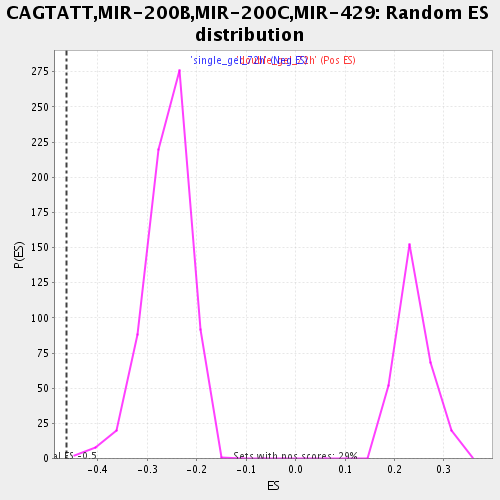
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | CAGTATT,MIR-200B,MIR-200C,MIR-429 |
| Enrichment Score (ES) | -0.46328148 |
| Normalized Enrichment Score (NES) | -1.78576 |
| Nominal p-value | 0.0014144272 |
| FDR q-value | 0.039636172 |
| FWER p-Value | 0.987 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FGFR2 | FGFR2 Entrez, Source | ffer syndrome, Jackson-Weiss syndrome) | 84 | 0.431 | 0.0126 | No |
| 2 | LEPR | LEPR Entrez, Source | leptin receptor | 146 | 0.357 | 0.0238 | No |
| 3 | NR5A2 | NR5A2 Entrez, Source | nuclear receptor subfamily 5, group A, member 2 | 206 | 0.307 | 0.0328 | No |
| 4 | ACACA | ACACA Entrez, Source | acetyl-Coenzyme A carboxylase alpha | 231 | 0.293 | 0.0440 | No |
| 5 | LRP4 | LRP4 Entrez, Source | low density lipoprotein receptor-related protein 4 | 323 | 0.258 | 0.0484 | No |
| 6 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 329 | 0.257 | 0.0594 | No |
| 7 | MGAT2 | MGAT2 Entrez, Source | mannosyl (alpha-1,6-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase | 671 | 0.177 | 0.0413 | No |
| 8 | SDC2 | SDC2 Entrez, Source | syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated, fibroglycan) | 817 | 0.161 | 0.0373 | No |
| 9 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 901 | 0.153 | 0.0378 | No |
| 10 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 1078 | 0.138 | 0.0305 | No |
| 11 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 1291 | 0.125 | 0.0198 | No |
| 12 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 1483 | 0.114 | 0.0103 | No |
| 13 | KLF9 | KLF9 Entrez, Source | Kruppel-like factor 9 | 1556 | 0.110 | 0.0097 | No |
| 14 | TUBB3 | TUBB3 Entrez, Source | tubulin, beta 3 | 1557 | 0.110 | 0.0146 | No |
| 15 | PCDH8 | PCDH8 Entrez, Source | protocadherin 8 | 1627 | 0.107 | 0.0141 | No |
| 16 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 1693 | 0.105 | 0.0138 | No |
| 17 | SLC6A1 | SLC6A1 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, GABA), member 1 | 1847 | 0.100 | 0.0065 | No |
| 18 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 1923 | 0.097 | 0.0051 | No |
| 19 | NUDT4 | NUDT4 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 4 | 1982 | 0.094 | 0.0048 | No |
| 20 | SYVN1 | SYVN1 Entrez, Source | synovial apoptosis inhibitor 1, synoviolin | 2057 | 0.092 | 0.0033 | No |
| 21 | NOG | NOG Entrez, Source | noggin | 2077 | 0.091 | 0.0059 | No |
| 22 | ITPR1 | ITPR1 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 1 | 2186 | 0.089 | 0.0016 | No |
| 23 | LRP1 | LRP1 Entrez, Source | low density lipoprotein-related protein 1 (alpha-2-macroglobulin receptor) | 2212 | 0.088 | 0.0035 | No |
| 24 | MAFG | MAFG Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog G (avian) | 2226 | 0.087 | 0.0064 | No |
| 25 | ACE2 | ACE2 Entrez, Source | angiotensin I converting enzyme (peptidyl-dipeptidase A) 2 | 2418 | 0.082 | -0.0045 | No |
