPS3A | RPS3A Entrez, Source | ribosomal protein S3A | 6320 | 0.009 | 0.1756 | No |
| 137 | RPL24 | RPL24 Entrez, Source | ribosomal protein L24 | 6383 | 0.008 | 0.1712 | No |
| 138 | FDXR | FDXR Entrez, Source | ferredoxin reductase | 6391 | 0.008 | 0.1710 | No |
| 139 | RPL37 | RPL37 Entrez, Source | ribosomal protein L37 | 6404 | 0.008 | 0.1704 | No |
| 140 | RPS9 | RPS9 Entrez, Source | ribosomal protein S9 | 6448 | 0.007 | 0.1674 | No |
| 141 | ST8SIA3 | ST8SIA3 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 3 | 6457 | 0.007 | 0.1670 | No |
| 142 | IL4 | IL4 Entrez, Source | interleukin 4 | 6466 | 0.007 | 0.1667 | No |
| 143 | RPS12 | RPS12 Entrez, Source | ribosomal protein S12 | 6467 | 0.007 | 0.1670 | No |
| 144 | BCL10 | BCL10 Entrez, Source | B-cell CLL/lymphoma 10 | 6471 | 0.007 | 0.1670 | No |
| 145 | DARS | DARS Entrez, Source | aspartyl-tRNA synthetase | 6478 | 0.007 | 0.1668 | No |
| 146 | RPL29 | RPL29 Entrez, Source | ribosomal protein L29 | 6480 | 0.007 | 0.1670 | No |
| 147 | RPS17 | RPS17 Entrez, Source | ribosomal protein S17 | 6495 | 0.006 | 0.1662 | No |
| 148 | ASMT | ASMT Entrez, Source | acetylserotonin O-methyltransferase | 6518 | 0.006 | 0.1647 | No |
| 149 | RPL18 | RPL18 Entrez, Source | ribosomal protein L18 | 6533 | 0.006 | 0.1639 | No |
| 150 | RPS3 | RPS3 Entrez, Source | ribosomal protein S3 | 6546 | 0.006 | 0.1632 | No |
| 151 | RPL28 | RPL28 Entrez, Source | ribosomal protein L28 | 6566 | 0.006 | 0.1620 | No |
| 152 | CYP19A1 | CYP19A1 Entrez, Source | cytochrome P450, family 19, subfamily A, polypeptide 1 | 6596 | 0.005 | 0.1600 | No |
| 153 | RPL7 | RPL7 Entrez, Source | ribosomal protein L7 | 6616 | 0.005 | 0.1587 | No |
| 154 | RPL31 | RPL31 Entrez, Source | ribosomal protein L31 | 6641 | 0.005 | 0.1571 | No |
| 155 | TRSPAP1 | TRSPAP1 Entrez, Source | tRNA selenocysteine associated protein 1 | 6644 | 0.005 | 0.1571 | No |
| 156 | GUK1 | GUK1 Entrez, Source | guanylate kinase 1 | 6715 | 0.004 | 0.1519 | No |
| 157 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6738 | 0.003 | 0.1503 | No |
| 158 | RPS11 | RPS11 Entrez, Source | ribosomal protein S11 | 6874 | 0.002 | 0.1401 | No |
| 159 | ALG8 | ALG8 Entrez, Source | asparagine-linked glycosylation 8 homolog (S. cerevisiae, alpha-1,3-glucosyltransferase) | 6902 | 0.001 | 0.1381 | No |
| 160 | RPL3 | RPL3 Entrez, Source | ribosomal protein L3 | 6956 | 0.000 | 0.1340 | No |
| 161 | ALG2 | ALG2 Entrez, Source | asparagine-linked glycosylation 2 homolog (S. cerevisiae, alpha-1,3-mannosyltransferase) | 6970 | 0.000 | 0.1330 | No |
| 162 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 6984 | -0.000 | 0.1320 | No |
| 163 | RPL30 | RPL30 Entrez, Source | ribosomal protein L30 | 6985 | -0.000 | 0.1320 | No |
| 164 | RPL8 | RPL8 Entrez, Source | ribosomal protein L8 | 7041 | -0.001 | 0.1279 | No |
