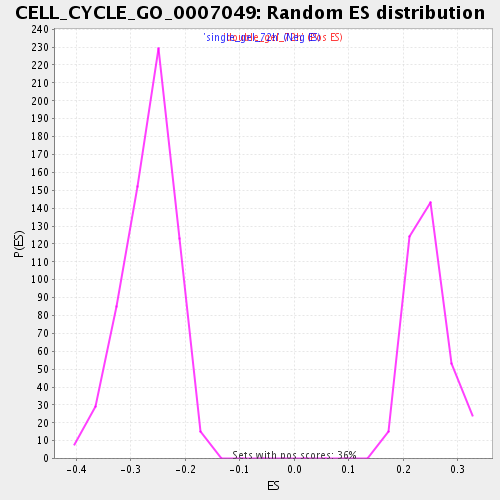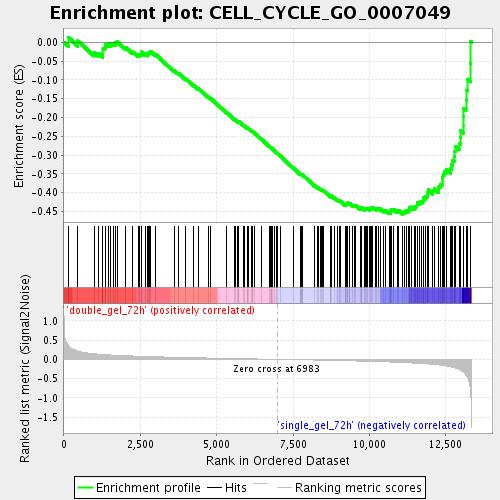
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | CELL_CYCLE_GO_0007049 |
| Enrichment Score (ES) | -0.45674753 |
| Normalized Enrichment Score (NES) | -1.7161803 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.057786725 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MDM4 | MDM4 Entrez, Source | Mdm4, transformed 3T3 cell double minute 4, p53 binding protein (mouse) | 141 | 0.366 | 0.0132 | No |
| 2 | CDK5R1 | CDK5R1 Entrez, Source | cyclin-dependent kinase 5, regulatory subunit 1 (p35) | 455 | 0.214 | 0.0035 | No |
| 3 | TGFA | TGFA Entrez, Source | transforming growth factor, alpha | 986 | 0.146 | -0.0271 | No |
| 4 | CENTD2 | CENTD2 Entrez, Source | centaurin, delta 2 | 1137 | 0.135 | -0.0297 | No |
| 5 | INHA | INHA Entrez, Source | inhibin, alpha | 1270 | 0.126 | -0.0315 | No |
| 6 | NOLC1 | NOLC1 Entrez, Source | nucleolar and coiled-body phosphoprotein 1 | 1277 | 0.125 | -0.0237 | No |
| 7 | CDK5RAP3 | CDK5RAP3 Entrez, Source | CDK5 regulatory subunit associated protein 3 | 1278 | 0.125 | -0.0155 | No |
| 8 | KATNA1 | KATNA1 Entrez, Source | katanin p60 (ATPase-containing) subunit A 1 | 1345 | 0.122 | -0.0126 | No |
| 9 | MAD2L2 | MAD2L2 Entrez, Source | MAD2 mitotic arrest deficient-like 2 (yeast) | 1355 | 0.121 | -0.0053 | No |
| 10 | GAS7 | GAS7 Entrez, Source | growth arrest-specific 7 | 1442 | 0.116 | -0.0042 | No |
| 11 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 1526 | 0.112 | -0.0032 | No |
| 12 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 1609 | 0.108 | -0.0023 | No |
| 13 | SUGT1 | SUGT1 Entrez, Source | SGT1, suppressor of G2 allele of SKP1 (S. cerevisiae) | 1681 | 0.106 | -0.0008 | No |
| 14 | EGF | EGF Entrez, Source | epidermal growth factor (beta-urogastrone) | 1748 | 0.103 | 0.0009 | No |
| 15 | TBRG4 | TBRG4 Entrez, Source | transforming growth factor beta regulator 4 | 2019 | 0.093 | -0.0134 | No |
| 16 | PA2G4 | PA2G4 Entrez, Source | proliferation-associated 2G4, 38kDa | 2235 | 0.087 | -0.0241 | No |
| 17 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 2426 | 0.082 | -0.0331 | No |
| 18 | GTPBP4 | GTPBP4 Entrez, Source | GTP binding protein 4 | 2484 | 0.080 | -0.0322 | No |
| 19 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 2527 | 0.079 | -0.0302 | No |
| 20 | SPDYA | SPDYA Entrez, Source | speedy homolog A (Drosophila) | 2536 | 0.079 | -0.0257 | No |
| 21 | BMP7 | BMP7 Entrez, Source | bone morphogenetic protein 7 (osteogenic protein 1) | 2669 | 0.074 | -0.0309 | No |
| 22 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 2746 | 0.073 | -0.0319 | No |
| 23 | RB1 | RB1 Entrez, Source | retinoblastoma 1 (including osteosarcoma) | 2765 | 0.072 | -0.0285 | No |
| 24 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 2800 | 0.071 | -0.0264 | No |
| 25 | EGFL6 | EGFL6 Entrez, Source | EGF-like-domain, multiple 6 | 2830 | 0.071 | -0.0240 | No |
| 26 | DUSP13 | DUSP13 Entrez, Source | dual specificity phosphatase 13 | 2997 | 0.067 | -0.0322 | No |
| 27 | RAD17 | RAD17 Entrez, Source | RAD17 homolog (S. pombe) | 3607 | 0.054 | -0.0749 | No |
| 28 | SMC1A | SMC1A Entrez, Source | structural maintenance of chromosomes 1A | 3744 | 0.051 | -0.0819 | No |
| 29 | NOTCH2 | NOTCH2 Entrez, Source | Notch homolog 2 (Drosophila) | 3989 | 0.047 | -0.0974 | No |