| 26 | ADIPOR2 | ADIPOR2 Entrez, Source | adiponectin receptor 2 | 2598 | 0.076 | -0.0148 | No |
| 27 | CNTN4 | CNTN4 Entrez, Source | contactin 4 | 2609 | 0.076 | -0.0122 | No |
| 28 | NDST1 | NDST1 Entrez, Source | N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 1 | 2645 | 0.075 | -0.0115 | No |
| 29 | MTUS1 | MTUS1 Entrez, Source | mitochondrial tumor suppressor 1 | 2646 | 0.075 | -0.0082 | No |
| 30 | XKR6 | XKR6 Entrez, Source | XK, Kell blood group complex subunit-related family, member 6 | 2668 | 0.074 | -0.0065 | No |
| 31 | NBR1 | NBR1 Entrez, Source | neighbor of BRCA1 gene 1 | 2837 | 0.071 | -0.0162 | No |
| 32 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 2854 | 0.070 | -0.0143 | No |
| 33 | ANKRD15 | ANKRD15 Entrez, Source | ankyrin repeat domain 15 | 2894 | 0.069 | -0.0143 | No |
| 34 | AKAP7 | AKAP7 Entrez, Source | A kinase (PRKA) anchor protein 7 | 2928 | 0.068 | -0.0138 | No |
| 35 | TEAD1 | TEAD1 Entrez, Source | TEA domain family member 1 (SV40 transcriptional enhancer factor) | 3114 | 0.064 | -0.0250 | No |
| 36 | UBQLN1 | UBQLN1 Entrez, Source | ubiquilin 1 | 3176 | 0.063 | -0.0269 | No |
| 37 | HS3ST1 | HS3ST1 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 1 | 3289 | 0.060 | -0.0327 | No |
| 38 | GOSR2 | GOSR2 Entrez, Source | golgi SNAP receptor complex member 2 | 3338 | 0.059 | -0.0338 | No |
| 39 | CNTFR | CNTFR Entrez, Source | ciliary neurotrophic factor receptor | 3340 | 0.059 | -0.0313 | No |
| 40 | EFNA1 | EFNA1 Entrez, Source | ephrin-A1 | 3378 | 0.058 | -0.0315 | No |
| 41 | GATA2 | GATA2 Entrez, Source | GATA binding protein 2 | 3541 | 0.055 | -0.0414 | No |
| 42 | RPS6KB1 | RPS6KB1 Entrez, Source | ribosomal protein S6 kinase, 70kDa, polypeptide 1 | 3665 | 0.053 | -0.0484 | No |
| 43 | MXD3 | MXD3 Entrez, Source | MAX dimerization protein 3 | 3701 | 0.052 | -0.0488 | No |
| 44 | UBE2I | UBE2I Entrez, Source | ubiquitin-conjugating enzyme E2I (UBC9 homolog, yeast) | 3706 | 0.052 | -0.0468 | No |
| 45 | COPS8 | COPS8 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 8 (Arabidopsis) | 3778 | 0.050 | -0.0500 | No |
| 46 | CDH11 | CDH11 Entrez, Source | cadherin 11, type 2, OB-cadherin (osteoblast) | 3828 | 0.049 | -0.0516 | No |
| 47 | ERRFI1 | ERRFI1 Entrez, Source | ERBB receptor feedback inhibitor 1 | 3954 | 0.047 | -0.0590 | No |
| 48 | STX1A | STX1A Entrez, Source | syntaxin 1A (brain) | 3958 | 0.047 | -0.0571 | No |
| 49 | PPP1R10 | PPP1R10 Entrez, Source | protein phosphatase 1, regulatory subunit 10 | 4035 | 0.046 | -0.0609 | No |
| 50 | PSIP1 | PSIP1 Entrez, Source | PC4 and SFRS1 interacting protein 1 | 4049 | 0.045 | -0.0599 | No |
| 51 | GLRA2 | GLRA2 Entrez, Source | glycine receptor, alpha 2 | 4090 | 0.045 | -0.0610 | No |
| 52 | SLC38A4 | SLC38A4 Entrez, Source | solute carrier family 38, member 4 | 4109 | 0.044 | -0.0604 | No |
| 53 | CACNA1C | CACNA1C Entrez, Source | calcium channel, voltage-dependent, L type, alpha 1C subunit | 4205 | 0.043 | -0.0657 | No |
