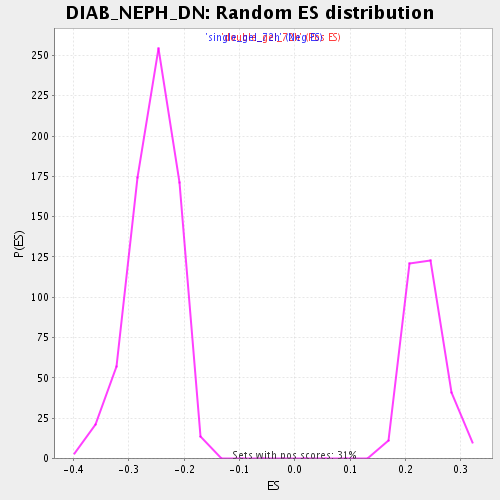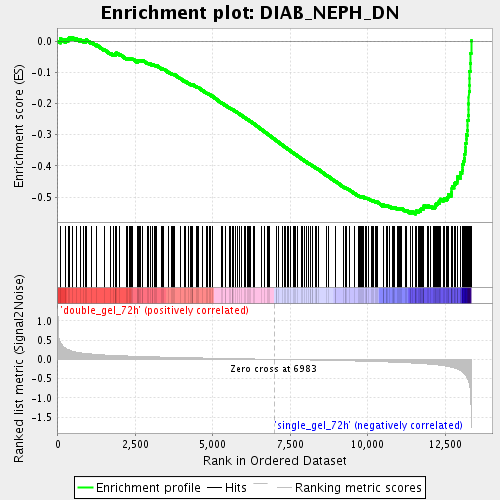
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | DIAB_NEPH_DN |
| Enrichment Score (ES) | -0.5547049 |
| Normalized Enrichment Score (NES) | -2.1734912 |
| Nominal p-value | 0.0 |
| FDR q-value | 4.987375E-4 |
| FWER p-Value | 0.0040 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FGFR2 | FGFR2 Entrez, Source | ffer syndrome, Jackson-Weiss syndrome) | 84 | 0.431 | 0.0077 | No |
| 2 | GHR | GHR Entrez, Source | growth hormone receptor | 250 | 0.287 | 0.0044 | No |
| 3 | MECP2 | MECP2 Entrez, Source | methyl CpG binding protein 2 (Rett syndrome) | 327 | 0.257 | 0.0070 | No |
| 4 | CYP51A1 | CYP51A1 Entrez, Source | cytochrome P450, family 51, subfamily A, polypeptide 1 | 366 | 0.244 | 0.0121 | No |
| 5 | IGFBP2 | IGFBP2 Entrez, Source | insulin-like growth factor binding protein 2, 36kDa | 473 | 0.209 | 0.0108 | No |
| 6 | GNE | GNE Entrez, Source | glucosamine (UDP-N-acetyl)-2-epimerase/N-acetylmannosamine kinase | 599 | 0.186 | 0.0074 | No |
| 7 | CFLAR | CFLAR Entrez, Source | CASP8 and FADD-like apoptosis regulator | 717 | 0.171 | 0.0040 | No |
| 8 | SDC2 | SDC2 Entrez, Source | syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated, fibroglycan) | 817 | 0.161 | 0.0017 | No |
| 9 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 901 | 0.153 | 0.0004 | No |
| 10 | HSPA5 | HSPA5 Entrez, Source | heat shock 70kDa protein 5 (glucose-regulated protein, 78kDa) | 909 | 0.152 | 0.0048 | No |
| 11 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 1078 | 0.138 | -0.0035 | No |
| 12 | TRIM23 | TRIM23 Entrez, Source | tripartite motif-containing 23 | 1245 | 0.127 | -0.0120 | No |
| 13 | RARRES2 | RARRES2 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 2 | 1501 | 0.113 | -0.0278 | No |
| 14 | TSPYL4 | TSPYL4 Entrez, Source | TSPY-like 4 | 1691 | 0.105 | -0.0388 | No |
| 15 | NPHS1 | NPHS1 Entrez, Source | nephrosis 1, congenital, Finnish type (nephrin) | 1792 | 0.101 | -0.0431 | No |
| 16 | OAS1 | OAS1 Entrez, Source | 2',5'-oligoadenylate synthetase 1, 40/46kDa | 1850 | 0.099 | -0.0442 | No |
| 17 | GULP1 | GULP1 Entrez, Source | GULP, engulfment adaptor PTB domain containing 1 | 1862 | 0.099 | -0.0418 | No |
| 18 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 1864 | 0.099 | -0.0387 | No |
| 19 | PBEF1 | PBEF1 Entrez, Source | pre-B-cell colony enhancing factor 1 | 1897 | 0.097 | -0.0379 | No |
| 20 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 1993 | 0.094 | -0.0421 | No |
| 21 | ARHGAP5 | ARHGAP5 Entrez, Source | Rho GTPase activating protein 5 | 2217 | 0.088 | -0.0563 | No |
| 22 | MAPK10 | MAPK10 Entrez, Source | mitogen-activated protein kinase 10 | 2261 | 0.086 | -0.0567 | No |
| 23 | WWP1 | WWP1 Entrez, Source | WW domain containing E3 ubiquitin protein ligase 1 | 2306 | 0.085 | -0.0573 | No |
| 24 | IRS1 | IRS1 Entrez, Source | insulin receptor substrate 1 | 2329 | 0.084 | -0.0562 | No |
| 25 | SLK | SLK Entrez, Source | STE20-like kinase (yeast) | 2387 | 0.083 | -0.0579 | No |
| 26 | PTPRO | PTPRO Entrez, Source | protein tyrosine phosphatase, receptor type, O | 2417 | 0.082 | -0.0574 | No |
| 27 | CEBPD | CEBPD Entrez, Source | CCAAT/enhancer binding protein (C/EBP), delta | 2567 | 0.078 | -0.0663 | No |