| 30 | ASNS | ASNS Entrez, Source | asparagine synthetase | 4245 | 0.042 | -0.1139 | No |
| 31 | CDKN3 | CDKN3 Entrez, Source | cyclin-dependent kinase inhibitor 3 (CDK2-associated dual specificity phosphatase) | 4390 | 0.040 | -0.1223 | No |
| 32 | PPP6C | PPP6C Entrez, Source | protein phosphatase 6, catalytic subunit | 4715 | 0.034 | -0.1446 | No |
| 33 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 4803 | 0.032 | -0.1491 | No |
| 34 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 5332 | 0.024 | -0.1876 | No |
| 35 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 5597 | 0.020 | -0.2063 | No |
| 36 | PTPRC | PTPRC Entrez, Source | protein tyrosine phosphatase, receptor type, C | 5608 | 0.020 | -0.2057 | No |
| 37 | PPM1G | PPM1G Entrez, Source | protein phosphatase 1G (formerly 2C), magnesium-dependent, gamma isoform | 5696 | 0.019 | -0.2111 | No |
| 38 | PPP5C | PPP5C Entrez, Source | protein phosphatase 5, catalytic subunit | 5703 | 0.019 | -0.2103 | No |
| 39 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 5720 | 0.018 | -0.2103 | No |
| 40 | LIG3 | LIG3 Entrez, Source | ligase III, DNA, ATP-dependent | 5865 | 0.016 | -0.2202 | No |
| 41 | BRCA2 | BRCA2 Entrez, Source | breast cancer 2, early onset | 5924 | 0.015 | -0.2236 | No |
| 42 | TTN | TTN Entrez, Source | titin | 5994 | 0.014 | -0.2279 | No |
| 43 | APBB1 | APBB1 Entrez, Source | amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65) | 6000 | 0.014 | -0.2273 | No |
| 44 | CDK5RAP1 | CDK5RAP1 Entrez, Source | CDK5 regulatory subunit associated protein 1 | 6027 | 0.014 | -0.2284 | No |
| 45 | AKAP8 | AKAP8 Entrez, Source | A kinase (PRKA) anchor protein 8 | 6151 | 0.012 | -0.2369 | No |
| 46 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 6158 | 0.012 | -0.2366 | No |
| 47 | CDC16 | CDC16 Entrez, Source | CDC16 cell division cycle 16 homolog (S. cerevisiae) | 6164 | 0.012 | -0.2362 | No |
| 48 | MRE11A | MRE11A Entrez, Source | MRE11 meiotic recombination 11 homolog A (S. cerevisiae) | 6221 | 0.011 | -0.2397 | No |
| 49 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 6476 | 0.007 | -0.2586 | No |
| 50 | GSPT1 | GSPT1 Entrez, Source | G1 to S phase transition 1 | 6717 | 0.004 | -0.2765 | No |
| 51 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 6755 | 0.003 | -0.2791 | No |
| 52 | NEK6 | NEK6 Entrez, Source | NIMA (never in mitosis gene a)-related kinase 6 | 6778 | 0.003 | -0.2806 | No |
| 53 | CDK10 | CDK10 Entrez, Source | cyclin-dependent kinase (CDC2-like) 10 | 6796 | 0.003 | -0.2817 | No |
| 54 | NPM2 | NPM2 Entrez, Source | nucleophosmin/nucleoplasmin, 2 | 6828 | 0.002 | -0.2839 | No |
| 55 | CHFR | CHFR Entrez, Source | checkpoint with forkhead and ring finger domains | 6893 | 0.001 | -0.2887 | No |
| 56 | DLG1 | DLG1 Entrez, Source | discs, large homolog 1 (Drosophila) | 6958 | 0.000 | -0.2935 | No |
| 57 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 6984 | -0.000 | -0.2954 | No |
| 58 | NPM1 | NPM1 Entrez, Source | nucleophosmin (nucleolar phosphoprotein B23, numatrin) | 7074 | -0.001 | -0.3021 | No |
| 59 | AIF1 | AIF1 Entrez, Source | allograft inflammatory factor 1 | 7500 | -0.007 | -0.3338 | No |
| 60 | CDC2L5 | CDC2L5 Entrez, Source | cell division cycle 2-like 5 (cholinesterase-related cell division controller) | 7727 | -0.011 | -0.3503 | No |