| 54 | EDG2 | EDG2 Entrez, Source | endothelial differentiation, lysophosphatidic acid G-protein-coupled receptor, 2 | 4234 | 0.042 | -0.0660 | No |
| 55 | NAB1 | NAB1 Entrez, Source | NGFI-A binding protein 1 (EGR1 binding protein 1) | 4286 | 0.041 | -0.0680 | No |
| 56 | TBP | TBP Entrez, Source | TATA box binding protein | 4369 | 0.040 | -0.0725 | No |
| 57 | OTUD4 | OTUD4 Entrez, Source | OTU domain containing 4 | 4532 | 0.037 | -0.0832 | No |
| 58 | PPM1B | PPM1B Entrez, Source | protein phosphatase 1B (formerly 2C), magnesium-dependent, beta isoform | 4644 | 0.035 | -0.0901 | No |
| 59 | EIF2B5 | EIF2B5 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 5 epsilon, 82kDa | 4827 | 0.032 | -0.1025 | No |
| 60 | NFIA | NFIA Entrez, Source | nuclear factor I/A | 4861 | 0.032 | -0.1037 | No |
| 61 | GATA4 | GATA4 Entrez, Source | GATA binding protein 4 | 5021 | 0.029 | -0.1145 | No |
| 62 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 5030 | 0.029 | -0.1139 | No |
| 63 | RPS6KA2 | RPS6KA2 Entrez, Source | ribosomal protein S6 kinase, 90kDa, polypeptide 2 | 5082 | 0.028 | -0.1165 | No |
| 64 | PTHLH | PTHLH Entrez, Source | parathyroid hormone-like hormone | 5115 | 0.027 | -0.1177 | No |
| 65 | FREQ | FREQ Entrez, Source | frequenin homolog (Drosophila) | 5177 | 0.027 | -0.1212 | No |
| 66 | NR3C1 | NR3C1 Entrez, Source | nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) | 5252 | 0.025 | -0.1257 | No |
| 67 | SLC14A1 | SLC14A1 Entrez, Source | solute carrier family 14 (urea transporter), member 1 (Kidd blood group) | 5281 | 0.025 | -0.1267 | No |
| 68 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 5332 | 0.024 | -0.1295 | No |
| 69 | LIN7B | LIN7B Entrez, Source | lin-7 homolog B (C. elegans) | 5360 | 0.024 | -0.1305 | No |
| 70 | EIF5B | EIF5B Entrez, Source | eukaryotic translation initiation factor 5B | 5435 | 0.023 | -0.1351 | No |
| 71 | SNAP25 | SNAP25 Entrez, Source | synaptosomal-associated protein, 25kDa | 5580 | 0.021 | -0.1452 | No |
| 72 | NCOA2 | NCOA2 Entrez, Source | nuclear receptor coactivator 2 | 5627 | 0.020 | -0.1478 | No |
| 73 | SULF1 | SULF1 Entrez, Source | sulfatase 1 | 5633 | 0.020 | -0.1473 | No |
| 74 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 5972 | 0.015 | -0.1724 | No |
| 75 | RSN | RSN Entrez, Source | restin (Reed-Steinberg cell-expressed intermediate filament-associated protein) | 6102 | 0.013 | -0.1817 | No |
| 76 | SLC1A2 | SLC1A2 Entrez, Source | solute carrier family 1 (glial high affinity glutamate transporter), member 2 | 6115 | 0.013 | -0.1820 | No |
| 77 | EIF2S1 | EIF2S1 Entrez, Source | eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa | 6150 | 0.012 | -0.1841 | No |
| 78 | KCNJ2 | KCNJ2 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 2 | 6345 | 0.009 | -0.1985 | No |
| 79 | ADCY2 | ADCY2 Entrez, Source | adenylate cyclase 2 (brain) | 6373 | 0.008 | -0.2002 | No |