| 28 | OPTN | OPTN Entrez, Source | optineurin | 2577 | 0.077 | -0.0644 | No |
| 29 | CAND2 | CAND2 Entrez, Source | cullin-associated and neddylation-dissociated 2 (putative) | 2588 | 0.077 | -0.0627 | No |
| 30 | ARL1 | ARL1 Entrez, Source | ADP-ribosylation factor-like 1 | 2616 | 0.076 | -0.0623 | No |
| 31 | GMFB | GMFB Entrez, Source | glia maturation factor, beta | 2636 | 0.075 | -0.0613 | No |
| 32 | BMP7 | BMP7 Entrez, Source | bone morphogenetic protein 7 (osteogenic protein 1) | 2669 | 0.074 | -0.0613 | No |
| 33 | CDS1 | CDS1 Entrez, Source | CDP-diacylglycerol synthase (phosphatidate cytidylyltransferase) 1 | 2733 | 0.073 | -0.0637 | No |
| 34 | GRK5 | GRK5 Entrez, Source | G protein-coupled receptor kinase 5 | 2743 | 0.073 | -0.0620 | No |
| 35 | ANKRD15 | ANKRD15 Entrez, Source | ankyrin repeat domain 15 | 2894 | 0.069 | -0.0712 | No |
| 36 | CSDA | CSDA Entrez, Source | cold shock domain protein A | 2927 | 0.068 | -0.0714 | No |
| 37 | TCF21 | TCF21 Entrez, Source | transcription factor 21 | 2974 | 0.067 | -0.0728 | No |
| 38 | DNAJB9 | DNAJB9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 9 | 3036 | 0.066 | -0.0753 | No |
| 39 | SUB1 | SUB1 Entrez, Source | SUB1 homolog (S. cerevisiae) | 3048 | 0.065 | -0.0740 | No |
| 40 | SACM1L | SACM1L Entrez, Source | SAC1 suppressor of actin mutations 1-like (yeast) | 3120 | 0.064 | -0.0773 | No |
| 41 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 3161 | 0.063 | -0.0783 | No |
| 42 | DNM1L | DNM1L Entrez, Source | dynamin 1-like | 3184 | 0.063 | -0.0779 | No |
| 43 | CANX | CANX Entrez, Source | calnexin | 3355 | 0.059 | -0.0890 | No |
| 44 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 3374 | 0.059 | -0.0884 | No |
| 45 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 3416 | 0.058 | -0.0897 | No |
| 46 | PEX14 | PEX14 Entrez, Source | peroxisomal biogenesis factor 14 | 3565 | 0.054 | -0.0992 | No |
| 47 | UBE2L6 | UBE2L6 Entrez, Source | ubiquitin-conjugating enzyme E2L 6 | 3662 | 0.053 | -0.1048 | No |
| 48 | AKAP11 | AKAP11 Entrez, Source | A kinase (PRKA) anchor protein 11 | 3700 | 0.052 | -0.1059 | No |
| 49 | CSPG2 | CSPG2 Entrez, Source | chondroitin sulfate proteoglycan 2 (versican) | 3728 | 0.051 | -0.1063 | No |
| 50 | TMEM23 | TMEM23 Entrez, Source | transmembrane protein 23 | 3767 | 0.050 | -0.1076 | No |
| 51 | ADORA1 | ADORA1 Entrez, Source | adenosine A1 receptor | 3951 | 0.047 | -0.1200 | No |
| 52 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 4076 | 0.045 | -0.1280 | No |
| 53 | DPP6 | DPP6 Entrez, Source | dipeptidyl-peptidase 6 | 4123 | 0.044 | -0.1301 | No |
| 54 | KLK7 | KLK7 Entrez, Source | kallikrein 7 (chymotryptic, stratum corneum) | 4220 | 0.043 | -0.1360 | No |
| 55 | NAB1 | NAB1 Entrez, Source | NGFI-A binding protein 1 (EGR1 binding protein 1) | 4286 | 0.041 | -0.1396 | No |
| 56 | PSMA6 | PSMA6 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 6 | 4318 | 0.041 | -0.1406 | No |
| 57 | GRIK2 | GRIK2 Entrez, Source | glutamate receptor, ionotropic, kainate 2 | 4329 | 0.041 | -0.1401 | No |
| 58 | XRCC5 | XRCC5 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining; Ku autoantigen, 80kDa) | 4343 | 0.040 | -0.1397 | No |