| 165 | RPS10 | RPS10 Entrez, Source | ribosomal protein S10 | 7076 | -0.001 | 0.1253 | No |
| 166 | RPL26 | RPL26 Entrez, Source | ribosomal protein L26 | 7117 | -0.002 | 0.1223 | No |
| 167 | TNFRSF8 | TNFRSF8 Entrez, Source | tumor necrosis factor receptor superfamily, member 8 | 7122 | -0.002 | 0.1221 | No |
| 168 | RPS2 | RPS2 Entrez, Source | ribosomal protein S2 | 7135 | -0.002 | 0.1213 | No |
| 169 | PABPC4 | PABPC4 Entrez, Source | poly(A) binding protein, cytoplasmic 4 (inducible form) | 7148 | -0.002 | 0.1205 | No |
| 170 | RPL9 | RPL9 Entrez, Source | ribosomal protein L9 | 7163 | -0.003 | 0.1195 | No |
| 171 | B4GALT7 | B4GALT7 Entrez, Source | xylosylprotein beta 1,4-galactosyltransferase, polypeptide 7 (galactosyltransferase I) | 7186 | -0.003 | 0.1179 | No |
| 172 | GLMN | GLMN Entrez, Source | glomulin, FKBP associated protein | 7210 | -0.003 | 0.1163 | No |
| 173 | RPL6 | RPL6 Entrez, Source | ribosomal protein L6 | 7233 | -0.003 | 0.1147 | No |
| 174 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 7239 | -0.004 | 0.1145 | No |
| 175 | KARS | KARS Entrez, Source | lysyl-tRNA synthetase | 7266 | -0.004 | 0.1127 | No |
| 176 | EEF1A1 | EEF1A1 Entrez, Source | eukaryotic translation elongation factor 1 alpha 1 | 7318 | -0.005 | 0.1090 | No |
| 177 | RPS27 | RPS27 Entrez, Source | ribosomal protein S27 (metallopanstimulin 1) | 7325 | -0.005 | 0.1087 | No |
| 178 | FNTB | FNTB Entrez, Source | farnesyltransferase, CAAX box, beta | 7331 | -0.005 | 0.1085 | No |
| 179 | RPL15 | RPL15 Entrez, Source | ribosomal protein L15 | 7353 | -0.005 | 0.1071 | No |
| 180 | RPS5 | RPS5 Entrez, Source | ribosomal protein S5 | 7405 | -0.006 | 0.1035 | No |
| 181 | ST8SIA2 | ST8SIA2 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 2 | 7470 | -0.007 | 0.0988 | No |
| 182 | DEFB103A | DEFB103A Entrez, Source | defensin, beta 103A | 7483 | -0.007 | 0.0982 | No |
| 183 | RPL5 | RPL5 Entrez, Source | ribosomal protein L5 | 7524 | -0.008 | 0.0954 | No |
| 184 | RPL22 | RPL22 Entrez, Source | ribosomal protein L22 | 7541 | -0.008 | 0.0945 | No |
| 185 | POFUT1 | POFUT1 Entrez, Source | protein O-fucosyltransferase 1 | 7549 | -0.008 | 0.0943 | No |
| 186 | RPS19 | RPS19 Entrez, Source | ribosomal protein S19 | 7575 | -0.009 | 0.0928 | No |
| 187 | B3GAT2 | B3GAT2 Entrez, Source | beta-1,3-glucuronyltransferase 2 (glucuronosyltransferase S) | 7679 | -0.010 | 0.0853 | No |
| 188 | CMPK | CMPK Entrez, Source | cytidylate kinase | 7736 | -0.011 | 0.0814 | No |
| 189 | CD28 | CD28 Entrez, Source | CD28 molecule | 7763 | -0.011 | 0.0799 | No |
| 190 | DDX25 | DDX25 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 25 | 7853 | -0.013 | 0.0736 | No |
| 191 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 7883 | -0.013 | 0.0719 | No |
| 192 | PIGV | PIGV Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class V | 7912 | -0.014 | 0.0703 | No |