| 61 | CUL5 | CUL5 Entrez, Source | cullin 5 | 7748 | -0.011 | -0.3511 | No |
| 62 | CD28 | CD28 Entrez, Source | CD28 molecule | 7763 | -0.011 | -0.3514 | No |
| 63 | SNF1LK | SNF1LK Entrez, Source | SNF1-like kinase | 7785 | -0.012 | -0.3522 | No |
| 64 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 7793 | -0.012 | -0.3519 | No |
| 65 | GPS1 | GPS1 Entrez, Source | G protein pathway suppressor 1 | 7796 | -0.012 | -0.3513 | No |
| 66 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 8216 | -0.018 | -0.3819 | No |
| 67 | UHMK1 | UHMK1 Entrez, Source | U2AF homology motif (UHM) kinase 1 | 8307 | -0.020 | -0.3874 | No |
| 68 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 8319 | -0.020 | -0.3869 | No |
| 69 | CCNA1 | CCNA1 Entrez, Source | cyclin A1 | 8384 | -0.021 | -0.3904 | No |
| 70 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 8416 | -0.022 | -0.3913 | No |
| 71 | CDK7 | CDK7 Entrez, Source | cyclin-dependent kinase 7 (MO15 homolog, Xenopus laevis, cdk-activating kinase) | 8462 | -0.022 | -0.3933 | No |
| 72 | RASSF1 | RASSF1 Entrez, Source | Ras association (RalGDS/AF-6) domain family 1 | 8495 | -0.023 | -0.3942 | No |
| 73 | NBN | NBN Entrez, Source | nibrin | 8726 | -0.027 | -0.4099 | No |
| 74 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 8736 | -0.027 | -0.4088 | No |
| 75 | CETN3 | CETN3 Entrez, Source | centrin, EF-hand protein, 3 (CDC31 homolog, yeast) | 8746 | -0.027 | -0.4077 | No |
| 76 | UBB | UBB Entrez, Source | ubiquitin B | 8839 | -0.029 | -0.4128 | No |
| 77 | EIF4G2 | EIF4G2 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 2 | 8954 | -0.031 | -0.4194 | No |
| 78 | MFN2 | MFN2 Entrez, Source | mitofusin 2 | 9004 | -0.032 | -0.4210 | No |
| 79 | MAP3K11 | MAP3K11 Entrez, Source | mitogen-activated protein kinase kinase kinase 11 | 9043 | -0.033 | -0.4217 | No |
| 80 | RAN | RAN Entrez, Source | RAN, member RAS oncogene family | 9203 | -0.036 | -0.4315 | No |
| 81 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 9229 | -0.036 | -0.4310 | No |
| 82 | MAP2K6 | MAP2K6 Entrez, Source | mitogen-activated protein kinase kinase 6 | 9252 | -0.037 | -0.4302 | No |
| 83 | ZW10 | ZW10 Entrez, Source | ZW10, kinetochore associated, homolog (Drosophila) | 9270 | -0.037 | -0.4291 | No |
| 84 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 9276 | -0.037 | -0.4270 | No |
| 85 | EREG | EREG Entrez, Source | epiregulin | 9343 | -0.039 | -0.4295 | No |
| 86 | TUBG1 | TUBG1 Entrez, Source | tubulin, gamma 1 | 9358 | -0.039 | -0.4280 | No |
| 87 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 9459 | -0.041 | -0.4330 | No |
| 88 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 9513 | -0.042 | -0.4342 | No |
| 89 | DDB1 | DDB1 Entrez, Source | damage-specific DNA binding protein 1, 127kDa | 9544 | -0.042 | -0.4337 | No |
| 90 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 9699 | -0.046 | -0.4424 | No |
| 91 | SYCP1 | SYCP1 Entrez, Source | synaptonemal complex protein 1 | 9738 | -0.046 | -0.4423 | No |
| 92 | PPP1R9B | PPP1R9B Entrez, Source | protein phosphatase 1, regulatory subunit 9B, spinophilin | 9743 | -0.047 | -0.4395 | No |
| 93 | APC | APC Entrez, Source | adenomatosis polyposis coli | 9845 | -0.049 | -0.4440 | No |
| 94 | CCND3 | CCND3 Entrez, Source | cyclin D3 | 9869 | -0.049 | -0.4425 | No |
