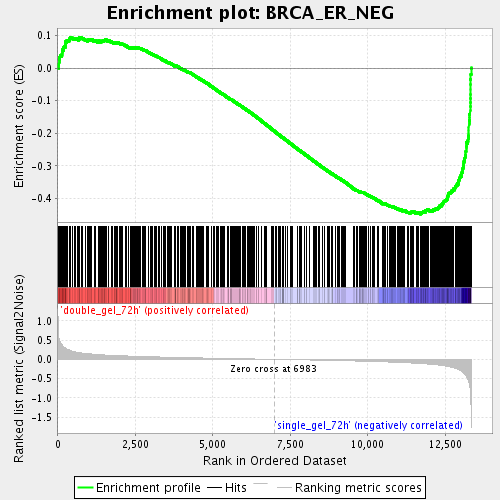
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | BRCA_ER_NEG |
| Enrichment Score (ES) | -0.44813165 |
| Normalized Enrichment Score (NES) | -1.8674895 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.02699792 |
| FWER p-Value | 0.739 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BBOX1 | BBOX1 Entrez, Source | butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 | 13 | 0.740 | 0.0106 | No |
| 2 | LTB | LTB Entrez, Source | lymphotoxin beta (TNF superfamily, member 3) | 35 | 0.539 | 0.0174 | No |
| 3 | AQP9 | AQP9 Entrez, Source | aquaporin 9 | 38 | 0.530 | 0.0256 | No |
| 4 | VCAM1 | VCAM1 Entrez, Source | vascular cell adhesion molecule 1 | 49 | 0.501 | 0.0326 | No |
| 5 | BCL2A1 | BCL2A1 Entrez, Source | BCL2-related protein A1 | 67 | 0.459 | 0.0385 | No |
| 6 | HAAO | HAAO Entrez, Source | 3-hydroxyanthranilate 3,4-dioxygenase | 120 | 0.390 | 0.0406 | No |
| 7 | LPIN1 | LPIN1 Entrez, Source | lipin 1 | 133 | 0.378 | 0.0456 | No |
| 8 | ACAA2 | ACAA2 Entrez, Source | acetyl-Coenzyme A acyltransferase 2 (mitochondrial 3-oxoacyl-Coenzyme A thiolase) | 135 | 0.374 | 0.0514 | No |
| 9 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 144 | 0.362 | 0.0564 | No |
| 10 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 180 | 0.328 | 0.0588 | No |
| 11 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 182 | 0.324 | 0.0638 | No |
| 12 | AADAT | AADAT Entrez, Source | aminoadipate aminotransferase | 200 | 0.309 | 0.0674 | No |
| 13 | SLC2A5 | SLC2A5 Entrez, Source | solute carrier family 2 (facilitated glucose/fructose transporter), member 5 | 238 | 0.290 | 0.0690 | No |
| 14 | RARRES1 | RARRES1 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 1 | 239 | 0.290 | 0.0736 | No |
| 15 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 243 | 0.289 | 0.0779 | No |
| 16 | ANGPTL4 | ANGPTL4 Entrez, Source | angiopoietin-like 4 | 251 | 0.287 | 0.0818 | No |
| 17 | ACN9 | ACN9 Entrez, Source | ACN9 homolog (S. cerevisiae) | 284 | 0.273 | 0.0836 | No |
| 18 | SLC7A7 | SLC7A7 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 7 | 320 | 0.260 | 0.0850 | No |
| 19 | TDO2 | TDO2 Entrez, Source | tryptophan 2,3-dioxygenase | 358 | 0.245 | 0.0859 | No |
| 20 | C1QB | C1QB Entrez, Source | complement component 1, q subcomponent, B chain | 383 | 0.238 | 0.0878 | No |
| 21 | RIPK2 | RIPK2 Entrez, Source | receptor-interacting serine-threonine kinase 2 | 384 | 0.238 | 0.0915 | No |
| 22 | CD14 | CD14 Entrez, Source | CD14 molecule | 400 | 0.231 | 0.0940 | No |
| 23 | PSPH | PSPH Entrez, Source | phosphoserine phosphatase | 459 | 0.213 | 0.0928 | No |
| 24 | HYOU1 | HYOU1 Entrez, Source | hypoxia up-regulated 1 | 540 | 0.194 | 0.0896 | No |
| 25 | MCCC1 | MCCC1 Entrez, Source | methylcrotonoyl-Coenzyme A carboxylase 1 (alpha) | 556 | 0.192 | 0.0914 | No |
| 26 | ARRDC2 | ARRDC2 Entrez, Source | arrestin domain containing 2 | 616 | 0.184 | 0.0897 | No |
| 27 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 678 | 0.176 | 0.0878 | No |
| 28 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 686 | 0.175 | 0.0900 | No |
| 29 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 693 | 0.174 | 0.0922 | No |
| 30 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 695 | 0.174 | 0.0949 | No |
| 31 | TYROBP | TYROBP Entrez, Source | TYRO protein tyrosine kinase binding protein | 747 | 0.168 | 0.0935 | No |
| 32 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 800 | 0.163 | 0.0920 | No |
| 33 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 901 | 0.153 | 0.0866 | No |
| 34 | LYZ | LYZ Entrez, Source | lysozyme (renal amyloidosis) | 961 | 0.148 | 0.0844 | No |
| 35 | SLC39A8 | SLC39A8 Entrez, Source | solute carrier family 39 (zinc transporter), member 8 | 969 | 0.148 | 0.0861 | No |
| 36 | TGFA | TGFA Entrez, Source | transforming growth factor, alpha | 986 | 0.146 | 0.0872 | No |
| 37 | LGALS9 | LGALS9 Entrez, Source | lectin, galactoside-binding, soluble, 9 (galectin 9) | 1031 | 0.142 | 0.0860 | No |
| 38 | CD86 | CD86 Entrez, Source | CD86 molecule | 1061 | 0.139 | 0.0859 | No |
| 39 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 1078 | 0.138 | 0.0868 | No |
| 40 | SLA | SLA Entrez, Source | Src-like-adaptor | 1080 | 0.138 | 0.0889 | No |
| 41 | SLC31A1 | SLC31A1 Entrez, Source | solute carrier family 31 (copper transporters), member 1 | 1173 | 0.132 | 0.0838 | No |
| 42 | EPRS | EPRS Entrez, Source | glutamyl-prolyl-tRNA synthetase | 1177 | 0.132 | 0.0857 | No |
| 43 | ST8SIA1 | ST8SIA1 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 1 | 1223 | 0.128 | 0.0842 | No |
| 44 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 1307 | 0.124 | 0.0797 | No |
| 45 | PIR | PIR Entrez, Source | pirin (iron-binding nuclear protein) | 1308 | 0.124 | 0.0816 | No |
| 46 | FGFBP1 | FGFBP1 Entrez, Source | fibroblast growth factor binding protein 1 | 1310 | 0.124 | 0.0835 | No |
| 47 | TRAF3IP3 | TRAF3IP3 Entrez, Source | TRAF3 interacting protein 3 | 1332 | 0.122 | 0.0837 | No |
| 48 | ASS1 | ASS1 Entrez, Source | argininosuccinate synthetase 1 | 1382 | 0.120 | 0.0818 | No |
| 49 | CCR1 | CCR1 Entrez, Source | chemokine (C-C motif) receptor 1 | 1404 | 0.119 | 0.0820 | No |
| 50 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 1413 | 0.118 | 0.0833 | No |
| 51 | SSR1 | SSR1 Entrez, Source | signal sequence receptor, alpha (translocon-associated protein alpha) | 1437 | 0.116 | 0.0833 | No |
| 52 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 1449 | 0.116 | 0.0843 | No |
| 53 | C3 | C3 Entrez, Source | complement component 3 | 1475 | 0.114 | 0.0841 | No |
| 54 | GSTA3 | GSTA3 Entrez, Source | glutathione S-transferase A3 | 1488 | 0.114 | 0.0850 | No |
| 55 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 1495 | 0.114 | 0.0863 | No |
| 56 | EDN2 | EDN2 Entrez, Source | endothelin 2 | 1518 | 0.112 | 0.0863 | No |
| 57 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 1524 | 0.112 | 0.0877 | No |
| 58 | TUBB3 | TUBB3 Entrez, Source | tubulin, beta 3 | 1557 | 0.110 | 0.0869 | No |
| 59 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 1617 | 0.108 | 0.0840 | No |
| 60 | PCDH8 | PCDH8 Entrez, Source | protocadherin 8 | 1627 | 0.107 | 0.0850 | No |
| 61 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 1644 | 0.107 | 0.0854 | No |
| 62 | RAB7L1 | RAB7L1 Entrez, Source | RAB7, member RAS oncogene family-like 1 | 1719 | 0.104 | 0.0813 | No |