| 80 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 6567 | 0.006 | -0.2146 | No |
| 81 | SCN3B | SCN3B Entrez, Source | sodium channel, voltage-gated, type III, beta | 6584 | 0.005 | -0.2156 | No |
| 82 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 6673 | 0.004 | -0.2221 | No |
| 83 | KCNK2 | KCNK2 Entrez, Source | potassium channel, subfamily K, member 2 | 6765 | 0.003 | -0.2289 | No |
| 84 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 6773 | 0.003 | -0.2293 | No |
| 85 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 6801 | 0.003 | -0.2312 | No |
| 86 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 6818 | 0.002 | -0.2323 | No |
| 87 | NPM1 | NPM1 Entrez, Source | nucleophosmin (nucleolar phosphoprotein B23, numatrin) | 7074 | -0.001 | -0.2517 | No |
| 88 | GIT2 | GIT2 Entrez, Source | G protein-coupled receptor kinase interactor 2 | 7104 | -0.002 | -0.2538 | No |
| 89 | NOVA1 | NOVA1 Entrez, Source | neuro-oncological ventral antigen 1 | 7115 | -0.002 | -0.2545 | No |
| 90 | PDPK1 | PDPK1 Entrez, Source | 3-phosphoinositide dependent protein kinase-1 | 7143 | -0.002 | -0.2565 | No |
| 91 | TCEB1 | TCEB1 Entrez, Source | transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) | 7156 | -0.002 | -0.2573 | No |
| 92 | ICK | ICK Entrez, Source | intestinal cell (MAK-like) kinase | 7184 | -0.003 | -0.2592 | No |
| 93 | MAP4K3 | MAP4K3 Entrez, Source | mitogen-activated protein kinase kinase kinase kinase 3 | 7583 | -0.009 | -0.2891 | No |
| 94 | CRKL | CRKL Entrez, Source | v-crk sarcoma virus CT10 oncogene homolog (avian)-like | 7585 | -0.009 | -0.2888 | No |
| 95 | NDN | NDN Entrez, Source | necdin homolog (mouse) | 7672 | -0.010 | -0.2949 | No |
| 96 | HLF | HLF Entrez, Source | hepatic leukemia factor | 7695 | -0.010 | -0.2961 | No |
| 97 | SERINC1 | SERINC1 Entrez, Source | serine incorporator 1 | 7788 | -0.012 | -0.3026 | No |
| 98 | LFNG | LFNG Entrez, Source | lunatic fringe homolog (Drosophila) | 7939 | -0.014 | -0.3134 | No |
| 99 | ACVR2A | ACVR2A Entrez, Source | activin A receptor, type IIA | 7984 | -0.015 | -0.3161 | No |
| 100 | EGLN1 | EGLN1 Entrez, Source | egl nine homolog 1 (C. elegans) | 8000 | -0.015 | -0.3166 | No |
| 101 | NRP2 | NRP2 Entrez, Source | neuropilin 2 | 8004 | -0.015 | -0.3162 | No |
| 102 | RDH10 | RDH10 Entrez, Source | retinol dehydrogenase 10 (all-trans) | 8061 | -0.016 | -0.3197 | No |
| 103 | MATR3 | MATR3 Entrez, Source | matrin 3 | 8066 | -0.016 | -0.3193 | No |
| 104 | SCAMP1 | SCAMP1 Entrez, Source | secretory carrier membrane protein 1 | 8178 | -0.018 | -0.3270 | No |
| 105 | UBE2B | UBE2B Entrez, Source | ubiquitin-conjugating enzyme E2B (RAD6 homolog) | 8205 | -0.018 | -0.3282 | No |
| 106 | PTBP1 | PTBP1 Entrez, Source | polypyrimidine tract binding protein 1 | 8339 | -0.020 | -0.3374 | No |
| 107 | DNAJB6 | DNAJB6 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 6 | 8356 | -0.021 | -0.3377 | No |