| 95 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 9909 | -0.050 | -0.4422 | No |
| 96 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 9946 | -0.051 | -0.4416 | No |
| 97 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 10008 | -0.052 | -0.4428 | No |
| 98 | CETN1 | CETN1 Entrez, Source | centrin, EF-hand protein, 1 | 10032 | -0.053 | -0.4411 | No |
| 99 | GFI1 | GFI1 Entrez, Source | growth factor independent 1 | 10061 | -0.053 | -0.4397 | No |
| 100 | HSPA2 | HSPA2 Entrez, Source | heat shock 70kDa protein 2 | 10104 | -0.054 | -0.4394 | No |
| 101 | PCAF | PCAF Entrez, Source | p300/CBP-associated factor | 10191 | -0.056 | -0.4422 | No |
| 102 | ZBTB17 | ZBTB17 Entrez, Source | zinc finger and BTB domain containing 17 | 10241 | -0.058 | -0.4421 | No |
| 103 | SC65 | SC65 Entrez, Source | - | 10286 | -0.059 | -0.4416 | No |
| 104 | DCTN2 | DCTN2 Entrez, Source | dynactin 2 (p50) | 10348 | -0.060 | -0.4423 | No |
| 105 | ANAPC4 | ANAPC4 Entrez, Source | anaphase promoting complex subunit 4 | 10467 | -0.063 | -0.4471 | No |
| 106 | CDC23 | CDC23 Entrez, Source | CDC23 (cell division cycle 23, yeast, homolog) | 10523 | -0.065 | -0.4471 | No |
| 107 | STAG3 | STAG3 Entrez, Source | stromal antigen 3 | 10647 | -0.068 | -0.4519 | Yes |
| 108 | SMC3 | SMC3 Entrez, Source | structural maintenance of chromosomes 3 | 10686 | -0.069 | -0.4503 | Yes |
| 109 | RAD50 | RAD50 Entrez, Source | RAD50 homolog (S. cerevisiae) | 10695 | -0.070 | -0.4463 | Yes |
| 110 | CDKN2B | CDKN2B Entrez, Source | cyclin-dependent kinase inhibitor 2B (p15, inhibits CDK4) | 10726 | -0.071 | -0.4440 | Yes |
| 111 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 10788 | -0.072 | -0.4439 | Yes |
| 112 | BTG4 | BTG4 Entrez, Source | B-cell translocation gene 4 | 10905 | -0.075 | -0.4477 | Yes |
| 113 | CDKN2D | CDKN2D Entrez, Source | cyclin-dependent kinase inhibitor 2D (p19, inhibits CDK4) | 10949 | -0.077 | -0.4460 | Yes |
| 114 | TBRG1 | TBRG1 Entrez, Source | transforming growth factor beta regulator 1 | 11092 | -0.082 | -0.4514 | Yes |
| 115 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 11138 | -0.083 | -0.4494 | Yes |
| 116 | USH1C | USH1C Entrez, Source | Usher syndrome 1C (autosomal recessive, severe) | 11196 | -0.086 | -0.4481 | Yes |
| 117 | ZWINT | ZWINT Entrez, Source | ZW10 interactor | 11263 | -0.089 | -0.4473 | Yes |
| 118 | STRN3 | STRN3 Entrez, Source | striatin, calmodulin binding protein 3 | 11294 | -0.090 | -0.4437 | Yes |
| 119 | SERTAD1 | SERTAD1 Entrez, Source | SERTA domain containing 1 | 11307 | -0.091 | -0.4387 | Yes |
| 120 | BTG3 | BTG3 Entrez, Source | BTG family, member 3 | 11377 | -0.094 | -0.4377 | Yes |
| 121 | AHR | AHR Entrez, Source | aryl hydrocarbon receptor | 11463 | -0.098 | -0.4378 | Yes |
| 122 | KHDRBS1 | KHDRBS1 Entrez, Source | KH domain containing, RNA binding, signal transduction associated 1 | 11520 | -0.100 | -0.4355 | Yes |
| 123 | MNAT1 | MNAT1 Entrez, Source | menage a trois homolog 1, cyclin H assembly factor (Xenopus laevis) | 11555 | -0.102 | -0.4314 | Yes |
| 124 | PLAGL1 | PLAGL1 Entrez, Source | pleiomorphic adenoma gene-like 1 | 11562 | -0.102 | -0.4251 | Yes |
| 125 | CKAP5 | CKAP5 Entrez, Source | cytoskeleton associated protein 5 | 11647 | -0.106 | -0.4245 | Yes |
| 126 | NUMA1 | NUMA1 Entrez, Source | nuclear mitotic apparatus protein 1 | 11712 | -0.110 | -0.4222 | Yes |