| 63 | IL1B | IL1B Entrez, Source | interleukin 1, beta | 1767 | 0.102 | 0.0793 | No |
| 64 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 1830 | 0.100 | 0.0760 | No |
| 65 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 1840 | 0.100 | 0.0769 | No |
| 66 | CCR2 | CCR2 Entrez, Source | chemokine (C-C motif) receptor 2 | 1845 | 0.100 | 0.0781 | No |
| 67 | TTC7A | TTC7A Entrez, Source | tetratricopeptide repeat domain 7A | 1888 | 0.098 | 0.0764 | No |
| 68 | RPP38 | RPP38 Entrez, Source | ribonuclease P/MRP 38kDa subunit | 1892 | 0.098 | 0.0777 | No |
| 69 | PBEF1 | PBEF1 Entrez, Source | pre-B-cell colony enhancing factor 1 | 1897 | 0.097 | 0.0789 | No |
| 70 | HCK | HCK Entrez, Source | hemopoietic cell kinase | 1900 | 0.097 | 0.0803 | No |
| 71 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 1921 | 0.097 | 0.0802 | No |
| 72 | BOP1 | BOP1 Entrez, Source | block of proliferation 1 | 1983 | 0.094 | 0.0770 | No |
| 73 | CYP27A1 | CYP27A1 Entrez, Source | cytochrome P450, family 27, subfamily A, polypeptide 1 | 2033 | 0.093 | 0.0746 | No |
| 74 | PGM1 | PGM1 Entrez, Source | phosphoglucomutase 1 | 2063 | 0.092 | 0.0738 | No |
| 75 | KCNN4 | KCNN4 Entrez, Source | potassium intermediate/small conductance calcium-activated channel, subfamily N, member 4 | 2085 | 0.091 | 0.0736 | No |
| 76 | HSPA14 | HSPA14 Entrez, Source | heat shock 70kDa protein 14 | 2096 | 0.091 | 0.0742 | No |
| 77 | CASP4 | CASP4 Entrez, Source | caspase 4, apoptosis-related cysteine peptidase | 2168 | 0.089 | 0.0701 | No |
| 78 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 2205 | 0.088 | 0.0687 | No |
| 79 | HIST1H1E | HIST1H1E Entrez, Source | histone cluster 1, H1e | 2288 | 0.085 | 0.0636 | No |
| 80 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 2338 | 0.084 | 0.0611 | No |
| 81 | SEC61A1 | SEC61A1 Entrez, Source | Sec61 alpha 1 subunit (S. cerevisiae) | 2340 | 0.084 | 0.0624 | No |
| 82 | BYSL | BYSL Entrez, Source | bystin-like | 2347 | 0.084 | 0.0632 | No |
| 83 | C10ORF7 | C10ORF7 Entrez, Source | chromosome 10 open reading frame 7 | 2355 | 0.084 | 0.0640 | No |
| 84 | DNMT3B | DNMT3B Entrez, Source | DNA (cytosine-5-)-methyltransferase 3 beta | 2369 | 0.083 | 0.0643 | No |
| 85 | PTS | PTS Entrez, Source | 6-pyruvoyltetrahydropterin synthase | 2407 | 0.082 | 0.0627 | No |
| 86 | UBD | UBD Entrez, Source | ubiquitin D | 2423 | 0.082 | 0.0628 | No |
| 87 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 2426 | 0.082 | 0.0639 | No |
| 88 | CD53 | CD53 Entrez, Source | CD53 molecule | 2467 | 0.081 | 0.0621 | No |
| 89 | GTPBP4 | GTPBP4 Entrez, Source | GTP binding protein 4 | 2484 | 0.080 | 0.0621 | No |
| 90 | ECHDC1 | ECHDC1 Entrez, Source | enoyl Coenzyme A hydratase domain containing 1 | 2494 | 0.080 | 0.0627 | No |
| 91 | NUMBL | NUMBL Entrez, Source | numb homolog (Drosophila)-like | 2500 | 0.080 | 0.0635 | No |
| 92 | NOB1 | NOB1 Entrez, Source | NIN1/RPN12 binding protein 1 homolog (S. cerevisiae) | 2520 | 0.079 | 0.0633 | No |
| 93 | SRD5A1 | SRD5A1 Entrez, Source | steroid-5-alpha-reductase, alpha polypeptide 1 (3-oxo-5 alpha-steroid delta 4-dehydrogenase alpha 1) | 2561 | 0.078 | 0.0614 | No |
| 94 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 2566 | 0.078 | 0.0623 | No |
| 95 | OPTN | OPTN Entrez, Source | optineurin | 2577 | 0.077 | 0.0627 | No |
| 96 | ENAH | ENAH Entrez, Source | enabled homolog (Drosophila) | 2600 | 0.076 | 0.0622 | No |
| 97 | GRAMD3 | GRAMD3 Entrez, Source | GRAM domain containing 3 | 2638 | 0.075 | 0.0605 | No |
| 98 | FGL2 | FGL2 Entrez, Source | fibrinogen-like 2 | 2640 | 0.075 | 0.0616 | No |
| 99 | LIMK2 | LIMK2 Entrez, Source | LIM domain kinase 2 | 2665 | 0.074 | 0.0609 | No |
| 100 | GLRX2 | GLRX2 Entrez, Source | glutaredoxin 2 | 2722 | 0.073 | 0.0577 | No |
| 101 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 2746 | 0.073 | 0.0571 | No |
| 102 | SOX9 | SOX9 Entrez, Source | SRY (sex determining region Y)-box 9 (campomelic dysplasia, autosomal sex-reversal) | 2781 | 0.072 | 0.0555 | No |
| 103 | LRRC8D | LRRC8D Entrez, Source | leucine rich repeat containing 8 family, member D | 2812 | 0.071 | 0.0543 | No |
| 104 | PLEKHB1 | PLEKHB1 Entrez, Source | pleckstrin homology domain containing, family B (evectins) member 1 | 2823 | 0.071 | 0.0547 | No |
| 105 | EGFL6 | EGFL6 Entrez, Source | EGF-like-domain, multiple 6 | 2830 | 0.071 | 0.0553 | No |
| 106 | CSDA | CSDA Entrez, Source | cold shock domain protein A | 2927 | 0.068 | 0.0489 | No |
| 107 | DSC2 | DSC2 Entrez, Source | desmocollin 2 | 2983 | 0.067 | 0.0457 | No |
| 108 | CENTG1 | CENTG1 Entrez, Source | centaurin, gamma 1 | 3016 | 0.066 | 0.0442 | No |
| 109 | ACOT7 | ACOT7 Entrez, Source | acyl-CoA thioesterase 7 | 3055 | 0.065 | 0.0423 | No |
| 110 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 3058 | 0.065 | 0.0431 | No |
| 111 | LAP3 | LAP3 Entrez, Source | leucine aminopeptidase 3 | 3125 | 0.064 | 0.0390 | No |
| 112 | PTBP2 | PTBP2 Entrez, Source | polypyrimidine tract binding protein 2 | 3139 | 0.064 | 0.0390 | No |
| 113 | DNM1L | DNM1L Entrez, Source | dynamin 1-like | 3184 | 0.063 | 0.0365 | No |
| 114 | MMP12 | MMP12 Entrez, Source | matrix metallopeptidase 12 (macrophage elastase) | 3186 | 0.063 | 0.0374 | No |
| 115 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 3253 | 0.061 | 0.0333 | No |
| 116 | EIF2B3 | EIF2B3 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 3 gamma, 58kDa | 3277 | 0.061 | 0.0324 | No |
| 117 | GIMAP4 | GIMAP4 Entrez, Source | GTPase, IMAP family member 4 | 3337 | 0.059 | 0.0288 | No |
| 118 | MRPL37 | MRPL37 Entrez, Source | mitochondrial ribosomal protein L37 | 3397 | 0.058 | 0.0251 | No |
| 119 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 3416 | 0.058 | 0.0246 | No |
| 120 | COPG | COPG Entrez, Source | coatomer protein complex, subunit gamma | 3452 | 0.057 | 0.0228 | No |
| 121 | KCNK1 | KCNK1 Entrez, Source | potassium channel, subfamily K, member 1 | 3465 | 0.057 | 0.0227 | No |
| 122 | MRPL11 | MRPL11 Entrez, Source | mitochondrial ribosomal protein L11 | 3529 | 0.055 | 0.0187 | No |
| 123 | PSAT1 | PSAT1 Entrez, Source | phosphoserine aminotransferase 1 | 3569 | 0.054 | 0.0165 | No |
| 124 | SH3KBP1 | SH3KBP1 Entrez, Source | SH3-domain kinase binding protein 1 | 3572 | 0.054 | 0.0172 | No |
| 125 | GMPS | GMPS Entrez, Source | guanine monphosphate synthetase | 3595 | 0.054 | 0.0163 | No |
| 126 | ACSL4 | ACSL4 Entrez, Source | acyl-CoA synthetase long-chain family member 4 | 3612 | 0.053 | 0.0159 | No |