| 59 | COL4A4 | COL4A4 Entrez, Source | collagen, type IV, alpha 4 | 4358 | 0.040 | -0.1395 | No |
| 60 | NAPG | NAPG Entrez, Source | N-ethylmaleimide-sensitive factor attachment protein, gamma | 4467 | 0.038 | -0.1465 | No |
| 61 | KLK6 | KLK6 Entrez, Source | kallikrein 6 (neurosin, zyme) | 4508 | 0.038 | -0.1483 | No |
| 62 | OTUD4 | OTUD4 Entrez, Source | OTU domain containing 4 | 4532 | 0.037 | -0.1489 | No |
| 63 | PCMT1 | PCMT1 Entrez, Source | protein-L-isoaspartate (D-aspartate) O-methyltransferase | 4670 | 0.035 | -0.1582 | No |
| 64 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 4803 | 0.032 | -0.1672 | No |
| 65 | HNRPDL | HNRPDL Entrez, Source | heterogeneous nuclear ribonucleoprotein D-like | 4822 | 0.032 | -0.1675 | No |
| 66 | TNNT2 | TNNT2 Entrez, Source | troponin T type 2 (cardiac) | 4891 | 0.031 | -0.1717 | No |
| 67 | RAD23B | RAD23B Entrez, Source | RAD23 homolog B (S. cerevisiae) | 4914 | 0.031 | -0.1724 | No |
| 68 | LOXL1 | LOXL1 Entrez, Source | lysyl oxidase-like 1 | 4974 | 0.030 | -0.1759 | No |
| 69 | STX7 | STX7 Entrez, Source | syntaxin 7 | 5278 | 0.025 | -0.1982 | No |
| 70 | ARF6 | ARF6 Entrez, Source | ADP-ribosylation factor 6 | 5321 | 0.024 | -0.2006 | No |
| 71 | ADD3 | ADD3 Entrez, Source | adducin 3 (gamma) | 5403 | 0.023 | -0.2061 | No |
| 72 | WT1 | WT1 Entrez, Source | Wilms tumor 1 | 5408 | 0.023 | -0.2056 | No |
| 73 | EXOSC7 | EXOSC7 Entrez, Source | exosome component 7 | 5539 | 0.021 | -0.2148 | No |
| 74 | FNBP1 | FNBP1 Entrez, Source | formin binding protein 1 | 5572 | 0.021 | -0.2166 | No |
| 75 | UGP2 | UGP2 Entrez, Source | UDP-glucose pyrophosphorylase 2 | 5626 | 0.020 | -0.2200 | No |
| 76 | SULF1 | SULF1 Entrez, Source | sulfatase 1 | 5633 | 0.020 | -0.2198 | No |
| 77 | COLQ | COLQ Entrez, Source | collagen-like tail subunit (single strand of homotrimer) of asymmetric acetylcholinesterase | 5644 | 0.020 | -0.2199 | No |
| 78 | CD164 | CD164 Entrez, Source | CD164 molecule, sialomucin | 5676 | 0.019 | -0.2217 | No |
| 79 | ST3GAL6 | ST3GAL6 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 6 | 5745 | 0.018 | -0.2263 | No |
| 80 | BST2 | BST2 Entrez, Source | bone marrow stromal cell antigen 2 | 5794 | 0.017 | -0.2294 | No |
| 81 | PSMA2 | PSMA2 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 2 | 5869 | 0.016 | -0.2345 | No |
| 82 | TBCA | TBCA Entrez, Source | tubulin-specific chaperone a | 5909 | 0.016 | -0.2370 | No |
| 83 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 6026 | 0.014 | -0.2454 | No |
| 84 | HNRPA1 | HNRPA1 Entrez, Source | heterogeneous nuclear ribonucleoprotein A1 | 6063 | 0.013 | -0.2477 | No |
| 85 | GJA1 | GJA1 Entrez, Source | gap junction protein, alpha 1, 43kDa (connexin 43) | 6120 | 0.013 | -0.2515 | No |
| 86 | RHOBTB2 | RHOBTB2 Entrez, Source | Rho-related BTB domain containing 2 | 6138 | 0.012 | -0.2524 | No |
| 87 | TMED10 | TMED10 Entrez, Source | transmembrane emp24-like trafficking protein 10 (yeast) | 6185 | 0.012 | -0.2556 | No |
| 88 | MTX2 | MTX2 Entrez, Source | metaxin 2 | 6206 | 0.011 | -0.2567 | No |
| 89 | PTPN7 | PTPN7 Entrez, Source | protein tyrosine phosphatase, non-receptor type 7 | 6306 | 0.009 | -0.2640 | No |