| 108 | BAT3 | BAT3 Entrez, Source | HLA-B associated transcript 3 | 8461 | -0.022 | -0.3447 | No |
| 109 | FGD1 | FGD1 Entrez, Source | FYVE, RhoGEF and PH domain containing 1 (faciogenital dysplasia) | 8576 | -0.024 | -0.3523 | No |
| 110 | PCTK1 | PCTK1 Entrez, Source | PCTAIRE protein kinase 1 | 8584 | -0.024 | -0.3517 | No |
| 111 | HNRPU | HNRPU Entrez, Source | heterogeneous nuclear ribonucleoprotein U (scaffold attachment factor A) | 8601 | -0.025 | -0.3518 | No |
| 112 | GOLGA7 | GOLGA7 Entrez, Source | golgi autoantigen, golgin subfamily a, 7 | 8621 | -0.025 | -0.3522 | No |
| 113 | ARHGDIA | ARHGDIA Entrez, Source | Rho GDP dissociation inhibitor (GDI) alpha | 8656 | -0.026 | -0.3536 | No |
| 114 | OGT | OGT Entrez, Source | O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) | 8678 | -0.026 | -0.3541 | No |
| 115 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 8693 | -0.026 | -0.3540 | No |
| 116 | CSNK1G3 | CSNK1G3 Entrez, Source | casein kinase 1, gamma 3 | 8705 | -0.026 | -0.3537 | No |
| 117 | OXR1 | OXR1 Entrez, Source | oxidation resistance 1 | 8815 | -0.028 | -0.3607 | No |
| 118 | YWHAB | YWHAB Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, beta polypeptide | 8834 | -0.029 | -0.3608 | No |
| 119 | KCND2 | KCND2 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 2 | 8996 | -0.032 | -0.3717 | No |
| 120 | GDI2 | GDI2 Entrez, Source | GDP dissociation inhibitor 2 | 9090 | -0.034 | -0.3772 | No |
| 121 | CRH | CRH Entrez, Source | corticotropin releasing hormone | 9147 | -0.035 | -0.3800 | No |
| 122 | DNMT3A | DNMT3A Entrez, Source | DNA (cytosine-5-)-methyltransferase 3 alpha | 9156 | -0.035 | -0.3790 | No |
| 123 | NFYA | NFYA Entrez, Source | nuclear transcription factor Y, alpha | 9205 | -0.036 | -0.3811 | No |
| 124 | NEGR1 | NEGR1 Entrez, Source | neuronal growth regulator 1 | 9235 | -0.036 | -0.3817 | No |
| 125 | BIRC4 | BIRC4 Entrez, Source | baculoviral IAP repeat-containing 4 | 9335 | -0.038 | -0.3875 | No |
| 126 | FMR1 | FMR1 Entrez, Source | fragile X mental retardation 1 | 9448 | -0.041 | -0.3943 | No |
| 127 | GNAI3 | GNAI3 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 3 | 9496 | -0.042 | -0.3960 | No |
| 128 | ESRRG | ESRRG Entrez, Source | estrogen-related receptor gamma | 9512 | -0.042 | -0.3953 | No |
| 129 | SLC38A2 | SLC38A2 Entrez, Source | solute carrier family 38, member 2 | 9549 | -0.043 | -0.3962 | No |
| 130 | LYPLA2 | LYPLA2 Entrez, Source | lysophospholipase II | 9600 | -0.043 | -0.3981 | No |
| 131 | PPP1R9B | PPP1R9B Entrez, Source | protein phosphatase 1, regulatory subunit 9B, spinophilin | 9743 | -0.047 | -0.4068 | No |
| 132 | ATP2A2 | ATP2A2 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 | 9784 | -0.048 | -0.4078 | No |
| 133 | FHL1 | FHL1 Entrez, Source | four and a half LIM domains 1 | 9787 | -0.048 | -0.4058 | No |
| 134 | NTF3 | NTF3 Entrez, Source | neurotrophin 3 | 9863 | -0.049 | -0.4094 | No |