| 127 | SFRP1 | SFRP1 Entrez, Source | secreted frizzled-related protein 1 | 3641 | 0.053 | 0.0146 | No |
| 128 | SLC4A1 | SLC4A1 Entrez, Source | solute carrier family 4, anion exchanger, member 1 (erythrocyte membrane protein band 3, Diego blood group) | 3668 | 0.053 | 0.0134 | No |
| 129 | UFD1L | UFD1L Entrez, Source | ubiquitin fusion degradation 1 like (yeast) | 3750 | 0.051 | 0.0079 | No |
| 130 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 3790 | 0.050 | 0.0056 | No |
| 131 | CD2 | CD2 Entrez, Source | CD2 molecule | 3794 | 0.050 | 0.0062 | No |
| 132 | CTSC | CTSC Entrez, Source | cathepsin C | 3796 | 0.050 | 0.0069 | No |
| 133 | IL10RA | IL10RA Entrez, Source | interleukin 10 receptor, alpha | 3799 | 0.050 | 0.0075 | No |
| 134 | FUT4 | FUT4 Entrez, Source | fucosyltransferase 4 (alpha (1,3) fucosyltransferase, myeloid-specific) | 3809 | 0.050 | 0.0076 | No |
| 135 | NMU | NMU Entrez, Source | neuromedin U | 3853 | 0.049 | 0.0050 | No |
| 136 | MMP9 | MMP9 Entrez, Source | matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) | 3861 | 0.049 | 0.0052 | No |
| 137 | GLIPR1 | GLIPR1 Entrez, Source | GLI pathogenesis-related 1 (glioma) | 3896 | 0.049 | 0.0033 | No |
| 138 | RNASE1 | RNASE1 Entrez, Source | ribonuclease, RNase A family, 1 (pancreatic) | 3953 | 0.047 | -0.0003 | No |
| 139 | DNAJB11 | DNAJB11 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 11 | 3986 | 0.047 | -0.0021 | No |
| 140 | PDZK1IP1 | PDZK1IP1 Entrez, Source | PDZK1 interacting protein 1 | 4023 | 0.046 | -0.0041 | No |
| 141 | CMAS | CMAS Entrez, Source | cytidine monophosphate N-acetylneuraminic acid synthetase | 4031 | 0.046 | -0.0040 | No |
| 142 | P2RY6 | P2RY6 Entrez, Source | pyrimidinergic receptor P2Y, G-protein coupled, 6 | 4048 | 0.045 | -0.0045 | No |
| 143 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 4076 | 0.045 | -0.0059 | No |
| 144 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 4079 | 0.045 | -0.0053 | No |
| 145 | RPN1 | RPN1 Entrez, Source | ribophorin I | 4101 | 0.045 | -0.0063 | No |
| 146 | GMFG | GMFG Entrez, Source | glia maturation factor, gamma | 4170 | 0.043 | -0.0109 | No |
| 147 | PHGDH | PHGDH Entrez, Source | phosphoglycerate dehydrogenase | 4199 | 0.043 | -0.0124 | No |
| 148 | KLK7 | KLK7 Entrez, Source | kallikrein 7 (chymotryptic, stratum corneum) | 4220 | 0.043 | -0.0133 | No |
| 149 | ATP5G3 | ATP5G3 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C3 (subunit 9) | 4222 | 0.043 | -0.0127 | No |
| 150 | ASNS | ASNS Entrez, Source | asparagine synthetase | 4245 | 0.042 | -0.0138 | No |
| 151 | KCNJ15 | KCNJ15 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 15 | 4250 | 0.042 | -0.0134 | No |
| 152 | SEC13L1 | SEC13L1 Entrez, Source | SEC13-like 1 (S. cerevisiae) | 4259 | 0.042 | -0.0134 | No |
| 153 | AARS | AARS Entrez, Source | alanyl-tRNA synthetase | 4275 | 0.042 | -0.0139 | No |
| 154 | SLC7A1 | SLC7A1 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 | 4346 | 0.040 | -0.0187 | No |
| 155 | GARS | GARS Entrez, Source | glycyl-tRNA synthetase | 4370 | 0.040 | -0.0199 | No |
| 156 | GIMAP5 | GIMAP5 Entrez, Source | GTPase, IMAP family member 5 | 4483 | 0.038 | -0.0280 | No |
| 157 | KLRD1 | KLRD1 Entrez, Source | killer cell lectin-like receptor subfamily D, member 1 | 4487 | 0.038 | -0.0276 | No |
| 158 | KLK6 | KLK6 Entrez, Source | kallikrein 6 (neurosin, zyme) | 4508 | 0.038 | -0.0286 | No |
| 159 | CYB5R2 | CYB5R2 Entrez, Source | cytochrome b5 reductase 2 | 4527 | 0.037 | -0.0294 | No |
| 160 | HIST1H4B | HIST1H4B Entrez, Source | histone cluster 1, H4b | 4584 | 0.036 | -0.0332 | No |
| 161 | SLC5A6 | SLC5A6 Entrez, Source | solute carrier family 5 (sodium-dependent vitamin transporter), member 6 | 4600 | 0.036 | -0.0338 | No |
| 162 | LYAR | LYAR Entrez, Source | - | 4619 | 0.036 | -0.0346 | No |
| 163 | LAPTM5 | LAPTM5 Entrez, Source | lysosomal associated multispanning membrane protein 5 | 4666 | 0.035 | -0.0377 | No |
| 164 | UCHL5 | UCHL5 Entrez, Source | ubiquitin carboxyl-terminal hydrolase L5 | 4673 | 0.035 | -0.0376 | No |
| 165 | CD247 | CD247 Entrez, Source | CD247 molecule | 4702 | 0.034 | -0.0392 | No |
| 166 | CCL3 | CCL3 Entrez, Source | chemokine (C-C motif) ligand 3 | 4784 | 0.033 | -0.0450 | No |
| 167 | ORM1 | ORM1 Entrez, Source | orosomucoid 1 | 4816 | 0.032 | -0.0469 | No |
| 168 | IGSF6 | IGSF6 Entrez, Source | immunoglobulin superfamily, member 6 | 4839 | 0.032 | -0.0481 | No |
| 169 | SELE | SELE Entrez, Source | selectin E (endothelial adhesion molecule 1) | 4846 | 0.032 | -0.0481 | No |
| 170 | WDR5 | WDR5 Entrez, Source | WD repeat domain 5 | 4959 | 0.030 | -0.0564 | No |
| 171 | SYK | SYK Entrez, Source | spleen tyrosine kinase | 5035 | 0.029 | -0.0618 | No |
| 172 | PSCD1 | PSCD1 Entrez, Source | pleckstrin homology, Sec7 and coiled-coil domains 1(cytohesin 1) | 5060 | 0.028 | -0.0632 | No |
| 173 | RIMS3 | RIMS3 Entrez, Source | regulating synaptic membrane exocytosis 3 | 5124 | 0.027 | -0.0677 | No |
| 174 | ESRRA | ESRRA Entrez, Source | estrogen-related receptor alpha | 5141 | 0.027 | -0.0685 | No |
| 175 | IL2RB | IL2RB Entrez, Source | interleukin 2 receptor, beta | 5178 | 0.027 | -0.0709 | No |
| 176 | COX4I1 | COX4I1 Entrez, Source | cytochrome c oxidase subunit IV isoform 1 | 5245 | 0.025 | -0.0756 | No |
| 177 | SH2D2A | SH2D2A Entrez, Source | SH2 domain protein 2A | 5263 | 0.025 | -0.0765 | No |
| 178 | PADI2 | PADI2 Entrez, Source | peptidyl arginine deiminase, type II | 5275 | 0.025 | -0.0770 | No |
| 179 | CYBB | CYBB Entrez, Source | cytochrome b-245, beta polypeptide (chronic granulomatous disease) | 5312 | 0.024 | -0.0794 | No |
| 180 | GALK1 | GALK1 Entrez, Source | galactokinase 1 | 5339 | 0.024 | -0.0811 | No |
| 181 | HSD17B12 | HSD17B12 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 12 | 5355 | 0.024 | -0.0818 | No |
| 182 | NPR3 | NPR3 Entrez, Source | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | 5375 | 0.024 | -0.0830 | No |
| 183 | ODC1 | ODC1 Entrez, Source | ornithine decarboxylase 1 | 5487 | 0.022 | -0.0913 | No |
| 184 | ANXA8 | ANXA8 Entrez, Source | annexin A8 | 5494 | 0.022 | -0.0914 | No |
| 185 | TPRKB | TPRKB Entrez, Source | TP53RK binding protein | 5505 | 0.022 | -0.0918 | No |
| 186 | PLEK | PLEK Entrez, Source | pleckstrin | 5509 | 0.022 | -0.0917 | No |
| 187 | FNBP1 | FNBP1 Entrez, Source | formin binding protein 1 | 5572 | 0.021 | -0.0962 | No |
| 188 | RNF138 | RNF138 Entrez, Source | ring finger protein 138 | 5579 | 0.021 | -0.0964 | No |