| 90 | NMI | NMI Entrez, Source | N-myc (and STAT) interactor | 6331 | 0.009 | -0.2655 | No |
| 91 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 6567 | 0.006 | -0.2833 | No |
| 92 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 6673 | 0.004 | -0.2911 | No |
| 93 | PRDX6 | PRDX6 Entrez, Source | peroxiredoxin 6 | 6768 | 0.003 | -0.2982 | No |
| 94 | TAX1BP1 | TAX1BP1 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 1 | 6803 | 0.003 | -0.3007 | No |
| 95 | MGAT5 | MGAT5 Entrez, Source | mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase | 6814 | 0.002 | -0.3014 | No |
| 96 | TOMM20 | TOMM20 Entrez, Source | translocase of outer mitochondrial membrane 20 homolog (yeast) | 7046 | -0.001 | -0.3190 | No |
| 97 | RHEB | RHEB Entrez, Source | Ras homolog enriched in brain | 7054 | -0.001 | -0.3195 | No |
| 98 | RAB1A | RAB1A Entrez, Source | RAB1A, member RAS oncogene family | 7133 | -0.002 | -0.3254 | No |
| 99 | CMAH | CMAH Entrez, Source | cytidine monophosphate-N-acetylneuraminic acid hydroxylase (CMP-N-acetylneuraminate monooxygenase) | 7234 | -0.004 | -0.3329 | No |
| 100 | PHYHIP | PHYHIP Entrez, Source | phytanoyl-CoA 2-hydroxylase interacting protein | 7298 | -0.005 | -0.3376 | No |
| 101 | LANCL1 | LANCL1 Entrez, Source | LanC lantibiotic synthetase component C-like 1 (bacterial) | 7352 | -0.005 | -0.3414 | No |
| 102 | SKP1A | SKP1A Entrez, Source | S-phase kinase-associated protein 1A (p19A) | 7418 | -0.006 | -0.3462 | No |
| 103 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 7445 | -0.007 | -0.3480 | No |
| 104 | AIF1 | AIF1 Entrez, Source | allograft inflammatory factor 1 | 7500 | -0.007 | -0.3518 | No |
| 105 | GLUL | GLUL Entrez, Source | glutamate-ammonia ligase (glutamine synthetase) | 7612 | -0.009 | -0.3600 | No |
| 106 | PPFIA4 | PPFIA4 Entrez, Source | protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 4 | 7620 | -0.009 | -0.3602 | No |
| 107 | NDN | NDN Entrez, Source | necdin homolog (mouse) | 7672 | -0.010 | -0.3638 | No |
| 108 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 7735 | -0.011 | -0.3682 | No |
| 109 | DDN | DDN Entrez, Source | dendrin | 7856 | -0.013 | -0.3769 | No |
| 110 | TSC22D3 | TSC22D3 Entrez, Source | TSC22 domain family, member 3 | 7885 | -0.013 | -0.3786 | No |
| 111 | VPS26A | VPS26A Entrez, Source | vacuolar protein sorting 26 homolog A (yeast) | 7970 | -0.015 | -0.3845 | No |
| 112 | CACNB2 | CACNB2 Entrez, Source | calcium channel, voltage-dependent, beta 2 subunit | 8013 | -0.015 | -0.3873 | No |
| 113 | PTPN11 | PTPN11 Entrez, Source | protein tyrosine phosphatase, non-receptor type 11 (Noonan syndrome 1) | 8096 | -0.016 | -0.3930 | No |
| 114 | MYO1B | MYO1B Entrez, Source | myosin IB | 8141 | -0.017 | -0.3958 | No |
| 115 | YWHAE | YWHAE Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, epsilon polypeptide | 8149 | -0.017 | -0.3958 | No |
| 116 | ENPEP | ENPEP Entrez, Source | glutamyl aminopeptidase (aminopeptidase A) | 8208 | -0.018 | -0.3996 | No |
| 117 | PARP1 | PARP1 Entrez, Source | poly (ADP-ribose) polymerase family, member 1 | 8224 | -0.018 | -0.4001 | No |
| 118 | ITGB5 | ITGB5 Entrez, Source | integrin, beta 5 | 8308 | -0.020 | -0.4058 | No |