| 135 | KRAS | KRAS Entrez, Source | v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog | 9882 | -0.049 | -0.4085 | No |
| 136 | GLI3 | GLI3 Entrez, Source | GLI-Kruppel family member GLI3 (Greig cephalopolysyndactyly syndrome) | 9905 | -0.050 | -0.4080 | No |
| 137 | PPP2CA | PPP2CA Entrez, Source | protein phosphatase 2 (formerly 2A), catalytic subunit, alpha isoform | 9927 | -0.051 | -0.4074 | No |
| 138 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 9940 | -0.051 | -0.4060 | No |
| 139 | FEZ2 | FEZ2 Entrez, Source | fasciculation and elongation protein zeta 2 (zygin II) | 9964 | -0.051 | -0.4055 | No |
| 140 | RAB7 | RAB7 Entrez, Source | RAB7, member RAS oncogene family | 10016 | -0.052 | -0.4071 | No |
| 141 | PUM2 | PUM2 Entrez, Source | pumilio homolog 2 (Drosophila) | 10034 | -0.053 | -0.4060 | No |
| 142 | FN1 | FN1 Entrez, Source | fibronectin 1 | 10062 | -0.053 | -0.4057 | No |
| 143 | MARCKS | MARCKS Entrez, Source | myristoylated alanine-rich protein kinase C substrate | 10092 | -0.054 | -0.4056 | No |
| 144 | HNRPK | HNRPK Entrez, Source | heterogeneous nuclear ribonucleoprotein K | 10138 | -0.055 | -0.4065 | No |
| 145 | CNOT4 | CNOT4 Entrez, Source | CCR4-NOT transcription complex, subunit 4 | 10146 | -0.055 | -0.4046 | No |
| 146 | TSGA14 | TSGA14 Entrez, Source | testis specific, 14 | 10166 | -0.056 | -0.4036 | No |
| 147 | PCAF | PCAF Entrez, Source | p300/CBP-associated factor | 10191 | -0.056 | -0.4030 | No |
| 148 | PLCL1 | PLCL1 Entrez, Source | phospholipase C-like 1 | 10259 | -0.058 | -0.4055 | No |
| 149 | ARHGAP20 | ARHGAP20 Entrez, Source | Rho GTPase activating protein 20 | 10392 | -0.061 | -0.4128 | No |
| 150 | PKIA | PKIA Entrez, Source | protein kinase (cAMP-dependent, catalytic) inhibitor alpha | 10441 | -0.063 | -0.4137 | No |
| 151 | CASR | CASR Entrez, Source | calcium-sensing receptor (hypocalciuric hypercalcemia 1, severe neonatal hyperparathyroidism) | 10471 | -0.064 | -0.4131 | No |
| 152 | RAC1 | RAC1 Entrez, Source | ras-related C3 botulinum toxin substrate 1 (rho family, small GTP binding protein Rac1) | 10733 | -0.071 | -0.4299 | No |
| 153 | SYNJ1 | SYNJ1 Entrez, Source | synaptojanin 1 | 10861 | -0.074 | -0.4363 | No |
| 154 | TNFRSF11B | TNFRSF11B Entrez, Source | tumor necrosis factor receptor superfamily, member 11b (osteoprotegerin) | 11055 | -0.080 | -0.4474 | No |
| 155 | IHPK1 | IHPK1 Entrez, Source | inositol hexaphosphate kinase 1 | 11145 | -0.083 | -0.4505 | No |
| 156 | ELF2 | ELF2 Entrez, Source | E74-like factor 2 (ets domain transcription factor) | 11297 | -0.090 | -0.4580 | No |
| 157 | YWHAG | YWHAG Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide | 11367 | -0.093 | -0.4592 | Yes |
| 158 | PKD1 | PKD1 Entrez, Source | polycystic kidney disease 1 (autosomal dominant) | 11375 | -0.094 | -0.4556 | Yes |
| 159 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 11431 | -0.096 | -0.4555 | Yes |