| 189 | GPI | GPI Entrez, Source | glucose phosphate isomerase | 5582 | 0.021 | -0.0962 | No |
| 190 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 5597 | 0.020 | -0.0970 | No |
| 191 | PDIA6 | PDIA6 Entrez, Source | protein disulfide isomerase family A, member 6 | 5601 | 0.020 | -0.0969 | No |
| 192 | PTPRC | PTPRC Entrez, Source | protein tyrosine phosphatase, receptor type, C | 5608 | 0.020 | -0.0970 | No |
| 193 | UGP2 | UGP2 Entrez, Source | UDP-glucose pyrophosphorylase 2 | 5626 | 0.020 | -0.0980 | No |
| 194 | FBL | FBL Entrez, Source | fibrillarin | 5673 | 0.019 | -0.1013 | No |
| 195 | DDX39 | DDX39 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 39 | 5687 | 0.019 | -0.1020 | No |
| 196 | EIF2C2 | EIF2C2 Entrez, Source | eukaryotic translation initiation factor 2C, 2 | 5725 | 0.018 | -0.1046 | No |
| 197 | CD3G | CD3G Entrez, Source | CD3g molecule, gamma (CD3-TCR complex) | 5771 | 0.018 | -0.1078 | No |
| 198 | PDHA1 | PDHA1 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 1 | 5797 | 0.017 | -0.1095 | No |
| 199 | C1ORF121 | C1ORF121 Entrez, Source | chromosome 1 open reading frame 121 | 5836 | 0.017 | -0.1122 | No |
| 200 | CUGBP2 | CUGBP2 Entrez, Source | CUG triplet repeat, RNA binding protein 2 | 5853 | 0.017 | -0.1132 | No |
| 201 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 5856 | 0.016 | -0.1131 | No |
| 202 | IFI30 | IFI30 Entrez, Source | interferon, gamma-inducible protein 30 | 5860 | 0.016 | -0.1131 | No |
| 203 | NUP93 | NUP93 Entrez, Source | nucleoporin 93kDa | 5890 | 0.016 | -0.1151 | No |
| 204 | ADFP | ADFP Entrez, Source | adipose differentiation-related protein | 5901 | 0.016 | -0.1156 | No |
| 205 | RHCG | RHCG Entrez, Source | Rh family, C glycoprotein | 5941 | 0.015 | -0.1184 | No |
| 206 | ART3 | ART3 Entrez, Source | ADP-ribosyltransferase 3 | 5951 | 0.015 | -0.1189 | No |
| 207 | C16ORF80 | C16ORF80 Entrez, Source | chromosome 16 open reading frame 80 | 5984 | 0.014 | -0.1211 | No |
| 208 | NF2 | NF2 Entrez, Source | neurofibromin 2 (bilateral acoustic neuroma) | 6018 | 0.014 | -0.1235 | No |
| 209 | PSMB2 | PSMB2 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 2 | 6045 | 0.014 | -0.1253 | No |
| 210 | DNAJA2 | DNAJA2 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 2 | 6068 | 0.013 | -0.1268 | No |
| 211 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 6114 | 0.013 | -0.1301 | No |
| 212 | GZMA | GZMA Entrez, Source | granzyme A (granzyme 1, cytotoxic T-lymphocyte-associated serine esterase 3) | 6121 | 0.012 | -0.1304 | No |
| 213 | GZMB | GZMB Entrez, Source | granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) | 6123 | 0.012 | -0.1303 | No |
| 214 | CALB2 | CALB2 Entrez, Source | calbindin 2, 29kDa (calretinin) | 6162 | 0.012 | -0.1330 | No |
| 215 | INPP5D | INPP5D Entrez, Source | inositol polyphosphate-5-phosphatase, 145kDa | 6188 | 0.011 | -0.1348 | No |
| 216 | BCL11A | BCL11A Entrez, Source | B-cell CLL/lymphoma 11A (zinc finger protein) | 6200 | 0.011 | -0.1355 | No |
| 217 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 6208 | 0.011 | -0.1359 | No |
| 218 | GALNT3 | GALNT3 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 3 (GalNAc-T3) | 6251 | 0.010 | -0.1390 | No |
| 219 | FASLG | FASLG Entrez, Source | Fas ligand (TNF superfamily, member 6) | 6273 | 0.010 | -0.1404 | No |
| 220 | STAT4 | STAT4 Entrez, Source | signal transducer and activator of transcription 4 | 6316 | 0.009 | -0.1436 | No |
| 221 | NMI | NMI Entrez, Source | N-myc (and STAT) interactor | 6331 | 0.009 | -0.1445 | No |
| 222 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 6402 | 0.008 | -0.1498 | No |
| 223 | HCLS1 | HCLS1 Entrez, Source | hematopoietic cell-specific Lyn substrate 1 | 6405 | 0.008 | -0.1499 | No |
| 224 | TGFBI | TGFBI Entrez, Source | transforming growth factor, beta-induced, 68kDa | 6461 | 0.007 | -0.1540 | No |
| 225 | DEFB1 | DEFB1 Entrez, Source | defensin, beta 1 | 6468 | 0.007 | -0.1544 | No |
| 226 | USP10 | USP10 Entrez, Source | ubiquitin specific peptidase 10 | 6559 | 0.006 | -0.1613 | No |
| 227 | LCP2 | LCP2 Entrez, Source | lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa) | 6665 | 0.004 | -0.1694 | No |
| 228 | CCL4 | CCL4 Entrez, Source | chemokine (C-C motif) ligand 4 | 6696 | 0.004 | -0.1717 | No |
| 229 | NARF | NARF Entrez, Source | nuclear prelamin A recognition factor | 6718 | 0.004 | -0.1733 | No |
| 230 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6738 | 0.003 | -0.1747 | No |
| 231 | TNF | TNF Entrez, Source | tumor necrosis factor (TNF superfamily, member 2) | 6739 | 0.003 | -0.1746 | No |
| 232 | SRPRB | SRPRB Entrez, Source | signal recognition particle receptor, B subunit | 6895 | 0.001 | -0.1867 | No |
| 233 | GAPDHS | GAPDHS Entrez, Source | glyceraldehyde-3-phosphate dehydrogenase, spermatogenic | 6929 | 0.001 | -0.1892 | No |
| 234 | FOLR1 | FOLR1 Entrez, Source | folate receptor 1 (adult) | 6957 | 0.000 | -0.1913 | No |
| 235 | CNGA1 | CNGA1 Entrez, Source | cyclic nucleotide gated channel alpha 1 | 7034 | -0.001 | -0.1972 | No |
| 236 | CXCR3 | CXCR3 Entrez, Source | chemokine (C-X-C motif) receptor 3 | 7038 | -0.001 | -0.1975 | No |
| 237 | RHEB | RHEB Entrez, Source | Ras homolog enriched in brain | 7054 | -0.001 | -0.1986 | No |
| 238 | GIT2 | GIT2 Entrez, Source | G protein-coupled receptor kinase interactor 2 | 7104 | -0.002 | -0.2024 | No |
| 239 | PGK1 | PGK1 Entrez, Source | phosphoglycerate kinase 1 | 7152 | -0.002 | -0.2060 | No |
| 240 | GATAD2A | GATAD2A Entrez, Source | GATA zinc finger domain containing 2A | 7168 | -0.003 | -0.2071 | No |
| 241 | NUP155 | NUP155 Entrez, Source | nucleoporin 155kDa | 7188 | -0.003 | -0.2086 | No |
| 242 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 7239 | -0.004 | -0.2124 | No |
| 243 | PSMB4 | PSMB4 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 4 | 7246 | -0.004 | -0.2128 | No |
| 244 | KLK10 | KLK10 Entrez, Source | kallikrein 10 | 7247 | -0.004 | -0.2128 | No |
| 245 | CD52 | CD52 Entrez, Source | CD52 molecule | 7261 | -0.004 | -0.2137 | No |
| 246 | KARS | KARS Entrez, Source | lysyl-tRNA synthetase | 7266 | -0.004 | -0.2140 | No |
| 247 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 7275 | -0.004 | -0.2145 | No |
| 248 | NUDT21 | NUDT21 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 21 | 7285 | -0.004 | -0.2151 | No |
| 249 | SOD3 | SOD3 Entrez, Source | superoxide dismutase 3, extracellular | 7289 | -0.004 | -0.2153 | No |