| 119 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 8340 | -0.020 | -0.4075 | No |
| 120 | ACBD3 | ACBD3 Entrez, Source | acyl-Coenzyme A binding domain containing 3 | 8347 | -0.021 | -0.4073 | No |
| 121 | AXL | AXL Entrez, Source | AXL receptor tyrosine kinase | 8404 | -0.021 | -0.4109 | No |
| 122 | GAB2 | GAB2 Entrez, Source | GRB2-associated binding protein 2 | 8676 | -0.026 | -0.4307 | No |
| 123 | NBN | NBN Entrez, Source | nibrin | 8726 | -0.027 | -0.4336 | No |
| 124 | HMGN3 | HMGN3 Entrez, Source | high mobility group nucleosomal binding domain 3 | 8947 | -0.031 | -0.4494 | No |
| 125 | HNRPC | HNRPC Entrez, Source | heterogeneous nuclear ribonucleoprotein C (C1/C2) | 8965 | -0.032 | -0.4496 | No |
| 126 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 9229 | -0.036 | -0.4685 | No |
| 127 | PFN2 | PFN2 Entrez, Source | profilin 2 | 9269 | -0.037 | -0.4703 | No |
| 128 | PAK1 | PAK1 Entrez, Source | p21/Cdc42/Rac1-activated kinase 1 (STE20 homolog, yeast) | 9287 | -0.038 | -0.4703 | No |
| 129 | DNAJA1 | DNAJA1 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 1 | 9325 | -0.038 | -0.4719 | No |
| 130 | TMED5 | TMED5 Entrez, Source | transmembrane emp24 protein transport domain containing 5 | 9395 | -0.040 | -0.4759 | No |
| 131 | RDX | RDX Entrez, Source | radixin | 9584 | -0.043 | -0.4888 | No |
| 132 | PDIA3 | PDIA3 Entrez, Source | protein disulfide isomerase family A, member 3 | 9691 | -0.045 | -0.4954 | No |
| 133 | APOD | APOD Entrez, Source | apolipoprotein D | 9731 | -0.046 | -0.4969 | No |
| 134 | BIRC2 | BIRC2 Entrez, Source | baculoviral IAP repeat-containing 2 | 9776 | -0.047 | -0.4987 | No |
| 135 | ATP2A2 | ATP2A2 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 | 9784 | -0.048 | -0.4977 | No |
| 136 | PLCG2 | PLCG2 Entrez, Source | phospholipase C, gamma 2 (phosphatidylinositol-specific) | 9827 | -0.048 | -0.4993 | No |
| 137 | GMDS | GMDS Entrez, Source | GDP-mannose 4,6-dehydratase | 9847 | -0.049 | -0.4992 | No |
| 138 | SYNPO | SYNPO Entrez, Source | synaptopodin | 9931 | -0.051 | -0.5039 | No |
| 139 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 9946 | -0.051 | -0.5033 | No |
| 140 | MAGI2 | MAGI2 Entrez, Source | membrane associated guanylate kinase, WW and PDZ domain containing 2 | 9967 | -0.051 | -0.5031 | No |
| 141 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 10008 | -0.052 | -0.5045 | No |
| 142 | EPS15 | EPS15 Entrez, Source | epidermal growth factor receptor pathway substrate 15 | 10115 | -0.055 | -0.5108 | No |
| 143 | FGF9 | FGF9 Entrez, Source | fibroblast growth factor 9 (glia-activating factor) | 10145 | -0.055 | -0.5112 | No |
| 144 | AQP3 | AQP3 Entrez, Source | aquaporin 3 (Gill blood group) | 10174 | -0.056 | -0.5115 | No |
| 145 | PRKAR1A | PRKAR1A Entrez, Source | protein kinase, cAMP-dependent, regulatory, type I, alpha (tissue specific extinguisher 1) | 10251 | -0.058 | -0.5154 | No |
| 146 | HMGN1 | HMGN1 Entrez, Source | high-mobility group nucleosome binding domain 1 | 10290 | -0.059 | -0.5164 | No |
| 147 | PLXNB2 | PLXNB2 Entrez, Source | plexin B2 | 10304 | -0.059 | -0.5154 | No |
| 148 | PTHR1 | PTHR1 Entrez, Source | parathyroid hormone receptor 1 | 10494 | -0.064 | -0.5278 | No |