| 160 | RHOA | RHOA Entrez, Source | ras homolog gene family, member A | 11466 | -0.098 | -0.4537 | Yes |
| 161 | KHDRBS1 | KHDRBS1 Entrez, Source | KH domain containing, RNA binding, signal transduction associated 1 | 11520 | -0.100 | -0.4533 | Yes |
| 162 | RAB21 | RAB21 Entrez, Source | RAB21, member RAS oncogene family | 11573 | -0.103 | -0.4528 | Yes |
| 163 | BCAP29 | BCAP29 Entrez, Source | B-cell receptor-associated protein 29 | 11617 | -0.105 | -0.4514 | Yes |
| 164 | TCF8 | TCF8 Entrez, Source | transcription factor 8 (represses interleukin 2 expression) | 11658 | -0.107 | -0.4497 | Yes |
| 165 | PLK2 | PLK2 Entrez, Source | polo-like kinase 2 (Drosophila) | 11671 | -0.108 | -0.4458 | Yes |
| 166 | SLC23A2 | SLC23A2 Entrez, Source | solute carrier family 23 (nucleobase transporters), member 2 | 11687 | -0.108 | -0.4422 | Yes |
| 167 | PPP2R2C | PPP2R2C Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), gamma isoform | 11700 | -0.109 | -0.4383 | Yes |
| 168 | TCF2 | TCF2 Entrez, Source | transcription factor 2, hepatic; LF-B3; variant hepatic nuclear factor | 11711 | -0.110 | -0.4342 | Yes |
| 169 | WASPIP | WASPIP Entrez, Source | Wiskott-Aldrich syndrome protein interacting protein | 11720 | -0.110 | -0.4300 | Yes |
| 170 | CNN3 | CNN3 Entrez, Source | calponin 3, acidic | 11730 | -0.111 | -0.4258 | Yes |
| 171 | NXPH1 | NXPH1 Entrez, Source | neurexophilin 1 | 11738 | -0.111 | -0.4214 | Yes |
| 172 | CLASP2 | CLASP2 Entrez, Source | cytoplasmic linker associated protein 2 | 11749 | -0.111 | -0.4172 | Yes |
| 173 | DAG1 | DAG1 Entrez, Source | dystroglycan 1 (dystrophin-associated glycoprotein 1) | 11814 | -0.114 | -0.4171 | Yes |
| 174 | IER5 | IER5 Entrez, Source | immediate early response 5 | 11841 | -0.116 | -0.4139 | Yes |
| 175 | ARID4B | ARID4B Entrez, Source | AT rich interactive domain 4B (RBP1- like) | 12049 | -0.128 | -0.4240 | Yes |
| 176 | RAP1B | RAP1B Entrez, Source | RAP1B, member of RAS oncogene family | 12148 | -0.134 | -0.4256 | Yes |
| 177 | PHLDB1 | PHLDB1 Entrez, Source | pleckstrin homology-like domain, family B, member 1 | 12160 | -0.135 | -0.4204 | Yes |
| 178 | QKI | QKI Entrez, Source | quaking homolog, KH domain RNA binding (mouse) | 12184 | -0.138 | -0.4161 | Yes |
| 179 | DGKA | DGKA Entrez, Source | diacylglycerol kinase, alpha 80kDa | 12191 | -0.138 | -0.4104 | Yes |
| 180 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 12198 | -0.139 | -0.4048 | Yes |
| 181 | HS2ST1 | HS2ST1 Entrez, Source | heparan sulfate 2-O-sulfotransferase 1 | 12206 | -0.140 | -0.3991 | Yes |
| 182 | PLS3 | PLS3 Entrez, Source | plastin 3 (T isoform) | 12321 | -0.150 | -0.4012 | Yes |
| 183 | SYT1 | SYT1 Entrez, Source | synaptotagmin I | 12373 | -0.155 | -0.3982 | Yes |
| 184 | PCTK2 | PCTK2 Entrez, Source | PCTAIRE protein kinase 2 | 12381 | -0.156 | -0.3918 | Yes |
| 185 | PPM1F | PPM1F Entrez, Source | protein phosphatase 1F (PP2C domain containing) | 12392 | -0.157 | -0.3856 | Yes |