| 250 | UQCRH | UQCRH Entrez, Source | ubiquinol-cytochrome c reductase hinge protein | 7358 | -0.005 | -0.2205 | No |
| 251 | SOX10 | SOX10 Entrez, Source | SRY (sex determining region Y)-box 10 | 7415 | -0.006 | -0.2248 | No |
| 252 | FUS | FUS Entrez, Source | fusion (involved in t(12;16) in malignant liposarcoma) | 7497 | -0.007 | -0.2310 | No |
| 253 | AIF1 | AIF1 Entrez, Source | allograft inflammatory factor 1 | 7500 | -0.007 | -0.2310 | No |
| 254 | PSMA4 | PSMA4 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 4 | 7529 | -0.008 | -0.2331 | No |
| 255 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 7577 | -0.009 | -0.2366 | No |
| 256 | CIB2 | CIB2 Entrez, Source | calcium and integrin binding family member 2 | 7716 | -0.011 | -0.2472 | No |
| 257 | CMPK | CMPK Entrez, Source | cytidylate kinase | 7736 | -0.011 | -0.2485 | No |
| 258 | GAPDH | GAPDH Entrez, Source | glyceraldehyde-3-phosphate dehydrogenase | 7743 | -0.011 | -0.2487 | No |
| 259 | CCNC | CCNC Entrez, Source | cyclin C | 7782 | -0.012 | -0.2515 | No |
| 260 | PTDSS1 | PTDSS1 Entrez, Source | phosphatidylserine synthase 1 | 7804 | -0.012 | -0.2530 | No |
| 261 | PLD1 | PLD1 Entrez, Source | phospholipase D1, phosphatidylcholine-specific | 7809 | -0.012 | -0.2531 | No |
| 262 | ADAM19 | ADAM19 Entrez, Source | ADAM metallopeptidase domain 19 (meltrin beta) | 7823 | -0.012 | -0.2539 | No |
| 263 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 7875 | -0.013 | -0.2577 | No |
| 264 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 7948 | -0.014 | -0.2630 | No |
| 265 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 7962 | -0.014 | -0.2638 | No |
| 266 | CEP57 | CEP57 Entrez, Source | centrosomal protein 57kDa | 8017 | -0.015 | -0.2678 | No |
| 267 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 8036 | -0.015 | -0.2690 | No |
| 268 | SERBP1 | SERBP1 Entrez, Source | SERPINE1 mRNA binding protein 1 | 8112 | -0.016 | -0.2745 | No |
| 269 | SRPK1 | SRPK1 Entrez, Source | SFRS protein kinase 1 | 8261 | -0.019 | -0.2857 | No |
| 270 | ITGB2 | ITGB2 Entrez, Source | integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) | 8281 | -0.020 | -0.2869 | No |
| 271 | PSME4 | PSME4 Entrez, Source | proteasome (prosome, macropain) activator subunit 4 | 8299 | -0.020 | -0.2879 | No |
| 272 | CD48 | CD48 Entrez, Source | CD48 molecule | 8344 | -0.021 | -0.2910 | No |
| 273 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 8416 | -0.022 | -0.2962 | No |
| 274 | EIF5A | EIF5A Entrez, Source | eukaryotic translation initiation factor 5A | 8430 | -0.022 | -0.2969 | No |
| 275 | CORO1A | CORO1A Entrez, Source | coronin, actin binding protein, 1A | 8440 | -0.022 | -0.2972 | No |
| 276 | PSMD2 | PSMD2 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 2 | 8545 | -0.024 | -0.3050 | No |
| 277 | CHAF1B | CHAF1B Entrez, Source | chromatin assembly factor 1, subunit B (p60) | 8597 | -0.025 | -0.3085 | No |
| 278 | LPXN | LPXN Entrez, Source | leupaxin | 8598 | -0.025 | -0.3082 | No |
| 279 | CLEC4A | CLEC4A Entrez, Source | C-type lectin domain family 4, member A | 8686 | -0.026 | -0.3145 | No |
| 280 | CDH3 | CDH3 Entrez, Source | cadherin 3, type 1, P-cadherin (placental) | 8724 | -0.027 | -0.3170 | No |
| 281 | SSRP1 | SSRP1 Entrez, Source | structure specific recognition protein 1 | 8733 | -0.027 | -0.3172 | No |
| 282 | RYK | RYK Entrez, Source | RYK receptor-like tyrosine kinase | 8768 | -0.028 | -0.3194 | No |
| 283 | TES | TES Entrez, Source | testis derived transcript (3 LIM domains) | 8843 | -0.029 | -0.3247 | No |
| 284 | PPP1CB | PPP1CB Entrez, Source | protein phosphatase 1, catalytic subunit, beta isoform | 8852 | -0.029 | -0.3249 | No |
| 285 | SLC1A3 | SLC1A3 Entrez, Source | solute carrier family 1 (glial high affinity glutamate transporter), member 3 | 8857 | -0.029 | -0.3247 | No |
| 286 | SELL | SELL Entrez, Source | selectin L (lymphocyte adhesion molecule 1) | 8867 | -0.029 | -0.3250 | No |
| 287 | HMGN3 | HMGN3 Entrez, Source | high mobility group nucleosomal binding domain 3 | 8947 | -0.031 | -0.3306 | No |
| 288 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 9012 | -0.032 | -0.3351 | No |
| 289 | CCT5 | CCT5 Entrez, Source | chaperonin containing TCP1, subunit 5 (epsilon) | 9022 | -0.033 | -0.3353 | No |
| 290 | RNASEH1 | RNASEH1 Entrez, Source | ribonuclease H1 | 9069 | -0.033 | -0.3383 | No |
| 291 | REXO2 | REXO2 Entrez, Source | REX2, RNA exonuclease 2 homolog (S. cerevisiae) | 9073 | -0.034 | -0.3380 | No |
| 292 | GDI2 | GDI2 Entrez, Source | GDP dissociation inhibitor 2 | 9090 | -0.034 | -0.3388 | No |
| 293 | DNMT3A | DNMT3A Entrez, Source | DNA (cytosine-5-)-methyltransferase 3 alpha | 9156 | -0.035 | -0.3433 | No |
| 294 | RTCD1 | RTCD1 Entrez, Source | RNA terminal phosphate cyclase domain 1 | 9183 | -0.035 | -0.3447 | No |
| 295 | CSPG4 | CSPG4 Entrez, Source | chondroitin sulfate proteoglycan 4 (melanoma-associated) | 9206 | -0.036 | -0.3459 | No |
| 296 | LIMK1 | LIMK1 Entrez, Source | LIM domain kinase 1 | 9254 | -0.037 | -0.3490 | No |
| 297 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 9276 | -0.037 | -0.3500 | No |
| 298 | B3GNT7 | B3GNT7 Entrez, Source | UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 7 | 9552 | -0.043 | -0.3708 | No |
| 299 | RDX | RDX Entrez, Source | radixin | 9584 | -0.043 | -0.3725 | No |
| 300 | TMEM158 | TMEM158 Entrez, Source | transmembrane protein 158 | 9629 | -0.044 | -0.3752 | No |
| 301 | LAMP2 | LAMP2 Entrez, Source | lysosomal-associated membrane protein 2 | 9636 | -0.044 | -0.3750 | No |
| 302 | SLC15A3 | SLC15A3 Entrez, Source | solute carrier family 15, member 3 | 9650 | -0.045 | -0.3753 | No |
| 303 | PALMD | PALMD Entrez, Source | palmdelphin | 9671 | -0.045 | -0.3762 | No |
| 304 | CCT4 | CCT4 Entrez, Source | chaperonin containing TCP1, subunit 4 (delta) | 9733 | -0.046 | -0.3802 | No |
| 305 | S100A1 | S100A1 Entrez, Source | S100 calcium binding protein A1 | 9735 | -0.046 | -0.3795 | No |
| 306 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 9737 | -0.046 | -0.3789 | No |
| 307 | SERPINB5 | SERPINB5 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 5 | 9769 | -0.047 | -0.3806 | No |
| 308 | BIRC2 | BIRC2 Entrez, Source | baculoviral IAP repeat-containing 2 | 9776 | -0.047 | -0.3803 | No |
| 309 | PITPNA | PITPNA Entrez, Source | phosphatidylinositol transfer protein, alpha | 9808 | -0.048 | -0.3819 | No |