| 193 | RPL19 | RPL19 Entrez, Source | ribosomal protein L19 | 7963 | -0.014 | 0.0670 | No |
| 194 | GHRL | GHRL Entrez, Source | ghrelin/obestatin preprohormone | 7982 | -0.015 | 0.0662 | No |
| 195 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 8036 | -0.015 | 0.0628 | No |
| 196 | HSD3B7 | HSD3B7 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 | 8089 | -0.016 | 0.0595 | No |
| 197 | LRP2 | LRP2 Entrez, Source | low density lipoprotein-related protein 2 | 8183 | -0.018 | 0.0530 | No |
| 198 | COG7 | COG7 Entrez, Source | component of oligomeric golgi complex 7 | 8215 | -0.018 | 0.0514 | No |
| 199 | RPL11 | RPL11 Entrez, Source | ribosomal protein L11 | 8243 | -0.019 | 0.0501 | No |
| 200 | LALBA | LALBA Entrez, Source | lactalbumin, alpha- | 8293 | -0.020 | 0.0471 | No |
| 201 | PAICS | PAICS Entrez, Source | phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoimidazole succinocarboxamide synthetase | 8385 | -0.021 | 0.0410 | No |
| 202 | EIF5A | EIF5A Entrez, Source | eukaryotic translation initiation factor 5A | 8430 | -0.022 | 0.0385 | No |
| 203 | DPM2 | DPM2 Entrez, Source | dolichyl-phosphate mannosyltransferase polypeptide 2, regulatory subunit | 8478 | -0.023 | 0.0358 | No |
| 204 | EIF4B | EIF4B Entrez, Source | eukaryotic translation initiation factor 4B | 8550 | -0.024 | 0.0313 | No |
| 205 | OGT | OGT Entrez, Source | O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) | 8678 | -0.026 | 0.0226 | No |
| 206 | EIF4A2 | EIF4A2 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 2 | 8760 | -0.027 | 0.0175 | No |
| 207 | POMGNT1 | POMGNT1 Entrez, Source | protein O-linked mannose beta1,2-N-acetylglucosaminyltransferase | 8813 | -0.028 | 0.0146 | No |
| 208 | EIF2AK1 | EIF2AK1 Entrez, Source | eukaryotic translation initiation factor 2-alpha kinase 1 | 8817 | -0.028 | 0.0155 | No |
| 209 | ALG5 | ALG5 Entrez, Source | asparagine-linked glycosylation 5 homolog (S. cerevisiae, dolichyl-phosphate beta-glucosyltransferase) | 8862 | -0.029 | 0.0133 | No |
| 210 | GHSR | GHSR Entrez, Source | growth hormone secretagogue receptor | 8872 | -0.029 | 0.0137 | No |
| 211 | EIF4G2 | EIF4G2 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 2 | 8954 | -0.031 | 0.0088 | No |
| 212 | XYLT1 | XYLT1 Entrez, Source | xylosyltransferase I | 8993 | -0.032 | 0.0071 | No |
| 213 | FUT2 | FUT2 Entrez, Source | fucosyltransferase 2 (secretor status included) | 9061 | -0.033 | 0.0033 | No |
| 214 | CYP7B1 | CYP7B1 Entrez, Source | cytochrome P450, family 7, subfamily B, polypeptide 1 | 9114 | -0.034 | 0.0007 | No |
| 215 | OAZ1 | OAZ1 Entrez, Source | ornithine decarboxylase antizyme 1 | 9198 | -0.036 | -0.0043 | No |
| 216 | FARS2 | FARS2 Entrez, Source | phenylalanine-tRNA synthetase 2 (mitochondrial) | 9209 | -0.036 | -0.0036 | No |