| 127 | CIT | CIT Entrez, Source | citron (rho-interacting, serine/threonine kinase 21) | 11752 | -0.111 | -0.4179 | Yes |
| 128 | ING4 | ING4 Entrez, Source | inhibitor of growth family, member 4 | 11779 | -0.113 | -0.4125 | Yes |
| 129 | ANAPC2 | ANAPC2 Entrez, Source | anaphase promoting complex subunit 2 | 11844 | -0.116 | -0.4097 | Yes |
| 130 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 11886 | -0.118 | -0.4051 | Yes |
| 131 | CDC27 | CDC27 Entrez, Source | cell division cycle 27 | 11905 | -0.119 | -0.3987 | Yes |
| 132 | AURKA | AURKA Entrez, Source | aurora kinase A | 11917 | -0.119 | -0.3918 | Yes |
| 133 | NDE1 | NDE1 Entrez, Source | nudE nuclear distribution gene E homolog 1 (A. nidulans) | 12075 | -0.130 | -0.3952 | Yes |
| 134 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 12112 | -0.132 | -0.3893 | Yes |
| 135 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 12256 | -0.144 | -0.3907 | Yes |
| 136 | CDK4 | CDK4 Entrez, Source | cyclin-dependent kinase 4 | 12272 | -0.146 | -0.3823 | Yes |
| 137 | MAPK12 | MAPK12 Entrez, Source | mitogen-activated protein kinase 12 | 12330 | -0.151 | -0.3768 | Yes |
| 138 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 12376 | -0.156 | -0.3700 | Yes |
| 139 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 12377 | -0.156 | -0.3598 | Yes |
| 140 | CDC37 | CDC37 Entrez, Source | CDC37 cell division cycle 37 homolog (S. cerevisiae) | 12406 | -0.158 | -0.3516 | Yes |
| 141 | CHEK2 | CHEK2 Entrez, Source | CHK2 checkpoint homolog (S. pombe) | 12445 | -0.163 | -0.3438 | Yes |
| 142 | SPHK1 | SPHK1 Entrez, Source | sphingosine kinase 1 | 12507 | -0.172 | -0.3372 | Yes |
| 143 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 12656 | -0.196 | -0.3356 | Yes |
| 144 | SH3BP4 | SH3BP4 Entrez, Source | SH3-domain binding protein 4 | 12682 | -0.201 | -0.3244 | Yes |
| 145 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 12725 | -0.209 | -0.3139 | Yes |
| 146 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 12774 | -0.218 | -0.3033 | Yes |
| 147 | BMP2 | BMP2 Entrez, Source | bone morphogenetic protein 2 | 12792 | -0.222 | -0.2900 | Yes |
| 148 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 12815 | -0.228 | -0.2768 | Yes |
| 149 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 12940 | -0.266 | -0.2688 | Yes |
| 150 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 12969 | -0.279 | -0.2526 | Yes |
| 151 | KIF22 | KIF22 Entrez, Source | kinesin family member 22 | 12985 | -0.285 | -0.2352 | Yes |
| 152 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 13078 | -0.337 | -0.2201 | Yes |
| 153 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 13080 | -0.340 | -0.1979 | Yes |
| 154 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 13085 | -0.343 | -0.1758 | Yes |
| 155 | PAM | PAM Entrez, Source | peptidylglycine alpha-amidating monooxygenase | 13160 | -0.412 | -0.1544 | Yes |
| 156 | BCAT1 | BCAT1 Entrez, Source | branched chain aminotransferase 1, cytosolic | 13181 | -0.442 | -0.1271 | Yes |
| 157 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 13211 | -0.481 | -0.0978 | Yes |
| 158 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 13302 | -0.755 | -0.0552 | Yes |
| 159 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 13314 | -0.890 | 0.0021 | Yes |