| 310 | CCR5 | CCR5 Entrez, Source | chemokine (C-C motif) receptor 5 | 9822 | -0.048 | -0.3822 | No |
| 311 | PLCG2 | PLCG2 Entrez, Source | phospholipase C, gamma 2 (phosphatidylinositol-specific) | 9827 | -0.048 | -0.3818 | No |
| 312 | FABP5 | FABP5 Entrez, Source | fatty acid binding protein 5 (psoriasis-associated) | 9829 | -0.048 | -0.3811 | No |
| 313 | GMDS | GMDS Entrez, Source | GDP-mannose 4,6-dehydratase | 9847 | -0.049 | -0.3816 | No |
| 314 | PITPNM1 | PITPNM1 Entrez, Source | phosphatidylinositol transfer protein, membrane-associated 1 | 9868 | -0.049 | -0.3824 | No |
| 315 | HDGF | HDGF Entrez, Source | hepatoma-derived growth factor (high-mobility group protein 1-like) | 9904 | -0.050 | -0.3844 | No |
| 316 | FBXL10 | FBXL10 Entrez, Source | F-box and leucine-rich repeat protein 10 | 9918 | -0.050 | -0.3846 | No |
| 317 | ZIC1 | ZIC1 Entrez, Source | Zic family member 1 (odd-paired homolog, Drosophila) | 9968 | -0.051 | -0.3876 | No |
| 318 | SCRG1 | SCRG1 Entrez, Source | - | 10012 | -0.052 | -0.3901 | No |
| 319 | PRDX4 | PRDX4 Entrez, Source | peroxiredoxin 4 | 10020 | -0.053 | -0.3898 | No |
| 320 | MARCKS | MARCKS Entrez, Source | myristoylated alanine-rich protein kinase C substrate | 10092 | -0.054 | -0.3945 | No |
| 321 | CLDN1 | CLDN1 Entrez, Source | claudin 1 | 10094 | -0.054 | -0.3937 | No |
| 322 | STEAP3 | STEAP3 Entrez, Source | STEAP family member 3 | 10141 | -0.055 | -0.3964 | No |
| 323 | ATP1B3 | ATP1B3 Entrez, Source | ATPase, Na+/K+ transporting, beta 3 polypeptide | 10160 | -0.056 | -0.3970 | No |
| 324 | MALL | MALL Entrez, Source | mal, T-cell differentiation protein-like | 10181 | -0.056 | -0.3977 | No |
| 325 | CAPN6 | CAPN6 Entrez, Source | calpain 6 | 10215 | -0.057 | -0.3993 | No |
| 326 | CARHSP1 | CARHSP1 Entrez, Source | calcium regulated heat stable protein 1, 24kDa | 10303 | -0.059 | -0.4052 | No |
| 327 | MRAS | MRAS Entrez, Source | muscle RAS oncogene homolog | 10349 | -0.060 | -0.4077 | No |
| 328 | KIF1B | KIF1B Entrez, Source | kinesin family member 1B | 10358 | -0.060 | -0.4074 | No |
| 329 | CLDN4 | CLDN4 Entrez, Source | claudin 4 | 10487 | -0.064 | -0.4164 | No |
| 330 | PLSCR1 | PLSCR1 Entrez, Source | phospholipid scramblase 1 | 10496 | -0.064 | -0.4160 | No |
| 331 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 10498 | -0.064 | -0.4151 | No |
| 332 | LSR | LSR Entrez, Source | lipolysis stimulated lipoprotein receptor | 10501 | -0.064 | -0.4142 | No |
| 333 | ACTL6A | ACTL6A Entrez, Source | actin-like 6A | 10539 | -0.065 | -0.4161 | No |
| 334 | SOSTDC1 | SOSTDC1 Entrez, Source | sclerostin domain containing 1 | 10561 | -0.066 | -0.4167 | No |
| 335 | CBR1 | CBR1 Entrez, Source | carbonyl reductase 1 | 10569 | -0.066 | -0.4162 | No |
| 336 | LAMP3 | LAMP3 Entrez, Source | lysosomal-associated membrane protein 3 | 10638 | -0.068 | -0.4204 | No |
| 337 | UGT8 | UGT8 Entrez, Source | UDP glycosyltransferase 8 (UDP-galactose ceramide galactosyltransferase) | 10649 | -0.068 | -0.4201 | No |
| 338 | DCPS | DCPS Entrez, Source | decapping enzyme, scavenger | 10711 | -0.070 | -0.4238 | No |
| 339 | TLR2 | TLR2 Entrez, Source | toll-like receptor 2 | 10723 | -0.070 | -0.4235 | No |
| 340 | PFKL | PFKL Entrez, Source | phosphofructokinase, liver | 10756 | -0.071 | -0.4249 | No |
| 341 | SOX11 | SOX11 Entrez, Source | SRY (sex determining region Y)-box 11 | 10773 | -0.072 | -0.4250 | No |
| 342 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 10798 | -0.072 | -0.4257 | No |
| 343 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 10801 | -0.073 | -0.4248 | No |
| 344 | PDE4B | PDE4B Entrez, Source | phosphodiesterase 4B, cAMP-specific (phosphodiesterase E4 dunce homolog, Drosophila) | 10834 | -0.073 | -0.4261 | No |
| 345 | DMN | DMN Entrez, Source | desmuslin | 10857 | -0.074 | -0.4266 | No |
| 346 | TIA1 | TIA1 Entrez, Source | TIA1 cytotoxic granule-associated RNA binding protein | 10903 | -0.075 | -0.4290 | No |
| 347 | CREB3L2 | CREB3L2 Entrez, Source | cAMP responsive element binding protein 3-like 2 | 10974 | -0.078 | -0.4332 | No |
| 348 | BACE2 | BACE2 Entrez, Source | beta-site APP-cleaving enzyme 2 | 10980 | -0.078 | -0.4324 | No |
| 349 | SDCBP | SDCBP Entrez, Source | syndecan binding protein (syntenin) | 11013 | -0.079 | -0.4336 | No |
| 350 | IFNGR1 | IFNGR1 Entrez, Source | interferon gamma receptor 1 | 11041 | -0.080 | -0.4345 | No |
| 351 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 11071 | -0.081 | -0.4355 | No |
| 352 | LXN | LXN Entrez, Source | latexin | 11080 | -0.081 | -0.4348 | No |
| 353 | NFIB | NFIB Entrez, Source | nuclear factor I/B | 11136 | -0.083 | -0.4378 | No |
| 354 | TGM1 | TGM1 Entrez, Source | transglutaminase 1 (K polypeptide epidermal type I, protein-glutamine-gamma-glutamyltransferase) | 11140 | -0.083 | -0.4367 | No |
| 355 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 11163 | -0.084 | -0.4371 | No |
| 356 | MPZL1 | MPZL1 Entrez, Source | myelin protein zero-like 1 | 11172 | -0.085 | -0.4364 | No |
| 357 | DR1 | DR1 Entrez, Source | down-regulator of transcription 1, TBP-binding (negative cofactor 2) | 11186 | -0.085 | -0.4361 | No |
| 358 | KLF6 | KLF6 Entrez, Source | Kruppel-like factor 6 | 11298 | -0.090 | -0.4433 | No |
| 359 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 11317 | -0.091 | -0.4433 | No |
| 360 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 11324 | -0.092 | -0.4423 | No |
| 361 | SPEG | SPEG Entrez, Source | SPEG complex locus | 11368 | -0.093 | -0.4442 | No |
| 362 | BTG3 | BTG3 Entrez, Source | BTG family, member 3 | 11377 | -0.094 | -0.4433 | No |
| 363 | PRKX | PRKX Entrez, Source | protein kinase, X-linked | 11385 | -0.094 | -0.4424 | No |
| 364 | PPP2R1B | PPP2R1B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), beta isoform | 11399 | -0.095 | -0.4419 | No |
| 365 | SDC1 | SDC1 Entrez, Source | syndecan 1 | 11411 | -0.095 | -0.4413 | No |
| 366 | GABRP | GABRP Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, pi | 11428 | -0.096 | -0.4410 | No |
| 367 | CD3D | CD3D Entrez, Source | CD3d molecule, delta (CD3-TCR complex) | 11471 | -0.098 | -0.4427 | No |
| 368 | ACTG2 | ACTG2 Entrez, Source | actin, gamma 2, smooth muscle, enteric | 11475 | -0.099 | -0.4414 | No |
| 369 | FDX1 | FDX1 Entrez, Source | ferredoxin 1 | 11489 | -0.099 | -0.4409 | No |
| 370 | ORC6L | ORC6L Entrez, Source | origin recognition complex, subunit 6 like (yeast) | 11568 | -0.103 | -0.4454 | No |
| 371 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 11588 | -0.104 | -0.4452 | No |