| 217 | PIGS | PIGS Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class S | 9263 | -0.037 | -0.0062 | No |
| 218 | SPR | SPR Entrez, Source | sepiapterin reductase (7,8-dihydrobiopterin:NADP+ oxidoreductase) | 9268 | -0.037 | -0.0051 | No |
| 219 | MIF | MIF Entrez, Source | macrophage migration inhibitory factor (glycosylation-inhibiting factor) | 9316 | -0.038 | -0.0072 | No |
| 220 | EREG | EREG Entrez, Source | epiregulin | 9343 | -0.039 | -0.0076 | No |
| 221 | HSP90AB1 | HSP90AB1 Entrez, Source | heat shock protein 90kDa alpha (cytosolic), class B member 1 | 9570 | -0.043 | -0.0232 | No |
| 222 | GALNT7 | GALNT7 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 7 (GalNAc-T7) | 9717 | -0.046 | -0.0326 | No |
| 223 | NR0B1 | NR0B1 Entrez, Source | nuclear receptor subfamily 0, group B, member 1 | 9783 | -0.047 | -0.0357 | No |
| 224 | IMPA1 | IMPA1 Entrez, Source | inositol(myo)-1(or 4)-monophosphatase 1 | 9804 | -0.048 | -0.0353 | No |
| 225 | GMDS | GMDS Entrez, Source | GDP-mannose 4,6-dehydratase | 9847 | -0.049 | -0.0366 | No |
| 226 | SLC5A7 | SLC5A7 Entrez, Source | solute carrier family 5 (choline transporter), member 7 | 9983 | -0.052 | -0.0449 | No |
| 227 | FNTA | FNTA Entrez, Source | farnesyltransferase, CAAX box, alpha | 10093 | -0.054 | -0.0511 | No |
| 228 | CYP11A1 | CYP11A1 Entrez, Source | cytochrome P450, family 11, subfamily A, polypeptide 1 | 10102 | -0.054 | -0.0496 | No |
| 229 | EIF2B2 | EIF2B2 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 2 beta, 39kDa | 10173 | -0.056 | -0.0527 | No |
| 230 | COX15 | COX15 Entrez, Source | COX15 homolog, cytochrome c oxidase assembly protein (yeast) | 10424 | -0.062 | -0.0694 | No |
| 231 | ELA2 | ELA2 Entrez, Source | elastase 2, neutrophil | 10427 | -0.062 | -0.0671 | No |
| 232 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 10430 | -0.062 | -0.0648 | No |
| 233 | AKT1 | AKT1 Entrez, Source | v-akt murine thymoma viral oncogene homolog 1 | 10634 | -0.068 | -0.0777 | No |
| 234 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 10674 | -0.069 | -0.0780 | No |
| 235 | HSP90AA1 | HSP90AA1 Entrez, Source | heat shock protein 90kDa alpha (cytosolic), class A member 1 | 10705 | -0.070 | -0.0775 | No |
| 236 | HPRT1 | HPRT1 Entrez, Source | hypoxanthine phosphoribosyltransferase 1 (Lesch-Nyhan syndrome) | 10730 | -0.071 | -0.0766 | No |
| 237 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 10801 | -0.073 | -0.0791 | No |
| 238 | AP3D1 | AP3D1 Entrez, Source | adaptor-related protein complex 3, delta 1 subunit | 10841 | -0.074 | -0.0792 | No |
| 239 | BACE2 | BACE2 Entrez, Source | beta-site APP-cleaving enzyme 2 | 10980 | -0.078 | -0.0866 | No |
| 240 | TSPO | TSPO Entrez, Source | translocator protein (18kDa) | 11004 | -0.079 | -0.0853 | No |
| 241 | CD81 | CD81 Entrez, Source | CD81 molecule | 11022 | -0.079 | -0.0835 | No |