| 149 | PLSCR1 | PLSCR1 Entrez, Source | phospholipid scramblase 1 | 10496 | -0.064 | -0.5257 | No |
| 150 | SLIT1 | SLIT1 Entrez, Source | slit homolog 1 (Drosophila) | 10513 | -0.064 | -0.5249 | No |
| 151 | TAPBP | TAPBP Entrez, Source | TAP binding protein (tapasin) | 10601 | -0.067 | -0.5293 | No |
| 152 | SEPT2 | SEPT2 Entrez, Source | septin 2 | 10614 | -0.067 | -0.5280 | No |
| 153 | PARD3 | PARD3 Entrez, Source | par-3 partitioning defective 3 homolog (C. elegans) | 10642 | -0.068 | -0.5279 | No |
| 154 | HSP90AA1 | HSP90AA1 Entrez, Source | heat shock protein 90kDa alpha (cytosolic), class A member 1 | 10705 | -0.070 | -0.5303 | No |
| 155 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 10801 | -0.073 | -0.5352 | No |
| 156 | PDE4B | PDE4B Entrez, Source | phosphodiesterase 4B, cAMP-specific (phosphodiesterase E4 dunce homolog, Drosophila) | 10834 | -0.073 | -0.5353 | No |
| 157 | CTBP1 | CTBP1 Entrez, Source | C-terminal binding protein 1 | 10835 | -0.073 | -0.5329 | No |
| 158 | LITAF | LITAF Entrez, Source | lipopolysaccharide-induced TNF factor | 10871 | -0.074 | -0.5331 | No |
| 159 | CREB3L2 | CREB3L2 Entrez, Source | cAMP responsive element binding protein 3-like 2 | 10974 | -0.078 | -0.5384 | No |
| 160 | LPL | LPL Entrez, Source | lipoprotein lipase | 10988 | -0.078 | -0.5368 | No |
| 161 | GTF2A2 | GTF2A2 Entrez, Source | general transcription factor IIA, 2, 12kDa | 11025 | -0.079 | -0.5369 | No |
| 162 | ATXN10 | ATXN10 Entrez, Source | ataxin 10 | 11049 | -0.080 | -0.5361 | No |
| 163 | PPP2CB | PPP2CB Entrez, Source | protein phosphatase 2 (formerly 2A), catalytic subunit, beta isoform | 11093 | -0.082 | -0.5367 | No |
| 164 | EXOC7 | EXOC7 Entrez, Source | exocyst complex component 7 | 11206 | -0.086 | -0.5424 | No |
| 165 | CAST | CAST Entrez, Source | calpastatin | 11239 | -0.087 | -0.5420 | No |
| 166 | NPTN | NPTN Entrez, Source | neuroplastin | 11383 | -0.094 | -0.5499 | No |
| 167 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 11390 | -0.094 | -0.5472 | No |
| 168 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 11431 | -0.096 | -0.5471 | No |
| 169 | KIF5B | KIF5B Entrez, Source | kinesin family member 5B | 11531 | -0.101 | -0.5514 | Yes |
| 170 | TGFBR3 | TGFBR3 Entrez, Source | transforming growth factor, beta receptor III (betaglycan, 300kDa) | 11551 | -0.102 | -0.5495 | Yes |
| 171 | ATP6AP2 | ATP6AP2 Entrez, Source | ATPase, H+ transporting, lysosomal accessory protein 2 | 11554 | -0.102 | -0.5464 | Yes |
| 172 | RAB21 | RAB21 Entrez, Source | RAB21, member RAS oncogene family | 11573 | -0.103 | -0.5444 | Yes |
| 173 | BIRC3 | BIRC3 Entrez, Source | baculoviral IAP repeat-containing 3 | 11637 | -0.106 | -0.5457 | Yes |
| 174 | NTRK2 | NTRK2 Entrez, Source | neurotrophic tyrosine kinase, receptor, type 2 | 11666 | -0.108 | -0.5444 | Yes |
| 175 | ING1 | ING1 Entrez, Source | inhibitor of growth family, member 1 | 11699 | -0.109 | -0.5432 | Yes |
| 176 | BCAM | BCAM Entrez, Source | basal cell adhesion molecule (Lutheran blood group) | 11705 | -0.109 | -0.5401 | Yes |
| 177 | YWHAZ | YWHAZ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide | 11724 | -0.110 | -0.5378 | Yes |
| 178 | GSTA4 | GSTA4 Entrez, Source | glutathione S-transferase A4 | 11757 | -0.112 | -0.5366 | Yes |