| 372 | TCN2 | TCN2 Entrez, Source | transcobalamin II; macrocytic anemia | 11593 | -0.104 | -0.4439 | No |
| 373 | SMYD2 | SMYD2 Entrez, Source | SET and MYND domain containing 2 | 11621 | -0.105 | -0.4443 | No |
| 374 | BIRC3 | BIRC3 Entrez, Source | baculoviral IAP repeat-containing 3 | 11637 | -0.106 | -0.4439 | No |
| 375 | CSTB | CSTB Entrez, Source | cystatin B (stefin B) | 11693 | -0.109 | -0.4464 | Yes |
| 376 | GATA6 | GATA6 Entrez, Source | GATA binding protein 6 | 11713 | -0.110 | -0.4462 | Yes |
| 377 | PTTG1IP | PTTG1IP Entrez, Source | pituitary tumor-transforming 1 interacting protein | 11723 | -0.110 | -0.4452 | Yes |
| 378 | CNN3 | CNN3 Entrez, Source | calponin 3, acidic | 11730 | -0.111 | -0.4439 | Yes |
| 379 | PRIM2A | PRIM2A Entrez, Source | primase, polypeptide 2A, 58kDa | 11731 | -0.111 | -0.4422 | Yes |
| 380 | TMSB10 | TMSB10 Entrez, Source | thymosin, beta 10 | 11750 | -0.111 | -0.4418 | Yes |
| 381 | CXADR | CXADR Entrez, Source | coxsackie virus and adenovirus receptor | 11788 | -0.113 | -0.4429 | Yes |
| 382 | ATP2C1 | ATP2C1 Entrez, Source | ATPase, Ca++ transporting, type 2C, member 1 | 11799 | -0.114 | -0.4419 | Yes |
| 383 | PSMB9 | PSMB9 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 9 (large multifunctional peptidase 2) | 11801 | -0.114 | -0.4402 | Yes |
| 384 | GNB4 | GNB4 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 4 | 11832 | -0.115 | -0.4407 | Yes |
| 385 | PHACTR2 | PHACTR2 Entrez, Source | phosphatase and actin regulator 2 | 11854 | -0.117 | -0.4405 | Yes |
| 386 | ST14 | ST14 Entrez, Source | suppression of tumorigenicity 14 (colon carcinoma) | 11861 | -0.117 | -0.4392 | Yes |
| 387 | TUBB6 | TUBB6 Entrez, Source | tubulin, beta 6 | 11867 | -0.117 | -0.4377 | Yes |
| 388 | PLA2G2A | PLA2G2A Entrez, Source | phospholipase A2, group IIA (platelets, synovial fluid) | 11878 | -0.118 | -0.4367 | Yes |
| 389 | MFGE8 | MFGE8 Entrez, Source | milk fat globule-EGF factor 8 protein | 11896 | -0.118 | -0.4361 | Yes |
| 390 | KLK3 | KLK3 Entrez, Source | kallikrein 3, (prostate specific antigen) | 11915 | -0.119 | -0.4356 | Yes |
| 391 | AURKA | AURKA Entrez, Source | aurora kinase A | 11917 | -0.119 | -0.4339 | Yes |
| 392 | DEK | DEK Entrez, Source | DEK oncogene (DNA binding) | 11951 | -0.122 | -0.4345 | Yes |
| 393 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 12036 | -0.127 | -0.4391 | Yes |
| 394 | IMPA2 | IMPA2 Entrez, Source | inositol(myo)-1(or 4)-monophosphatase 2 | 12041 | -0.127 | -0.4374 | Yes |
| 395 | UCHL1 | UCHL1 Entrez, Source | ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) | 12088 | -0.130 | -0.4389 | Yes |
| 396 | RSRC1 | RSRC1 Entrez, Source | arginine/serine-rich coiled-coil 1 | 12104 | -0.132 | -0.4380 | Yes |
| 397 | KLF5 | KLF5 Entrez, Source | Kruppel-like factor 5 (intestinal) | 12107 | -0.132 | -0.4361 | Yes |
| 398 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 12113 | -0.132 | -0.4344 | Yes |
| 399 | RGS10 | RGS10 Entrez, Source | regulator of G-protein signalling 10 | 12133 | -0.133 | -0.4338 | Yes |
| 400 | CSRP2 | CSRP2 Entrez, Source | cysteine and glycine-rich protein 2 | 12164 | -0.135 | -0.4340 | Yes |
| 401 | MTF2 | MTF2 Entrez, Source | metal response element binding transcription factor 2 | 12173 | -0.137 | -0.4325 | Yes |
| 402 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 12198 | -0.139 | -0.4322 | Yes |
| 403 | S100A8 | S100A8 Entrez, Source | S100 calcium binding protein A8 | 12223 | -0.142 | -0.4318 | Yes |
| 404 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 12256 | -0.144 | -0.4321 | Yes |
| 405 | CRLF3 | CRLF3 Entrez, Source | cytokine receptor-like factor 3 | 12266 | -0.145 | -0.4305 | Yes |
| 406 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 12269 | -0.146 | -0.4284 | Yes |
| 407 | MX2 | MX2 Entrez, Source | myxovirus (influenza virus) resistance 2 (mouse) | 12287 | -0.147 | -0.4274 | Yes |
| 408 | S100A10 | S100A10 Entrez, Source | S100 calcium binding protein A10 | 12310 | -0.149 | -0.4268 | Yes |
| 409 | PLS3 | PLS3 Entrez, Source | plastin 3 (T isoform) | 12321 | -0.150 | -0.4252 | Yes |
| 410 | KHDRBS3 | KHDRBS3 Entrez, Source | KH domain containing, RNA binding, signal transduction associated 3 | 12325 | -0.150 | -0.4231 | Yes |
| 411 | ITGB4 | ITGB4 Entrez, Source | integrin, beta 4 | 12348 | -0.153 | -0.4224 | Yes |
| 412 | NOS3 | NOS3 Entrez, Source | nitric oxide synthase 3 (endothelial cell) | 12369 | -0.155 | -0.4215 | Yes |
| 413 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 12377 | -0.156 | -0.4196 | Yes |
| 414 | BIN1 | BIN1 Entrez, Source | bridging integrator 1 | 12397 | -0.157 | -0.4186 | Yes |
| 415 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 12413 | -0.160 | -0.4173 | Yes |
| 416 | PLA2G4A | PLA2G4A Entrez, Source | phospholipase A2, group IVA (cytosolic, calcium-dependent) | 12414 | -0.160 | -0.4148 | Yes |
| 417 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 12432 | -0.162 | -0.4136 | Yes |
| 418 | GGH | GGH Entrez, Source | gamma-glutamyl hydrolase (conjugase, folylpolygammaglutamyl hydrolase) | 12437 | -0.162 | -0.4113 | Yes |
| 419 | USP18 | USP18 Entrez, Source | ubiquitin specific peptidase 18 | 12457 | -0.165 | -0.4102 | Yes |
| 420 | MMP7 | MMP7 Entrez, Source | matrix metallopeptidase 7 (matrilysin, uterine) | 12465 | -0.166 | -0.4082 | Yes |
| 421 | RND3 | RND3 Entrez, Source | Rho family GTPase 3 | 12492 | -0.169 | -0.4076 | Yes |
| 422 | STK10 | STK10 Entrez, Source | serine/threonine kinase 10 | 12500 | -0.171 | -0.4054 | Yes |
| 423 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 12528 | -0.175 | -0.4048 | Yes |
| 424 | ARPC1B | ARPC1B Entrez, Source | actin related protein 2/3 complex, subunit 1B, 41kDa | 12548 | -0.178 | -0.4035 | Yes |
| 425 | IL1R2 | IL1R2 Entrez, Source | interleukin 1 receptor, type II | 12559 | -0.179 | -0.4014 | Yes |
| 426 | CX3CL1 | CX3CL1 Entrez, Source | chemokine (C-X3-C motif) ligand 1 | 12570 | -0.180 | -0.3994 | Yes |
| 427 | CLIC4 | CLIC4 Entrez, Source | chloride intracellular channel 4 | 12577 | -0.181 | -0.3970 | Yes |
| 428 | TRIM2 | TRIM2 Entrez, Source | tripartite motif-containing 2 | 12579 | -0.181 | -0.3942 | Yes |
| 429 | GLS | GLS Entrez, Source | glutaminase | 12588 | -0.183 | -0.3920 | Yes |
| 430 | KIF3C | KIF3C Entrez, Source | kinesin family member 3C | 12599 | -0.184 | -0.3899 | Yes |
| 431 | YWHAQ | YWHAQ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta polypeptide | 12601 | -0.184 | -0.3871 | Yes |