| 242 | IL18 | IL18 Entrez, Source | interleukin 18 (interferon-gamma-inducing factor) | 11246 | -0.088 | -0.0971 | No |
| 243 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 11324 | -0.092 | -0.0994 | No |
| 244 | GATM | GATM Entrez, Source | glycine amidinotransferase (L-arginine:glycine amidinotransferase) | 11580 | -0.103 | -0.1148 | No |
| 245 | IL17F | IL17F Entrez, Source | interleukin 17F | 11826 | -0.115 | -0.1290 | No |
| 246 | ATG7 | ATG7 Entrez, Source | ATG7 autophagy related 7 homolog (S. cerevisiae) | 11870 | -0.117 | -0.1277 | No |
| 247 | ALK | ALK Entrez, Source | anaplastic lymphoma kinase (Ki-1) | 12048 | -0.128 | -0.1362 | No |
| 248 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 12112 | -0.132 | -0.1358 | No |
| 249 | CHM | CHM Entrez, Source | choroideremia (Rab escort protein 1) | 12137 | -0.133 | -0.1324 | No |
| 250 | AGPAT6 | AGPAT6 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 6 (lysophosphatidic acid acyltransferase, zeta) | 12308 | -0.149 | -0.1396 | No |
| 251 | EEF2K | EEF2K Entrez, Source | eukaryotic elongation factor-2 kinase | 12382 | -0.156 | -0.1390 | No |
| 252 | EIF2AK3 | EIF2AK3 Entrez, Source | eukaryotic translation initiation factor 2-alpha kinase 3 | 12512 | -0.172 | -0.1421 | No |
| 253 | TM4SF4 | TM4SF4 Entrez, Source | transmembrane 4 L six family member 4 | 12583 | -0.182 | -0.1403 | No |
| 254 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 12607 | -0.185 | -0.1348 | No |
| 255 | PLOD1 | PLOD1 Entrez, Source | procollagen-lysine 1, 2-oxoglutarate 5-dioxygenase 1 | 12626 | -0.189 | -0.1287 | No |
| 256 | HSPB1 | HSPB1 Entrez, Source | heat shock 27kDa protein 1 | 12847 | -0.236 | -0.1363 | No |
| 257 | B3GALNT1 | B3GALNT1 Entrez, Source | beta-1,3-N-acetylgalactosaminyltransferase 1 (globoside blood group) | 12867 | -0.242 | -0.1282 | No |
| 258 | DDAH2 | DDAH2 Entrez, Source | dimethylarginine dimethylaminohydrolase 2 | 12900 | -0.253 | -0.1207 | No |
| 259 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 12906 | -0.254 | -0.1111 | No |
| 260 | GCNT1 | GCNT1 Entrez, Source | glucosaminyl (N-acetyl) transferase 1, core 2 (beta-1,6-N-acetylglucosaminyltransferase) | 13037 | -0.312 | -0.1088 | No |
| 261 | CD276 | CD276 Entrez, Source | CD276 molecule | 13052 | -0.320 | -0.0973 | No |
| 262 | PLCE1 | PLCE1 Entrez, Source | phospholipase C, epsilon 1 | 13110 | -0.363 | -0.0874 | No |
| 263 | TSPAN8 | TSPAN8 Entrez, Source | tetraspanin 8 | 13161 | -0.417 | -0.0748 | No |
| 264 | BCAT1 | BCAT1 Entrez, Source | branched chain aminotransferase 1, cytosolic | 13181 | -0.442 | -0.0589 | No |
| 265 | GCNT3 | GCNT3 Entrez, Source | glucosaminyl (N-acetyl) transferase 3, mucin type | 13194 | -0.458 | -0.0418 | No |
| 266 | LIPC | LIPC Entrez, Source | lipase, hepatic | 13275 | -0.650 | -0.0224 | No |
| 267 | ADM | ADM Entrez, Source | adrenomedullin | 13287 | -0.697 | 0.0042 | No |