| 179 | CXADR | CXADR Entrez, Source | coxsackie virus and adenovirus receptor | 11788 | -0.113 | -0.5352 | Yes |
| 180 | PSMB9 | PSMB9 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 9 (large multifunctional peptidase 2) | 11801 | -0.114 | -0.5324 | Yes |
| 181 | AGRIN | AGRIN Entrez, Source | agrin | 11810 | -0.114 | -0.5293 | Yes |
| 182 | DAG1 | DAG1 Entrez, Source | dystroglycan 1 (dystrophin-associated glycoprotein 1) | 11814 | -0.114 | -0.5258 | Yes |
| 183 | CAPN2 | CAPN2 Entrez, Source | calpain 2, (m/II) large subunit | 11927 | -0.120 | -0.5304 | Yes |
| 184 | NEDD9 | NEDD9 Entrez, Source | neural precursor cell expressed, developmentally down-regulated 9 | 11950 | -0.122 | -0.5282 | Yes |
| 185 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 12036 | -0.127 | -0.5305 | Yes |
| 186 | RGS10 | RGS10 Entrez, Source | regulator of G-protein signalling 10 | 12133 | -0.133 | -0.5335 | Yes |
| 187 | HTRA1 | HTRA1 Entrez, Source | HtrA serine peptidase 1 | 12167 | -0.136 | -0.5316 | Yes |
| 188 | PTGER4 | PTGER4 Entrez, Source | prostaglandin E receptor 4 (subtype EP4) | 12179 | -0.137 | -0.5279 | Yes |
| 189 | GYG1 | GYG1 Entrez, Source | glycogenin 1 | 12194 | -0.139 | -0.5245 | Yes |
| 190 | PTPRD | PTPRD Entrez, Source | protein tyrosine phosphatase, receptor type, D | 12216 | -0.141 | -0.5215 | Yes |
| 191 | PAWR | PAWR Entrez, Source | PRKC, apoptosis, WT1, regulator | 12264 | -0.145 | -0.5203 | Yes |
| 192 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 12269 | -0.146 | -0.5159 | Yes |
| 193 | PLS3 | PLS3 Entrez, Source | plastin 3 (T isoform) | 12321 | -0.150 | -0.5149 | Yes |
| 194 | NFASC | NFASC Entrez, Source | neurofascin homolog (chicken) | 12327 | -0.150 | -0.5103 | Yes |
| 195 | CALM2 | CALM2 Entrez, Source | calmodulin 2 (phosphorylase kinase, delta) | 12340 | -0.152 | -0.5063 | Yes |
| 196 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 12432 | -0.162 | -0.5079 | Yes |
| 197 | ARMCX2 | ARMCX2 Entrez, Source | armadillo repeat containing, X-linked 2 | 12467 | -0.166 | -0.5051 | Yes |
| 198 | COL3A1 | COL3A1 Entrez, Source | collagen, type III, alpha 1 (Ehlers-Danlos syndrome type IV, autosomal dominant) | 12541 | -0.177 | -0.5049 | Yes |
| 199 | PLEKHC1 | PLEKHC1 Entrez, Source | pleckstrin homology domain containing, family C (with FERM domain) member 1 | 12574 | -0.181 | -0.5015 | Yes |
| 200 | CBLB | CBLB Entrez, Source | Cas-Br-M (murine) ecotropic retroviral transforming sequence b | 12594 | -0.184 | -0.4969 | Yes |
| 201 | ABI1 | ABI1 Entrez, Source | abl-interactor 1 | 12600 | -0.184 | -0.4913 | Yes |
| 202 | CALD1 | CALD1 Entrez, Source | caldesmon 1 | 12690 | -0.203 | -0.4914 | Yes |
| 203 | CD47 | CD47 Entrez, Source | CD47 molecule | 12700 | -0.205 | -0.4854 | Yes |
| 204 | DSTN | DSTN Entrez, Source | destrin (actin depolymerizing factor) | 12715 | -0.208 | -0.4797 | Yes |
| 205 | PRKCI | PRKCI Entrez, Source | protein kinase C, iota | 12716 | -0.208 | -0.4729 | Yes |
| 206 | MYO1E | MYO1E Entrez, Source | myosin IE | 12726 | -0.209 | -0.4668 | Yes |
| 207 | ALDH3B1 | ALDH3B1 Entrez, Source | aldehyde dehydrogenase 3 family, member B1 | 12784 | -0.220 | -0.4640 | Yes |