| 186 | VLDLR | VLDLR Entrez, Source | very low density lipoprotein receptor | 12472 | -0.167 | -0.3843 | Yes |
| 187 | RND3 | RND3 Entrez, Source | Rho family GTPase 3 | 12492 | -0.169 | -0.3783 | Yes |
| 188 | PLEKHC1 | PLEKHC1 Entrez, Source | pleckstrin homology domain containing, family C (with FERM domain) member 1 | 12574 | -0.181 | -0.3765 | Yes |
| 189 | CLIC4 | CLIC4 Entrez, Source | chloride intracellular channel 4 | 12577 | -0.181 | -0.3686 | Yes |
| 190 | TRIM2 | TRIM2 Entrez, Source | tripartite motif-containing 2 | 12579 | -0.181 | -0.3607 | Yes |
| 191 | CLASP1 | CLASP1 Entrez, Source | cytoplasmic linker associated protein 1 | 12914 | -0.257 | -0.3748 | Yes |
| 192 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 12930 | -0.262 | -0.3643 | Yes |
| 193 | TUBB | TUBB Entrez, Source | tubulin, beta | 13001 | -0.292 | -0.3568 | Yes |
| 194 | LAMC1 | LAMC1 Entrez, Source | laminin, gamma 1 (formerly LAMB2) | 13018 | -0.300 | -0.3448 | Yes |
| 195 | PRKAR2B | PRKAR2B Entrez, Source | protein kinase, cAMP-dependent, regulatory, type II, beta | 13042 | -0.314 | -0.3326 | Yes |
| 196 | HNRPD | HNRPD Entrez, Source | heterogeneous nuclear ribonucleoprotein D (AU-rich element RNA binding protein 1, 37kDa) | 13048 | -0.319 | -0.3189 | Yes |
| 197 | PTPRZ1 | PTPRZ1 Entrez, Source | protein tyrosine phosphatase, receptor-type, Z polypeptide 1 | 13074 | -0.336 | -0.3060 | Yes |
| 198 | TSC22D1 | TSC22D1 Entrez, Source | TSC22 domain family, member 1 | 13081 | -0.341 | -0.2914 | Yes |
| 199 | SLC6A6 | SLC6A6 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, taurine), member 6 | 13097 | -0.354 | -0.2769 | Yes |
| 200 | AKAP2 | AKAP2 Entrez, Source | A kinase (PRKA) anchor protein 2 | 13107 | -0.362 | -0.2616 | Yes |
| 201 | BASP1 | BASP1 Entrez, Source | brain abundant, membrane attached signal protein 1 | 13112 | -0.366 | -0.2458 | Yes |
| 202 | CDR2 | CDR2 Entrez, Source | cerebellar degeneration-related protein 2, 62kDa | 13150 | -0.398 | -0.2310 | Yes |
| 203 | PAM | PAM Entrez, Source | peptidylglycine alpha-amidating monooxygenase | 13160 | -0.412 | -0.2135 | Yes |
| 204 | LCP1 | LCP1 Entrez, Source | lymphocyte cytosolic protein 1 (L-plastin) | 13166 | -0.425 | -0.1951 | Yes |
| 205 | CYLN2 | CYLN2 Entrez, Source | cytoplasmic linker 2 | 13179 | -0.436 | -0.1767 | Yes |
| 206 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 13180 | -0.438 | -0.1573 | Yes |
| 207 | LMO7 | LMO7 Entrez, Source | LIM domain 7 | 13209 | -0.477 | -0.1384 | Yes |
| 208 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 13211 | -0.481 | -0.1172 | Yes |
| 209 | GPM6A | GPM6A Entrez, Source | glycoprotein M6A | 13245 | -0.543 | -0.0958 | Yes |
| 210 | MSN | MSN Entrez, Source | moesin | 13329 | -1.080 | -0.0544 | Yes |
| 211 | BHLHB3 | BHLHB3 Entrez, Source | basic helix-loop-helix domain containing, class B, 3 | 13335 | -1.251 | 0.0005 | Yes |