| 432 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 12607 | -0.185 | -0.3846 | Yes |
| 433 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 12628 | -0.189 | -0.3832 | Yes |
| 434 | NMT2 | NMT2 Entrez, Source | N-myristoyltransferase 2 | 12660 | -0.196 | -0.3825 | Yes |
| 435 | POLA2 | POLA2 Entrez, Source | polymerase (DNA directed), alpha 2 (70kD subunit) | 12673 | -0.198 | -0.3803 | Yes |
| 436 | CAV2 | CAV2 Entrez, Source | caveolin 2 | 12694 | -0.203 | -0.3787 | Yes |
| 437 | PRKCI | PRKCI Entrez, Source | protein kinase C, iota | 12716 | -0.208 | -0.3771 | Yes |
| 438 | SNN | SNN Entrez, Source | stannin | 12747 | -0.213 | -0.3761 | Yes |
| 439 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 12752 | -0.214 | -0.3730 | Yes |
| 440 | KIFC1 | KIFC1 Entrez, Source | kinesin family member C1 | 12767 | -0.216 | -0.3707 | Yes |
| 441 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 12815 | -0.228 | -0.3708 | Yes |
| 442 | CDCA8 | CDCA8 Entrez, Source | cell division cycle associated 8 | 12816 | -0.228 | -0.3672 | Yes |
| 443 | CAPG | CAPG Entrez, Source | capping protein (actin filament), gelsolin-like | 12818 | -0.229 | -0.3637 | Yes |
| 444 | DTX2 | DTX2 Entrez, Source | deltex homolog 2 (Drosophila) | 12845 | -0.236 | -0.3620 | Yes |
| 445 | TRIP13 | TRIP13 Entrez, Source | thyroid hormone receptor interactor 13 | 12851 | -0.238 | -0.3587 | Yes |
| 446 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 12889 | -0.247 | -0.3577 | Yes |
| 447 | CEP170 | CEP170 Entrez, Source | centrosomal protein 170kDa | 12890 | -0.249 | -0.3538 | Yes |
| 448 | CYBA | CYBA Entrez, Source | cytochrome b-245, alpha polypeptide | 12921 | -0.260 | -0.3520 | Yes |
| 449 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 12930 | -0.262 | -0.3486 | Yes |
| 450 | IFI44 | IFI44 Entrez, Source | interferon-induced protein 44 | 12935 | -0.264 | -0.3447 | Yes |
| 451 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 12949 | -0.272 | -0.3415 | Yes |
| 452 | SLC2A1 | SLC2A1 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 1 | 12962 | -0.277 | -0.3381 | Yes |
| 453 | SPBC25 | SPBC25 Entrez, Source | spindle pole body component 25 homolog (S. cerevisiae) | 12974 | -0.280 | -0.3345 | Yes |
| 454 | SLC16A3 | SLC16A3 Entrez, Source | solute carrier family 16, member 3 (monocarboxylic acid transporter 4) | 12983 | -0.284 | -0.3307 | Yes |
| 455 | HN1 | HN1 Entrez, Source | hematological and neurological expressed 1 | 13029 | -0.307 | -0.3294 | Yes |
| 456 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 13034 | -0.309 | -0.3249 | Yes |
| 457 | PRNP | PRNP Entrez, Source | prion protein (p27-30) (Creutzfeldt-Jakob disease, Gerstmann-Strausler-Scheinker syndrome, fatal familial insomnia) | 13038 | -0.313 | -0.3202 | Yes |
| 458 | RRAS2 | RRAS2 Entrez, Source | related RAS viral (r-ras) oncogene homolog 2 | 13055 | -0.322 | -0.3164 | Yes |
| 459 | TMEM123 | TMEM123 Entrez, Source | transmembrane protein 123 | 13062 | -0.326 | -0.3117 | Yes |
| 460 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 13069 | -0.331 | -0.3070 | Yes |
| 461 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 13078 | -0.337 | -0.3024 | Yes |
| 462 | GPNMB | GPNMB Entrez, Source | glycoprotein (transmembrane) nmb | 13090 | -0.347 | -0.2978 | Yes |
| 463 | S100A11 | S100A11 Entrez, Source | S100 calcium binding protein A11 | 13099 | -0.355 | -0.2928 | Yes |
| 464 | ADORA2B | ADORA2B Entrez, Source | adenosine A2b receptor | 13103 | -0.358 | -0.2874 | Yes |
| 465 | AURKB | AURKB Entrez, Source | aurora kinase B | 13115 | -0.366 | -0.2826 | Yes |
| 466 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 13129 | -0.381 | -0.2776 | Yes |
| 467 | PROM1 | PROM1 Entrez, Source | prominin 1 | 13138 | -0.389 | -0.2721 | Yes |
| 468 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 13153 | -0.404 | -0.2669 | Yes |
| 469 | S100A6 | S100A6 Entrez, Source | S100 calcium binding protein A6 | 13165 | -0.424 | -0.2611 | Yes |
| 470 | LCP1 | LCP1 Entrez, Source | lymphocyte cytosolic protein 1 (L-plastin) | 13166 | -0.425 | -0.2544 | Yes |
| 471 | EDN1 | EDN1 Entrez, Source | endothelin 1 | 13172 | -0.431 | -0.2480 | Yes |
| 472 | CD44 | CD44 Entrez, Source | CD44 molecule (Indian blood group) | 13174 | -0.431 | -0.2414 | Yes |
| 473 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 13180 | -0.438 | -0.2349 | Yes |
| 474 | FABP7 | FABP7 Entrez, Source | fatty acid binding protein 7, brain | 13182 | -0.442 | -0.2280 | Yes |
| 475 | VIM | VIM Entrez, Source | vimentin | 13227 | -0.506 | -0.2235 | Yes |
| 476 | FXYD5 | FXYD5 Entrez, Source | FXYD domain containing ion transport regulator 5 | 13236 | -0.522 | -0.2160 | Yes |
| 477 | SOAT1 | SOAT1 Entrez, Source | sterol O-acyltransferase (acyl-Coenzyme A: cholesterol acyltransferase) 1 | 13248 | -0.551 | -0.2082 | Yes |
| 478 | LAMC2 | LAMC2 Entrez, Source | laminin, gamma 2 | 13264 | -0.594 | -0.2000 | Yes |
| 479 | DUSP9 | DUSP9 Entrez, Source | dual specificity phosphatase 9 | 13265 | -0.606 | -0.1905 | Yes |
| 480 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 13266 | -0.607 | -0.1810 | Yes |
| 481 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13267 | -0.614 | -0.1714 | Yes |
| 482 | GBP2 | GBP2 Entrez, Source | guanylate binding protein 2, interferon-inducible | 13270 | -0.639 | -0.1615 | Yes |
| 483 | FBLIM1 | FBLIM1 Entrez, Source | filamin binding LIM protein 1 | 13284 | -0.687 | -0.1517 | Yes |
| 484 | ADM | ADM Entrez, Source | adrenomedullin | 13287 | -0.697 | -0.1410 | Yes |
| 485 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 13300 | -0.747 | -0.1302 | Yes |
| 486 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 13302 | -0.755 | -0.1184 | Yes |
| 487 | TIMP2 | TIMP2 Entrez, Source | TIMP metallopeptidase inhibitor 2 | 13306 | -0.786 | -0.1063 | Yes |
| 488 | CD69 | CD69 Entrez, Source | CD69 molecule | 13310 | -0.833 | -0.0935 | Yes |
| 489 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13314 | -0.890 | -0.0798 | Yes |
| 490 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 13317 | -0.925 | -0.0655 | Yes |
| 491 | DBN1 | DBN1 Entrez, Source | drebrin 1 | 13318 | -0.939 | -0.0507 | Yes |
| 492 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 13321 | -0.988 | -0.0354 | Yes |
| 493 | MSN | MSN Entrez, Source | moesin | 13329 | -1.080 | -0.0190 | Yes |
| 494 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 13336 | -1.272 | 0.0005 | Yes |