| 208 | BMP2 | BMP2 Entrez, Source | bone morphogenetic protein 2 | 12792 | -0.222 | -0.4572 | Yes |
| 209 | PODXL | PODXL Entrez, Source | podocalyxin-like | 12834 | -0.233 | -0.4528 | Yes |
| 210 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 12881 | -0.245 | -0.4483 | Yes |
| 211 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 12889 | -0.247 | -0.4408 | Yes |
| 212 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 12906 | -0.254 | -0.4337 | Yes |
| 213 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 12984 | -0.284 | -0.4303 | Yes |
| 214 | RAI14 | RAI14 Entrez, Source | retinoic acid induced 14 | 12994 | -0.287 | -0.4216 | Yes |
| 215 | PRKAR2B | PRKAR2B Entrez, Source | protein kinase, cAMP-dependent, regulatory, type II, beta | 13042 | -0.314 | -0.4149 | Yes |
| 216 | TMEM123 | TMEM123 Entrez, Source | transmembrane protein 123 | 13062 | -0.326 | -0.4057 | Yes |
| 217 | CLIC5 | CLIC5 Entrez, Source | chloride intracellular channel 5 | 13065 | -0.327 | -0.3952 | Yes |
| 218 | GPNMB | GPNMB Entrez, Source | glycoprotein (transmembrane) nmb | 13090 | -0.347 | -0.3857 | Yes |
| 219 | PLCE1 | PLCE1 Entrez, Source | phospholipase C, epsilon 1 | 13110 | -0.363 | -0.3753 | Yes |
| 220 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 13125 | -0.378 | -0.3640 | Yes |
| 221 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 13140 | -0.392 | -0.3523 | Yes |
| 222 | DPYSL3 | DPYSL3 Entrez, Source | dihydropyrimidinase-like 3 | 13155 | -0.408 | -0.3401 | Yes |
| 223 | PAM | PAM Entrez, Source | peptidylglycine alpha-amidating monooxygenase | 13160 | -0.412 | -0.3269 | Yes |
| 224 | PDGFA | PDGFA Entrez, Source | platelet-derived growth factor alpha polypeptide | 13176 | -0.433 | -0.3139 | Yes |
| 225 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 13180 | -0.438 | -0.2998 | Yes |
| 226 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 13211 | -0.481 | -0.2864 | Yes |
| 227 | SERINC5 | SERINC5 Entrez, Source | serine incorporator 5 | 13225 | -0.503 | -0.2710 | Yes |
| 228 | MME | MME Entrez, Source | membrane metallo-endopeptidase (neutral endopeptidase, enkephalinase) | 13226 | -0.505 | -0.2545 | Yes |
| 229 | TSPAN3 | TSPAN3 Entrez, Source | tetraspanin 3 | 13247 | -0.547 | -0.2382 | Yes |
| 230 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13260 | -0.571 | -0.2204 | Yes |
| 231 | PDLIM5 | PDLIM5 Entrez, Source | PDZ and LIM domain 5 | 13261 | -0.575 | -0.2016 | Yes |
| 232 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 13266 | -0.607 | -0.1821 | Yes |
| 233 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13267 | -0.614 | -0.1621 | Yes |
| 234 | F2R | F2R Entrez, Source | coagulation factor II (thrombin) receptor | 13282 | -0.665 | -0.1415 | Yes |
| 235 | ADM | ADM Entrez, Source | adrenomedullin | 13287 | -0.697 | -0.1190 | Yes |
| 236 | GPRC5A | GPRC5A Entrez, Source | G protein-coupled receptor, family C, group 5, member A | 13289 | -0.705 | -0.0961 | Yes |
| 237 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 13308 | -0.800 | -0.0714 | Yes |
| 238 | IGFBP7 | IGFBP7 Entrez, Source | insulin-like growth factor binding protein 7 | 13322 | -0.989 | -0.0401 | Yes |
| 239 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 13336 | -1.272 | 0.0005 | Yes |